Yongyi Lu
CLIP-Driven Universal Model for Organ Segmentation and Tumor Detection
Jan 06, 2023



Abstract:An increasing number of public datasets have shown a marked clinical impact on assessing anatomical structures. However, each of the datasets is small, partially labeled, and rarely investigates severe tumor subjects. Moreover, current models are limited to segmenting specific organs/tumors, which can not be extended to novel domains and classes. To tackle these limitations, we introduce embedding learned from Contrastive Language-Image Pre-training (CLIP) to segmentation models, dubbed the CLIP-Driven Universal Model. The Universal Model can better segment 25 organs and 6 types of tumors by exploiting the semantic relationship between abdominal structures. The model is developed from an assembly of 14 datasets with 3,410 CT scans and evaluated on 6,162 external CT scans from 3 datasets. We achieve the state-of-the-art results on Beyond The Cranial Vault (BTCV). Compared with dataset-specific models, the Universal Model is computationally more efficient (6x faster), generalizes better to CT scans from varying sites, and shows stronger transfer learning performance on novel tasks. The design of CLIP embedding enables the Universal Model to be easily extended to new classes without catastrophically forgetting the previously learned classes.
Making Your First Choice: To Address Cold Start Problem in Vision Active Learning
Oct 05, 2022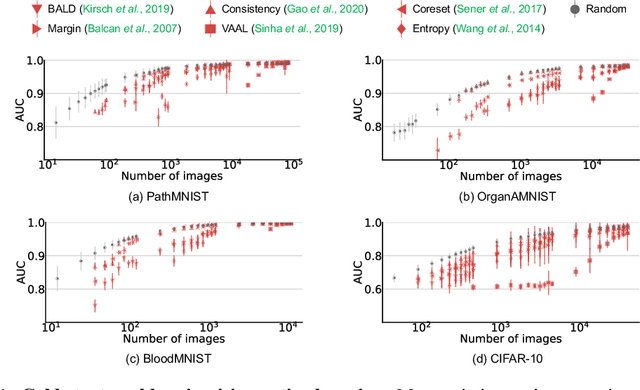
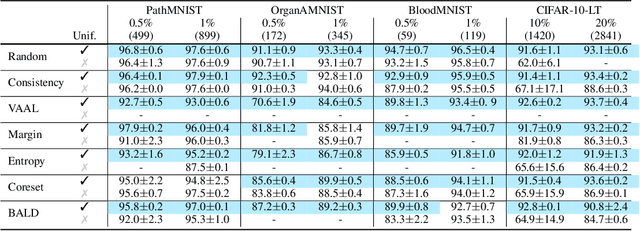
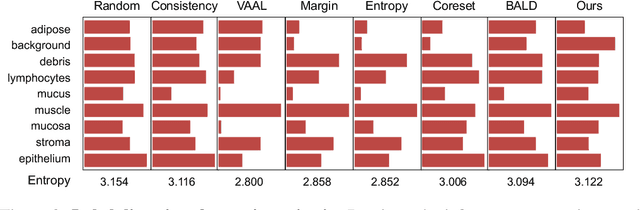
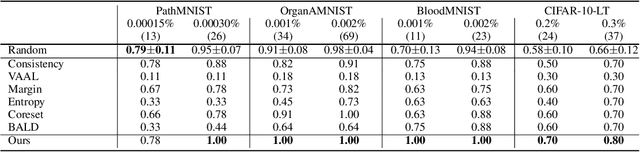
Abstract:Active learning promises to improve annotation efficiency by iteratively selecting the most important data to be annotated first. However, we uncover a striking contradiction to this promise: active learning fails to select data as efficiently as random selection at the first few choices. We identify this as the cold start problem in vision active learning, caused by a biased and outlier initial query. This paper seeks to address the cold start problem by exploiting the three advantages of contrastive learning: (1) no annotation is required; (2) label diversity is ensured by pseudo-labels to mitigate bias; (3) typical data is determined by contrastive features to reduce outliers. Experiments are conducted on CIFAR-10-LT and three medical imaging datasets (i.e. Colon Pathology, Abdominal CT, and Blood Cell Microscope). Our initial query not only significantly outperforms existing active querying strategies but also surpasses random selection by a large margin. We foresee our solution to the cold start problem as a simple yet strong baseline to choose the initial query for vision active learning. Code is available: https://github.com/c-liangyu/CSVAL
Unsupervised Domain Adaptation through Shape Modeling for Medical Image Segmentation
Jul 06, 2022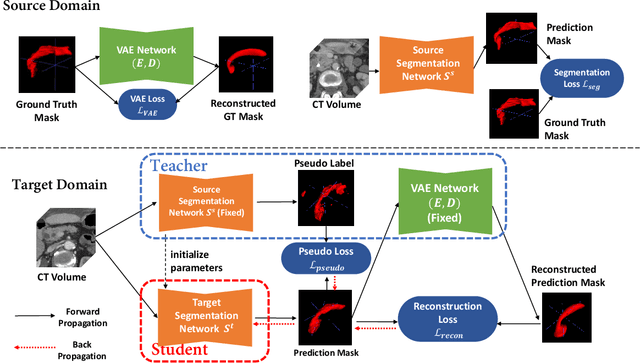
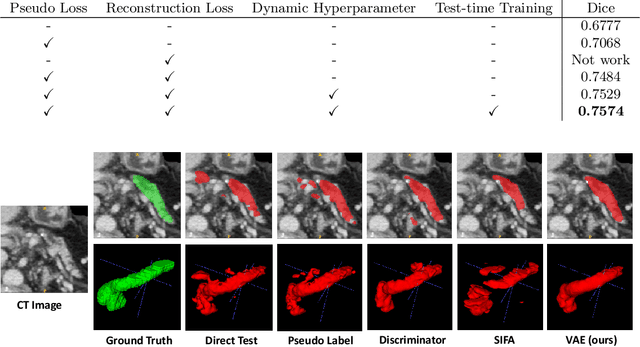
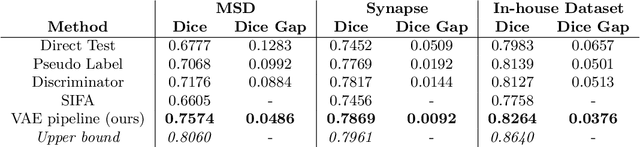
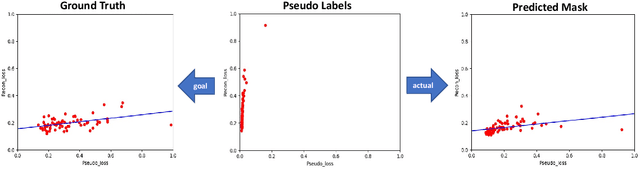
Abstract:Shape information is a strong and valuable prior in segmenting organs in medical images. However, most current deep learning based segmentation algorithms have not taken shape information into consideration, which can lead to bias towards texture. We aim at modeling shape explicitly and using it to help medical image segmentation. Previous methods proposed Variational Autoencoder (VAE) based models to learn the distribution of shape for a particular organ and used it to automatically evaluate the quality of a segmentation prediction by fitting it into the learned shape distribution. Based on which we aim at incorporating VAE into current segmentation pipelines. Specifically, we propose a new unsupervised domain adaptation pipeline based on a pseudo loss and a VAE reconstruction loss under a teacher-student learning paradigm. Both losses are optimized simultaneously and, in return, boost the segmentation task performance. Extensive experiments on three public Pancreas segmentation datasets as well as two in-house Pancreas segmentation datasets show consistent improvements with at least 2.8 points gain in the Dice score, demonstrating the effectiveness of our method in challenging unsupervised domain adaptation scenarios for medical image segmentation. We hope this work will advance shape analysis and geometric learning in medical imaging.
Semantic-Aware Representation Blending for Multi-Label Image Recognition with Partial Labels
May 26, 2022
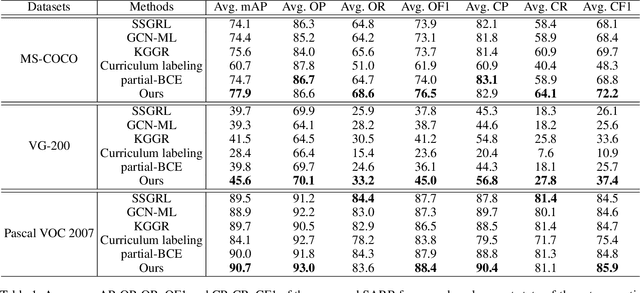
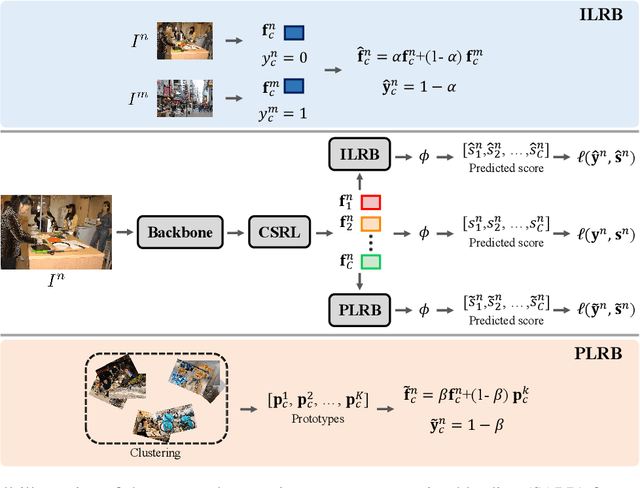
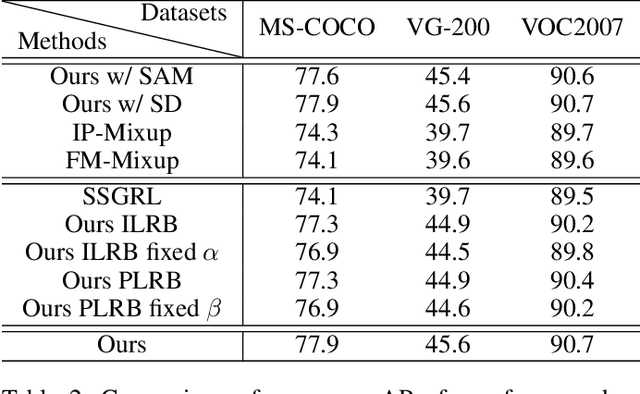
Abstract:Despite achieving impressive progress, current multi-label image recognition (MLR) algorithms heavily depend on large-scale datasets with complete labels, making collecting large-scale datasets extremely time-consuming and labor-intensive. Training the multi-label image recognition models with partial labels (MLR-PL) is an alternative way to address this issue, in which merely some labels are known while others are unknown for each image (see Figure 1). However, current MLP-PL algorithms mainly rely on the pre-trained image classification or similarity models to generate pseudo labels for the unknown labels. Thus, they depend on a certain amount of data annotations and inevitably suffer from obvious performance drops, especially when the known label proportion is low. To address this dilemma, we propose a unified semantic-aware representation blending (SARB) that consists of two crucial modules to blend multi-granularity category-specific semantic representation across different images to transfer information of known labels to complement unknown labels. Extensive experiments on the MS-COCO, Visual Genome, and Pascal VOC 2007 datasets show that the proposed SARB consistently outperforms current state-of-the-art algorithms on all known label proportion settings. Concretely, it obtain the average mAP improvement of 1.9%, 4.5%, 1.0% on the three benchmark datasets compared with the second-best algorithm.
Exploring Negatives in Contrastive Learning for Unpaired Image-to-Image Translation
Apr 23, 2022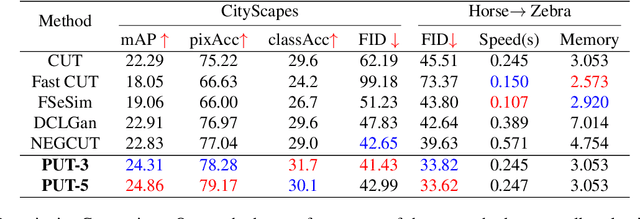
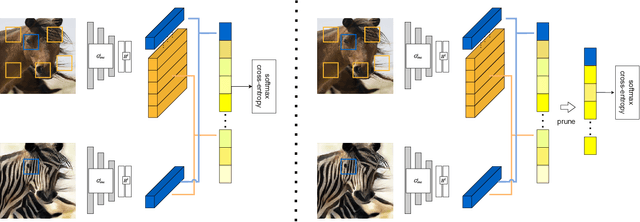
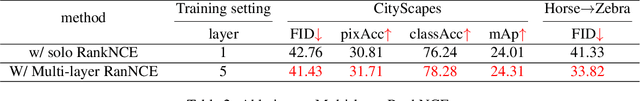
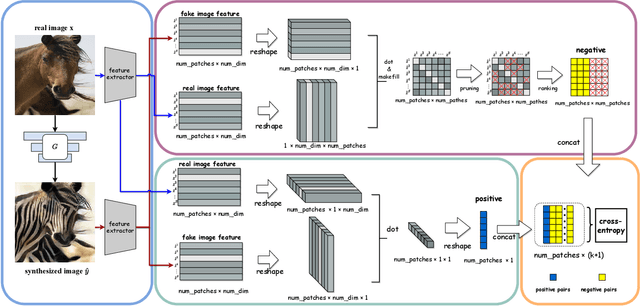
Abstract:Unpaired image-to-image translation aims to find a mapping between the source domain and the target domain. To alleviate the problem of the lack of supervised labels for the source images, cycle-consistency based methods have been proposed for image structure preservation by assuming a reversible relationship between unpaired images. However, this assumption only uses limited correspondence between image pairs. Recently, contrastive learning (CL) has been used to further investigate the image correspondence in unpaired image translation by using patch-based positive/negative learning. Patch-based contrastive routines obtain the positives by self-similarity computation and recognize the rest patches as negatives. This flexible learning paradigm obtains auxiliary contextualized information at a low cost. As the negatives own an impressive sample number, with curiosity, we make an investigation based on a question: are all negatives necessary for feature contrastive learning? Unlike previous CL approaches that use negatives as much as possible, in this paper, we study the negatives from an information-theoretic perspective and introduce a new negative Pruning technology for Unpaired image-to-image Translation (PUT) by sparsifying and ranking the patches. The proposed algorithm is efficient, flexible and enables the model to learn essential information between corresponding patches stably. By putting quality over quantity, only a few negative patches are required to achieve better results. Lastly, we validate the superiority, stability, and versatility of our model through comparative experiments.
MT-TransUNet: Mediating Multi-Task Tokens in Transformers for Skin Lesion Segmentation and Classification
Dec 03, 2021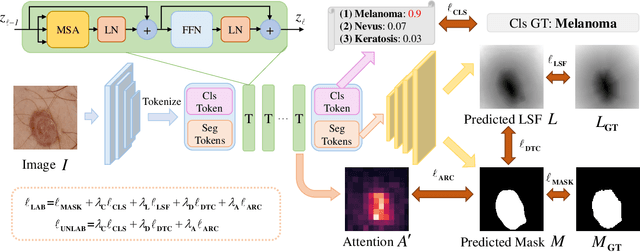
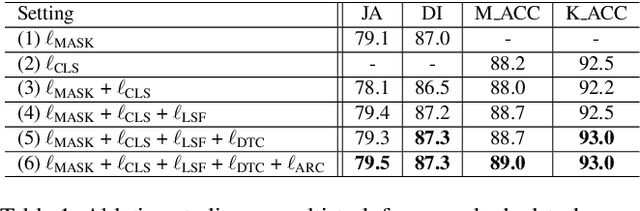
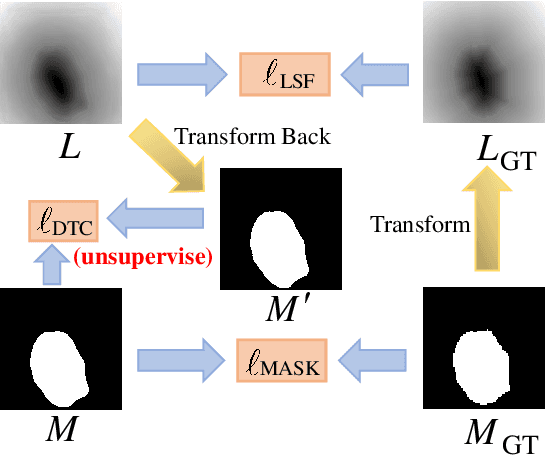
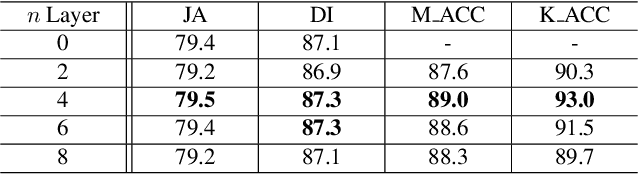
Abstract:Recent advances in automated skin cancer diagnosis have yielded performance on par with board-certified dermatologists. However, these approaches formulated skin cancer diagnosis as a simple classification task, dismissing the potential benefit from lesion segmentation. We argue that an accurate lesion segmentation can supplement the classification task with additive lesion information, such as asymmetry, border, intensity, and physical size; in turn, a faithful lesion classification can support the segmentation task with discriminant lesion features. To this end, this paper proposes a new multi-task framework, named MT-TransUNet, which is capable of segmenting and classifying skin lesions collaboratively by mediating multi-task tokens in Transformers. Furthermore, we have introduced dual-task and attended region consistency losses to take advantage of those images without pixel-level annotation, ensuring the model's robustness when it encounters the same image with an account of augmentation. Our MT-TransUNet exceeds the previous state of the art for lesion segmentation and classification tasks in ISIC-2017 and PH2; more importantly, it preserves compelling computational efficiency regarding model parameters (48M~vs.~130M) and inference speed (0.17s~vs.~2.02s per image). Code will be available at https://github.com/JingyeChen/MT-TransUNet.
In-painting Radiography Images for Unsupervised Anomaly Detection
Nov 30, 2021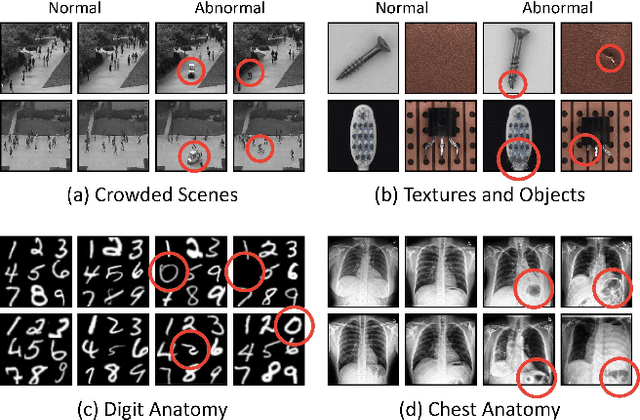
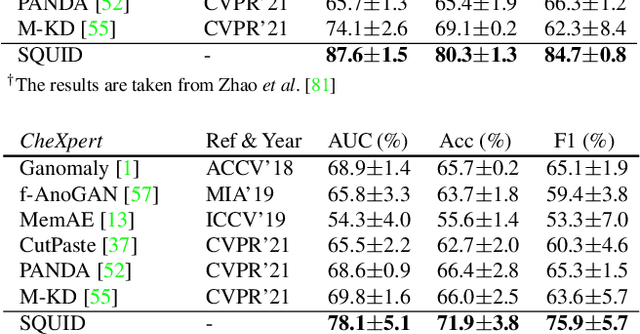
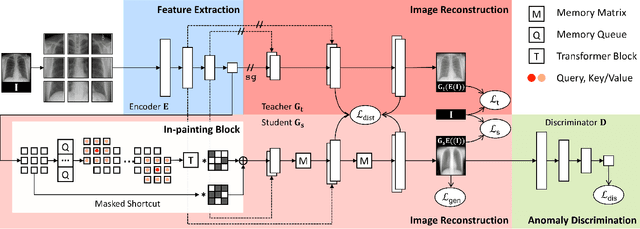
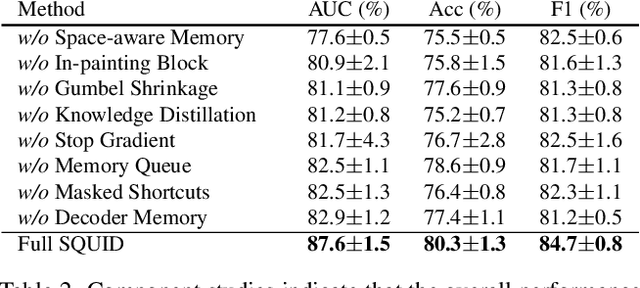
Abstract:We propose space-aware memory queues for in-painting and detecting anomalies from radiography images (abbreviated as SQUID). Radiography imaging protocols focus on particular body regions, therefore producing images of great similarity and yielding recurrent anatomical structures across patients. To exploit this structured information, our SQUID consists of a new Memory Queue and a novel in-painting block in the feature space. We show that SQUID can taxonomize the ingrained anatomical structures into recurrent patterns; and in the inference, SQUID can identify anomalies (unseen/modified patterns) in the image. SQUID surpasses the state of the art in unsupervised anomaly detection by over 5 points on two chest X-ray benchmark datasets. Additionally, we have created a new dataset (DigitAnatomy), which synthesizes the spatial correlation and consistent shape in chest anatomy. We hope DigitAnatomy can prompt the development, evaluation, and interpretability of anomaly detection methods, particularly for radiography imaging.
Data, Assemble: Leveraging Multiple Datasets with Heterogeneous and Partial Labels
Sep 25, 2021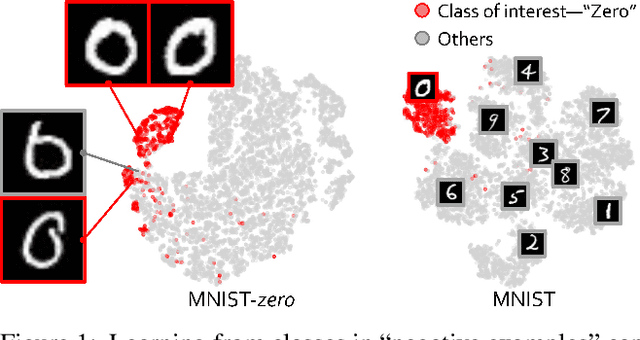
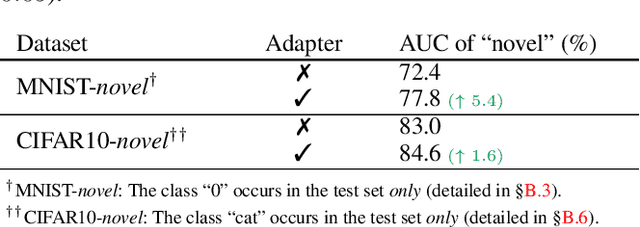
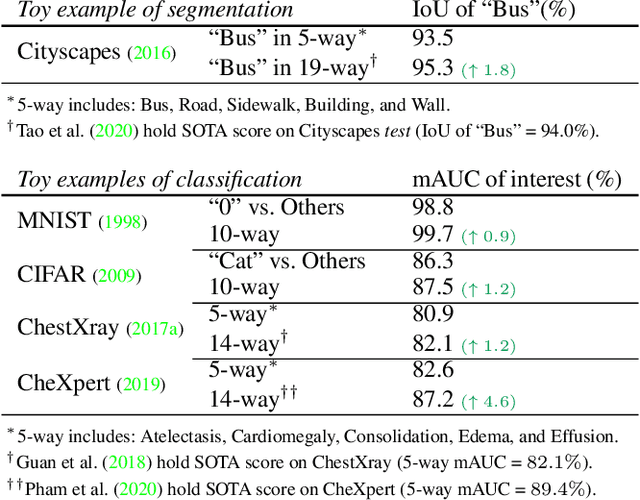
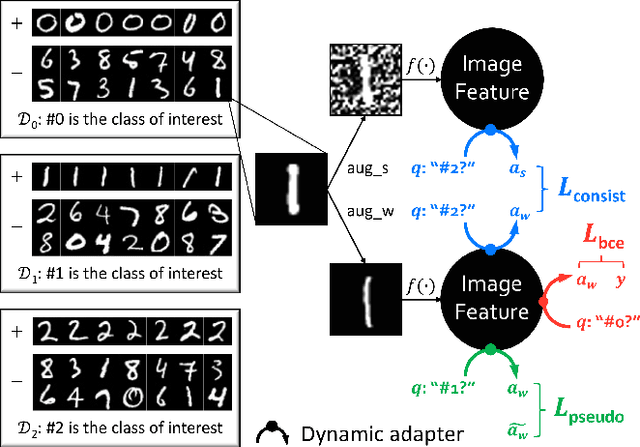

Abstract:The success of deep learning relies heavily on large datasets with extensive labels, but we often only have access to several small, heterogeneous datasets associated with partial labels, particularly in the field of medical imaging. When learning from multiple datasets, existing challenges include incomparable, heterogeneous, or even conflicting labeling protocols across datasets. In this paper, we propose a new initiative--"data, assemble"--which aims to unleash the full potential of partially labeled data and enormous unlabeled data from an assembly of datasets. To accommodate the supervised learning paradigm to partial labels, we introduce a dynamic adapter that encodes multiple visual tasks and aggregates image features in a question-and-answer manner. Furthermore, we employ pseudo-labeling and consistency constraints to harness images with missing labels and to mitigate the domain gap across datasets. From proof-of-concept studies on three natural imaging datasets and rigorous evaluations on two large-scale thorax X-ray benchmarks, we discover that learning from "negative examples" facilitates both classification and segmentation of classes of interest. This sheds new light on the computer-aided diagnosis of rare diseases and emerging pandemics, wherein "positive examples" are hard to collect, yet "negative examples" are relatively easier to assemble. As a result, besides exceeding the prior art in the NIH ChestXray benchmark, our model is particularly strong in identifying diseases of minority classes, yielding over 3-point improvement on average. Remarkably, when using existing partial labels, our model performance is on-par (p>0.05) with that using a fully curated dataset with exhaustive labels, eliminating the need for additional 40% annotation costs.
Glance-and-Gaze Vision Transformer
Jun 04, 2021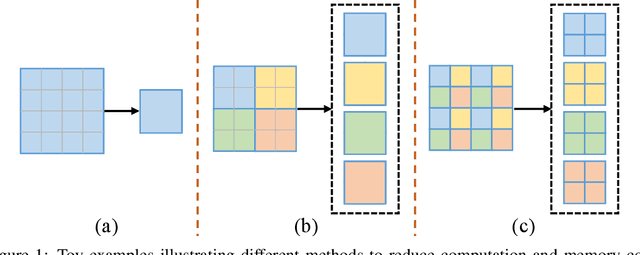
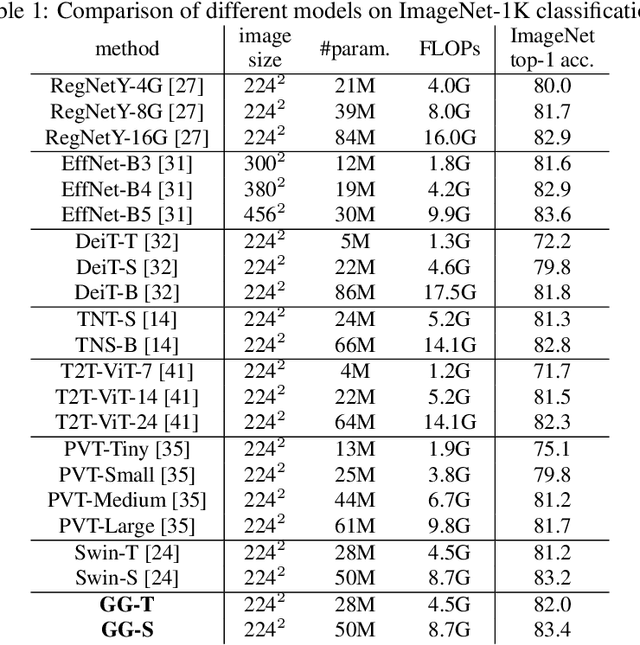
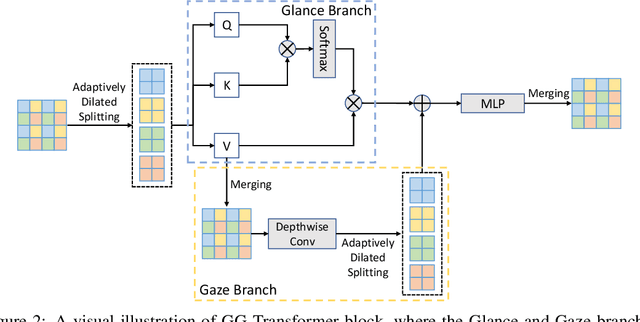
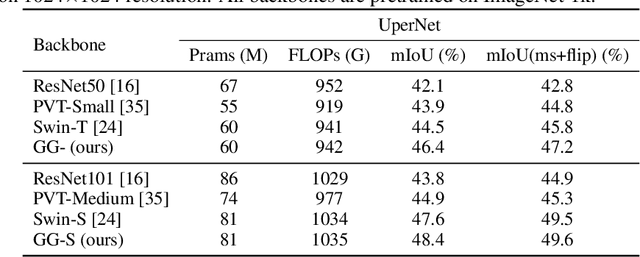
Abstract:Recently, there emerges a series of vision Transformers, which show superior performance with a more compact model size than conventional convolutional neural networks, thanks to the strong ability of Transformers to model long-range dependencies. However, the advantages of vision Transformers also come with a price: Self-attention, the core part of Transformer, has a quadratic complexity to the input sequence length. This leads to a dramatic increase of computation and memory cost with the increase of sequence length, thus introducing difficulties when applying Transformers to the vision tasks that require dense predictions based on high-resolution feature maps. In this paper, we propose a new vision Transformer, named Glance-and-Gaze Transformer (GG-Transformer), to address the aforementioned issues. It is motivated by the Glance and Gaze behavior of human beings when recognizing objects in natural scenes, with the ability to efficiently model both long-range dependencies and local context. In GG-Transformer, the Glance and Gaze behavior is realized by two parallel branches: The Glance branch is achieved by performing self-attention on the adaptively-dilated partitions of the input, which leads to a linear complexity while still enjoying a global receptive field; The Gaze branch is implemented by a simple depth-wise convolutional layer, which compensates local image context to the features obtained by the Glance mechanism. We empirically demonstrate our method achieves consistently superior performance over previous state-of-the-art Transformers on various vision tasks and benchmarks. The codes and models will be made available at https://github.com/yucornetto/GG-Transformer.
TransUNet: Transformers Make Strong Encoders for Medical Image Segmentation
Feb 08, 2021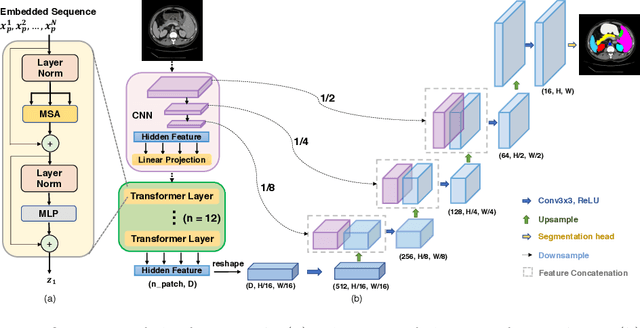
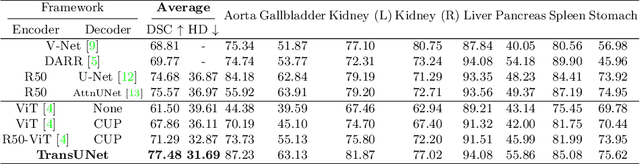
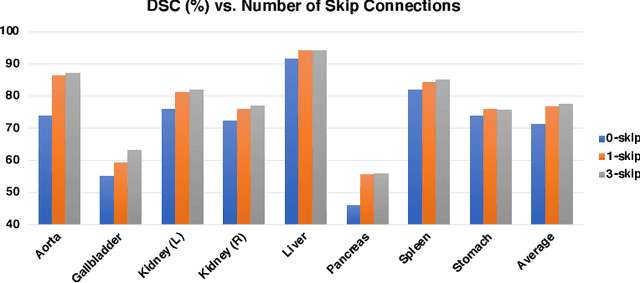
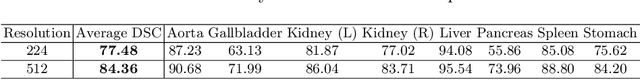
Abstract:Medical image segmentation is an essential prerequisite for developing healthcare systems, especially for disease diagnosis and treatment planning. On various medical image segmentation tasks, the u-shaped architecture, also known as U-Net, has become the de-facto standard and achieved tremendous success. However, due to the intrinsic locality of convolution operations, U-Net generally demonstrates limitations in explicitly modeling long-range dependency. Transformers, designed for sequence-to-sequence prediction, have emerged as alternative architectures with innate global self-attention mechanisms, but can result in limited localization abilities due to insufficient low-level details. In this paper, we propose TransUNet, which merits both Transformers and U-Net, as a strong alternative for medical image segmentation. On one hand, the Transformer encodes tokenized image patches from a convolution neural network (CNN) feature map as the input sequence for extracting global contexts. On the other hand, the decoder upsamples the encoded features which are then combined with the high-resolution CNN feature maps to enable precise localization. We argue that Transformers can serve as strong encoders for medical image segmentation tasks, with the combination of U-Net to enhance finer details by recovering localized spatial information. TransUNet achieves superior performances to various competing methods on different medical applications including multi-organ segmentation and cardiac segmentation. Code and models are available at https://github.com/Beckschen/TransUNet.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge