Dakai Jin
MUSE: Multi-Scale Dense Self-Distillation for Nucleus Detection and Classification
Nov 07, 2025



Abstract:Nucleus detection and classification (NDC) in histopathology analysis is a fundamental task that underpins a wide range of high-level pathology applications. However, existing methods heavily rely on labor-intensive nucleus-level annotations and struggle to fully exploit large-scale unlabeled data for learning discriminative nucleus representations. In this work, we propose MUSE (MUlti-scale denSE self-distillation), a novel self-supervised learning method tailored for NDC. At its core is NuLo (Nucleus-based Local self-distillation), a coordinate-guided mechanism that enables flexible local self-distillation based on predicted nucleus positions. By removing the need for strict spatial alignment between augmented views, NuLo allows critical cross-scale alignment, thus unlocking the capacity of models for fine-grained nucleus-level representation. To support MUSE, we design a simple yet effective encoder-decoder architecture and a large field-of-view semi-supervised fine-tuning strategy that together maximize the value of unlabeled pathology images. Extensive experiments on three widely used benchmarks demonstrate that MUSE effectively addresses the core challenges of histopathological NDC. The resulting models not only surpass state-of-the-art supervised baselines but also outperform generic pathology foundation models.
Anatomy-Aware Low-Dose CT Denoising via Pretrained Vision Models and Semantic-Guided Contrastive Learning
Aug 11, 2025Abstract:To reduce radiation exposure and improve the diagnostic efficacy of low-dose computed tomography (LDCT), numerous deep learning-based denoising methods have been developed to mitigate noise and artifacts. However, most of these approaches ignore the anatomical semantics of human tissues, which may potentially result in suboptimal denoising outcomes. To address this problem, we propose ALDEN, an anatomy-aware LDCT denoising method that integrates semantic features of pretrained vision models (PVMs) with adversarial and contrastive learning. Specifically, we introduce an anatomy-aware discriminator that dynamically fuses hierarchical semantic features from reference normal-dose CT (NDCT) via cross-attention mechanisms, enabling tissue-specific realism evaluation in the discriminator. In addition, we propose a semantic-guided contrastive learning module that enforces anatomical consistency by contrasting PVM-derived features from LDCT, denoised CT and NDCT, preserving tissue-specific patterns through positive pairs and suppressing artifacts via dual negative pairs. Extensive experiments conducted on two LDCT denoising datasets reveal that ALDEN achieves the state-of-the-art performance, offering superior anatomy preservation and substantially reducing over-smoothing issue of previous work. Further validation on a downstream multi-organ segmentation task (encompassing 117 anatomical structures) affirms the model's ability to maintain anatomical awareness.
HarmonySeg: Tubular Structure Segmentation with Deep-Shallow Feature Fusion and Growth-Suppression Balanced Loss
Apr 10, 2025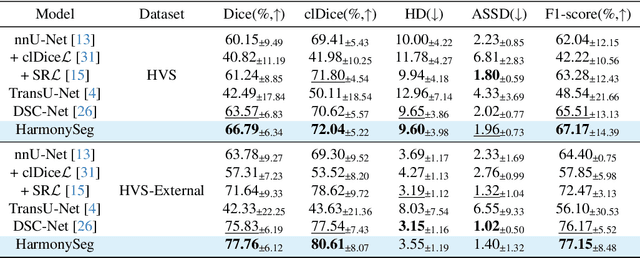
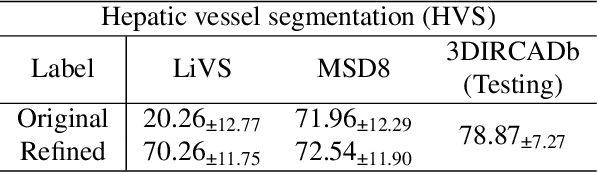
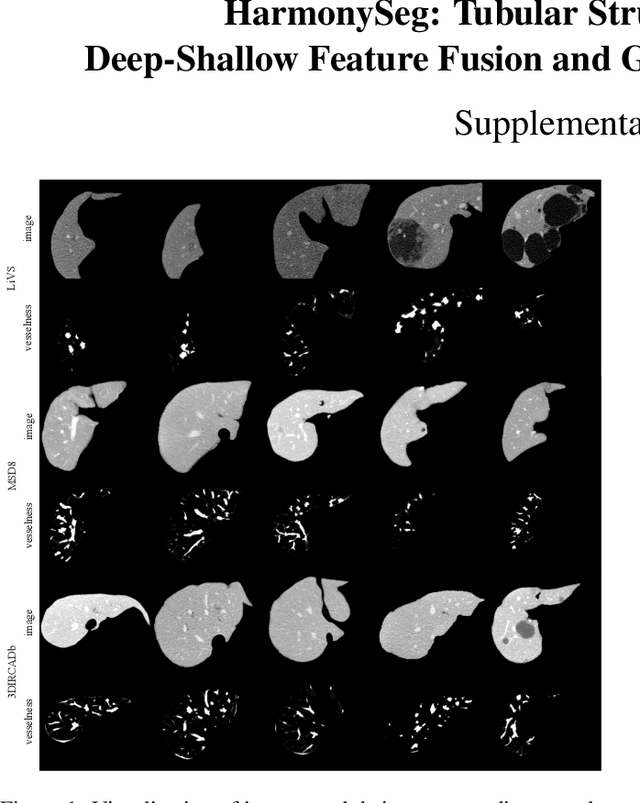

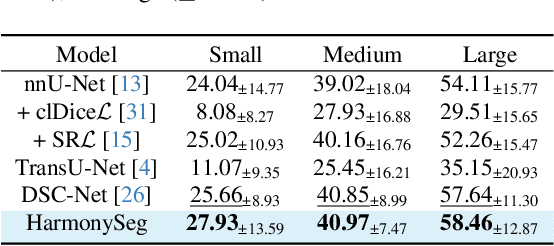
Abstract:Accurate segmentation of tubular structures in medical images, such as vessels and airway trees, is crucial for computer-aided diagnosis, radiotherapy, and surgical planning. However, significant challenges exist in algorithm design when faced with diverse sizes, complex topologies, and (often) incomplete data annotation of these structures. We address these difficulties by proposing a new tubular structure segmentation framework named HarmonySeg. First, we design a deep-to-shallow decoder network featuring flexible convolution blocks with varying receptive fields, which enables the model to effectively adapt to tubular structures of different scales. Second, to highlight potential anatomical regions and improve the recall of small tubular structures, we incorporate vesselness maps as auxiliary information. These maps are aligned with image features through a shallow-and-deep fusion module, which simultaneously eliminates unreasonable candidates to maintain high precision. Finally, we introduce a topology-preserving loss function that leverages contextual and shape priors to balance the growth and suppression of tubular structures, which also allows the model to handle low-quality and incomplete annotations. Extensive quantitative experiments are conducted on four public datasets. The results show that our model can accurately segment 2D and 3D tubular structures and outperform existing state-of-the-art methods. External validation on a private dataset also demonstrates good generalizability.
UniReg: Foundation Model for Controllable Medical Image Registration
Mar 17, 2025



Abstract:Learning-based medical image registration has achieved performance parity with conventional methods while demonstrating a substantial advantage in computational efficiency. However, learning-based registration approaches lack generalizability across diverse clinical scenarios, requiring the laborious development of multiple isolated networks for specific registration tasks, e.g., inter-/intra-subject registration or organ-specific alignment. % To overcome this limitation, we propose \textbf{UniReg}, the first interactive foundation model for medical image registration, which combines the precision advantages of task-specific learning methods with the generalization of traditional optimization methods. Our key innovation is a unified framework for diverse registration scenarios, achieved through a conditional deformation field estimation within a unified registration model. This is realized through a dynamic learning paradigm that explicitly encodes: (1) anatomical structure priors, (2) registration type constraints (inter/intra-subject), and (3) instance-specific features, enabling the generation of scenario-optimal deformation fields. % Through comprehensive experiments encompassing $90$ anatomical structures at different body regions, our UniReg model demonstrates comparable performance with contemporary state-of-the-art methodologies while achieving ~50\% reduction in required training iterations relative to the conventional learning-based paradigm. This optimization contributes to a significant reduction in computational resources, such as training time. Code and model will be available.
A Continual Learning-driven Model for Accurate and Generalizable Segmentation of Clinically Comprehensive and Fine-grained Whole-body Anatomies in CT
Mar 16, 2025Abstract:Precision medicine in the quantitative management of chronic diseases and oncology would be greatly improved if the Computed Tomography (CT) scan of any patient could be segmented, parsed and analyzed in a precise and detailed way. However, there is no such fully annotated CT dataset with all anatomies delineated for training because of the exceptionally high manual cost, the need for specialized clinical expertise, and the time required to finish the task. To this end, we proposed a novel continual learning-driven CT model that can segment complete anatomies presented using dozens of previously partially labeled datasets, dynamically expanding its capacity to segment new ones without compromising previously learned organ knowledge. Existing multi-dataset approaches are not able to dynamically segment new anatomies without catastrophic forgetting and would encounter optimization difficulty or infeasibility when segmenting hundreds of anatomies across the whole range of body regions. Our single unified CT segmentation model, CL-Net, can highly accurately segment a clinically comprehensive set of 235 fine-grained whole-body anatomies. Composed of a universal encoder, multiple optimized and pruned decoders, CL-Net is developed using 13,952 CT scans from 20 public and 16 private high-quality partially labeled CT datasets of various vendors, different contrast phases, and pathologies. Extensive evaluation demonstrates that CL-Net consistently outperforms the upper limit of an ensemble of 36 specialist nnUNets trained per dataset with the complexity of 5% model size and significantly surpasses the segmentation accuracy of recent leading Segment Anything-style medical image foundation models by large margins. Our continual learning-driven CL-Net model would lay a solid foundation to facilitate many downstream tasks of oncology and chronic diseases using the most widely adopted CT imaging.
From Slices to Sequences: Autoregressive Tracking Transformer for Cohesive and Consistent 3D Lymph Node Detection in CT Scans
Mar 11, 2025



Abstract:Lymph node (LN) assessment is an essential task in the routine radiology workflow, providing valuable insights for cancer staging, treatment planning and beyond. Identifying scatteredly-distributed and low-contrast LNs in 3D CT scans is highly challenging, even for experienced clinicians. Previous lesion and LN detection methods demonstrate effectiveness of 2.5D approaches (i.e, using 2D network with multi-slice inputs), leveraging pretrained 2D model weights and showing improved accuracy as compared to separate 2D or 3D detectors. However, slice-based 2.5D detectors do not explicitly model inter-slice consistency for LN as a 3D object, requiring heuristic post-merging steps to generate final 3D LN instances, which can involve tuning a set of parameters for each dataset. In this work, we formulate 3D LN detection as a tracking task and propose LN-Tracker, a novel LN tracking transformer, for joint end-to-end detection and 3D instance association. Built upon DETR-based detector, LN-Tracker decouples transformer decoder's query into the track and detection groups, where the track query autoregressively follows previously tracked LN instances along the z-axis of a CT scan. We design a new transformer decoder with masked attention module to align track query's content to the context of current slice, meanwhile preserving detection query's high accuracy in current slice. An inter-slice similarity loss is introduced to encourage cohesive LN association between slices. Extensive evaluation on four lymph node datasets shows LN-Tracker's superior performance, with at least 2.7% gain in average sensitivity when compared to other top 3D/2.5D detectors. Further validation on public lung nodule and prostate tumor detection tasks confirms the generalizability of LN-Tracker as it achieves top performance on both tasks. Datasets will be released upon acceptance.
From Pixels to Gigapixels: Bridging Local Inductive Bias and Long-Range Dependencies with Pixel-Mamba
Dec 21, 2024



Abstract:Histopathology plays a critical role in medical diagnostics, with whole slide images (WSIs) offering valuable insights that directly influence clinical decision-making. However, the large size and complexity of WSIs may pose significant challenges for deep learning models, in both computational efficiency and effective representation learning. In this work, we introduce Pixel-Mamba, a novel deep learning architecture designed to efficiently handle gigapixel WSIs. Pixel-Mamba leverages the Mamba module, a state-space model (SSM) with linear memory complexity, and incorporates local inductive biases through progressively expanding tokens, akin to convolutional neural networks. This enables Pixel-Mamba to hierarchically combine both local and global information while efficiently addressing computational challenges. Remarkably, Pixel-Mamba achieves or even surpasses the quantitative performance of state-of-the-art (SOTA) foundation models that were pretrained on millions of WSIs or WSI-text pairs, in a range of tumor staging and survival analysis tasks, {\bf even without requiring any pathology-specific pretraining}. Extensive experiments demonstrate the efficacy of Pixel-Mamba as a powerful and efficient framework for end-to-end WSI analysis.
From Histopathology Images to Cell Clouds: Learning Slide Representations with Hierarchical Cell Transformer
Dec 21, 2024Abstract:It is clinically crucial and potentially very beneficial to be able to analyze and model directly the spatial distributions of cells in histopathology whole slide images (WSI). However, most existing WSI datasets lack cell-level annotations, owing to the extremely high cost over giga-pixel images. Thus, it remains an open question whether deep learning models can directly and effectively analyze WSIs from the semantic aspect of cell distributions. In this work, we construct a large-scale WSI dataset with more than 5 billion cell-level annotations, termed WSI-Cell5B, and a novel hierarchical Cell Cloud Transformer (CCFormer) to tackle these challenges. WSI-Cell5B is based on 6,998 WSIs of 11 cancers from The Cancer Genome Atlas Program, and all WSIs are annotated per cell by coordinates and types. To the best of our knowledge, WSI-Cell5B is the first WSI-level large-scale dataset integrating cell-level annotations. On the other hand, CCFormer formulates the collection of cells in each WSI as a cell cloud and models cell spatial distribution. Specifically, Neighboring Information Embedding (NIE) is proposed to characterize the distribution of cells within the neighborhood of each cell, and a novel Hierarchical Spatial Perception (HSP) module is proposed to learn the spatial relationship among cells in a bottom-up manner. The clinical analysis indicates that WSI-Cell5B can be used to design clinical evaluation metrics based on counting cells that effectively assess the survival risk of patients. Extensive experiments on survival prediction and cancer staging show that learning from cell spatial distribution alone can already achieve state-of-the-art (SOTA) performance, i.e., CCFormer strongly outperforms other competing methods.
Leveraging Semantic Asymmetry for Precise Gross Tumor Volume Segmentation of Nasopharyngeal Carcinoma in Planning CT
Nov 27, 2024



Abstract:In the radiation therapy of nasopharyngeal carcinoma (NPC), clinicians typically delineate the gross tumor volume (GTV) using non-contrast planning computed tomography to ensure accurate radiation dose delivery. However, the low contrast between tumors and adjacent normal tissues necessitates that radiation oncologists manually delineate the tumors, often relying on diagnostic MRI for guidance. % In this study, we propose a novel approach to directly segment NPC gross tumors on non-contrast planning CT images, circumventing potential registration errors when aligning MRI or MRI-derived tumor masks to planning CT. To address the low contrast issues between tumors and adjacent normal structures in planning CT, we introduce a 3D Semantic Asymmetry Tumor segmentation (SATs) method. Specifically, we posit that a healthy nasopharyngeal region is characteristically bilaterally symmetric, whereas the emergence of nasopharyngeal carcinoma disrupts this symmetry. Then, we propose a Siamese contrastive learning segmentation framework that minimizes the voxel-wise distance between original and flipped areas without tumor and encourages a larger distance between original and flipped areas with tumor. Thus, our approach enhances the sensitivity of features to semantic asymmetries. % Extensive experiments demonstrate that the proposed SATs achieves the leading NPC GTV segmentation performance in both internal and external testing, \emph{e.g.}, with at least 2\% absolute Dice score improvement and 12\% average distance error reduction when compared to other state-of-the-art methods in the external testing.
Low-Rank Continual Pyramid Vision Transformer: Incrementally Segment Whole-Body Organs in CT with Light-Weighted Adaptation
Oct 07, 2024Abstract:Deep segmentation networks achieve high performance when trained on specific datasets. However, in clinical practice, it is often desirable that pretrained segmentation models can be dynamically extended to enable segmenting new organs without access to previous training datasets or without training from scratch. This would ensure a much more efficient model development and deployment paradigm accounting for the patient privacy and data storage issues. This clinically preferred process can be viewed as a continual semantic segmentation (CSS) problem. Previous CSS works would either experience catastrophic forgetting or lead to unaffordable memory costs as models expand. In this work, we propose a new continual whole-body organ segmentation model with light-weighted low-rank adaptation (LoRA). We first train and freeze a pyramid vision transformer (PVT) base segmentation model on the initial task, then continually add light-weighted trainable LoRA parameters to the frozen model for each new learning task. Through a holistically exploration of the architecture modification, we identify three most important layers (i.e., patch-embedding, multi-head attention and feed forward layers) that are critical in adapting to the new segmentation tasks, while retaining the majority of the pretrained parameters fixed. Our proposed model continually segments new organs without catastrophic forgetting and meanwhile maintaining a low parameter increasing rate. Continually trained and tested on four datasets covering different body parts of a total of 121 organs, results show that our model achieves high segmentation accuracy, closely reaching the PVT and nnUNet upper bounds, and significantly outperforms other regularization-based CSS methods. When comparing to the leading architecture-based CSS method, our model has a substantial lower parameter increasing rate while achieving comparable performance.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge