Wei Fan
HGAN: Hierarchical Graph Alignment Network for Image-Text Retrieval
Dec 16, 2022Abstract:Image-text retrieval (ITR) is a challenging task in the field of multimodal information processing due to the semantic gap between different modalities. In recent years, researchers have made great progress in exploring the accurate alignment between image and text. However, existing works mainly focus on the fine-grained alignment between image regions and sentence fragments, which ignores the guiding significance of context background information. Actually, integrating the local fine-grained information and global context background information can provide more semantic clues for retrieval. In this paper, we propose a novel Hierarchical Graph Alignment Network (HGAN) for image-text retrieval. First, to capture the comprehensive multimodal features, we construct the feature graphs for the image and text modality respectively. Then, a multi-granularity shared space is established with a designed Multi-granularity Feature Aggregation and Rearrangement (MFAR) module, which enhances the semantic corresponding relations between the local and global information, and obtains more accurate feature representations for the image and text modalities. Finally, the ultimate image and text features are further refined through three-level similarity functions to achieve the hierarchical alignment. To justify the proposed model, we perform extensive experiments on MS-COCO and Flickr30K datasets. Experimental results show that the proposed HGAN outperforms the state-of-the-art methods on both datasets, which demonstrates the effectiveness and superiority of our model.
Background-Mixed Augmentation for Weakly Supervised Change Detection
Dec 01, 2022Abstract:Change detection (CD) is to decouple object changes (i.e., object missing or appearing) from background changes (i.e., environment variations) like light and season variations in two images captured in the same scene over a long time span, presenting critical applications in disaster management, urban development, etc. In particular, the endless patterns of background changes require detectors to have a high generalization against unseen environment variations, making this task significantly challenging. Recent deep learning-based methods develop novel network architectures or optimization strategies with paired-training examples, which do not handle the generalization issue explicitly and require huge manual pixel-level annotation efforts. In this work, for the first attempt in the CD community, we study the generalization issue of CD from the perspective of data augmentation and develop a novel weakly supervised training algorithm that only needs image-level labels. Different from general augmentation techniques for classification, we propose the background-mixed augmentation that is specifically designed for change detection by augmenting examples under the guidance of a set of background-changing images and letting deep CD models see diverse environment variations. Moreover, we propose the augmented & real data consistency loss that encourages the generalization increase significantly. Our method as a general framework can enhance a wide range of existing deep learning-based detectors. We conduct extensive experiments in two public datasets and enhance four state-of-the-art methods, demonstrating the advantages of our method. We release the code at https://github.com/tsingqguo/bgmix.
DiMBERT: Learning Vision-Language Grounded Representations with Disentangled Multimodal-Attention
Oct 28, 2022



Abstract:Vision-and-language (V-L) tasks require the system to understand both vision content and natural language, thus learning fine-grained joint representations of vision and language (a.k.a. V-L representations) is of paramount importance. Recently, various pre-trained V-L models are proposed to learn V-L representations and achieve improved results in many tasks. However, the mainstream models process both vision and language inputs with the same set of attention matrices. As a result, the generated V-L representations are entangled in one common latent space. To tackle this problem, we propose DiMBERT (short for Disentangled Multimodal-Attention BERT), which is a novel framework that applies separated attention spaces for vision and language, and the representations of multi-modalities can thus be disentangled explicitly. To enhance the correlation between vision and language in disentangled spaces, we introduce the visual concepts to DiMBERT which represent visual information in textual format. In this manner, visual concepts help to bridge the gap between the two modalities. We pre-train DiMBERT on a large amount of image-sentence pairs on two tasks: bidirectional language modeling and sequence-to-sequence language modeling. After pre-train, DiMBERT is further fine-tuned for the downstream tasks. Experiments show that DiMBERT sets new state-of-the-art performance on three tasks (over four datasets), including both generation tasks (image captioning and visual storytelling) and classification tasks (referring expressions). The proposed DiM (short for Disentangled Multimodal-Attention) module can be easily incorporated into existing pre-trained V-L models to boost their performance, up to a 5% increase on the representative task. Finally, we conduct a systematic analysis and demonstrate the effectiveness of our DiM and the introduced visual concepts.
Graph Soft-Contrastive Learning via Neighborhood Ranking
Sep 28, 2022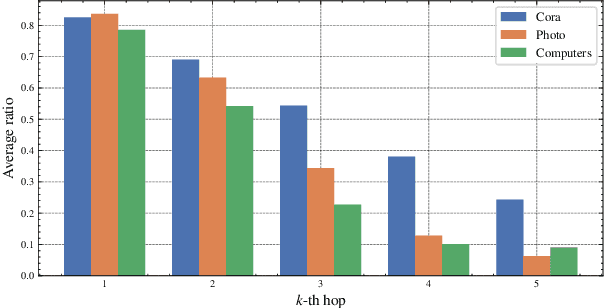
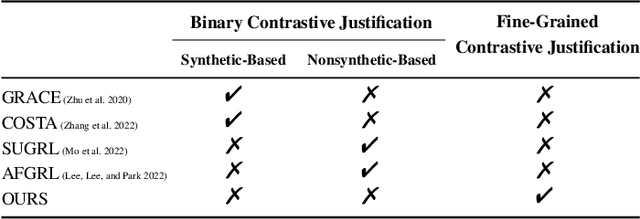
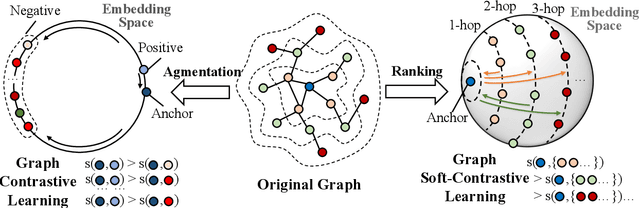
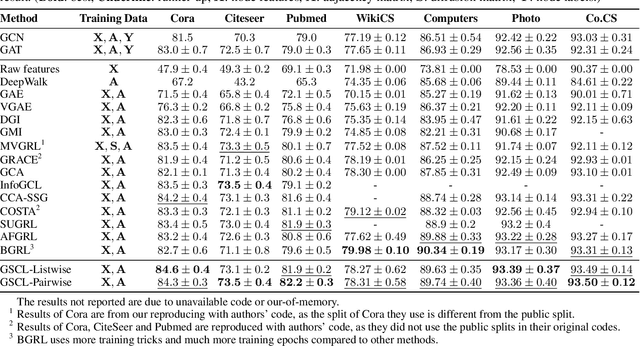
Abstract:Graph contrastive learning (GCL) has been an emerging solution for graph self-supervised learning. The core principle of GCL is to reduce the distance between samples in the positive view, but increase the distance between samples in the negative view. While achieving promising performances, current GCL methods still suffer from two limitations: (1) uncontrollable validity of augmentation, that graph perturbation may produce invalid views against semantics and feature-topology correspondence of graph data; and (2) unreliable binary contrastive justification, that the positiveness and negativeness of the constructed views are difficult to be determined for non-euclidean graph data. To tackle the above limitations, we propose a new contrastive learning paradigm for graphs, namely Graph Soft-Contrastive Learning (GSCL), that conducts contrastive learning in a finer-granularity via ranking neighborhoods without any augmentations and binary contrastive justification. GSCL is built upon the fundamental assumption of graph proximity that connected neighbors are more similar than far-distant nodes. Specifically, we develop pair-wise and list-wise Gated Ranking infoNCE Loss functions to preserve the relative ranking relationship in the neighborhood. Moreover, as the neighborhood size exponentially expands with more hops considered, we propose neighborhood sampling strategies to improve learning efficiency. The extensive experimental results show that our proposed GSCL can consistently achieve state-of-the-art performances on various public datasets with comparable practical complexity to GCL.
A Knowledge Distillation-Based Backdoor Attack in Federated Learning
Aug 12, 2022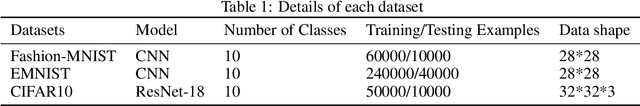
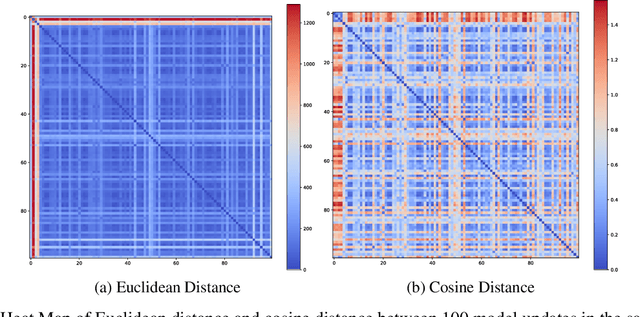
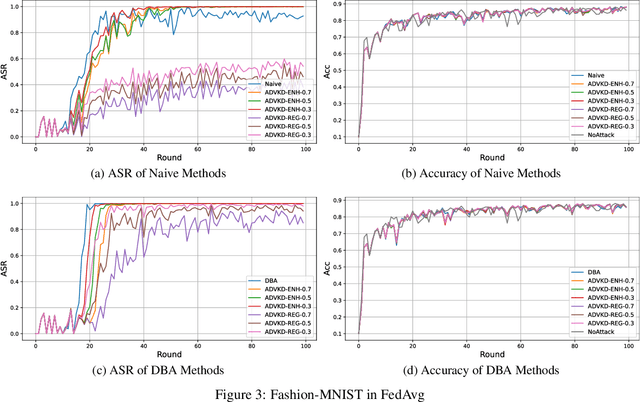
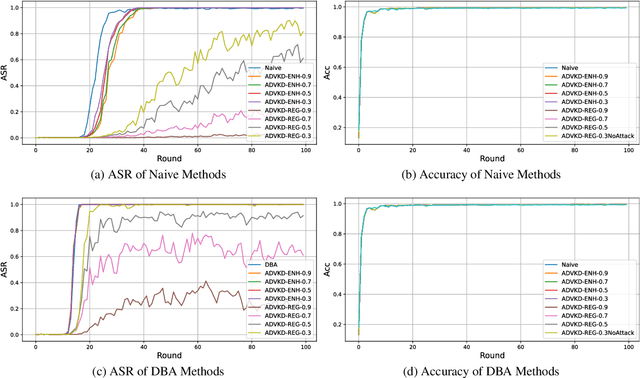
Abstract:Federated Learning (FL) is a novel framework of decentralized machine learning. Due to the decentralized feature of FL, it is vulnerable to adversarial attacks in the training procedure, e.g. , backdoor attacks. A backdoor attack aims to inject a backdoor into the machine learning model such that the model will make arbitrarily incorrect behavior on the test sample with some specific backdoor trigger. Even though a range of backdoor attack methods of FL has been introduced, there are also methods defending against them. Many of the defending methods utilize the abnormal characteristics of the models with backdoor or the difference between the models with backdoor and the regular models. To bypass these defenses, we need to reduce the difference and the abnormal characteristics. We find a source of such abnormality is that backdoor attack would directly flip the label of data when poisoning the data. However, current studies of the backdoor attack in FL are not mainly focus on reducing the difference between the models with backdoor and the regular models. In this paper, we propose Adversarial Knowledge Distillation(ADVKD), a method combine knowledge distillation with backdoor attack in FL. With knowledge distillation, we can reduce the abnormal characteristics in model result from the label flipping, thus the model can bypass the defenses. Compared to current methods, we show that ADVKD can not only reach a higher attack success rate, but also successfully bypass the defenses when other methods fails. To further explore the performance of ADVKD, we test how the parameters affect the performance of ADVKD under different scenarios. According to the experiment result, we summarize how to adjust the parameter for better performance under different scenarios. We also use several methods to visualize the effect of different attack and explain the effectiveness of ADVKD.
On the Feasibility of Out-of-Band Spatial Channel Information for Millimeter-Wave Beam Search
Aug 11, 2022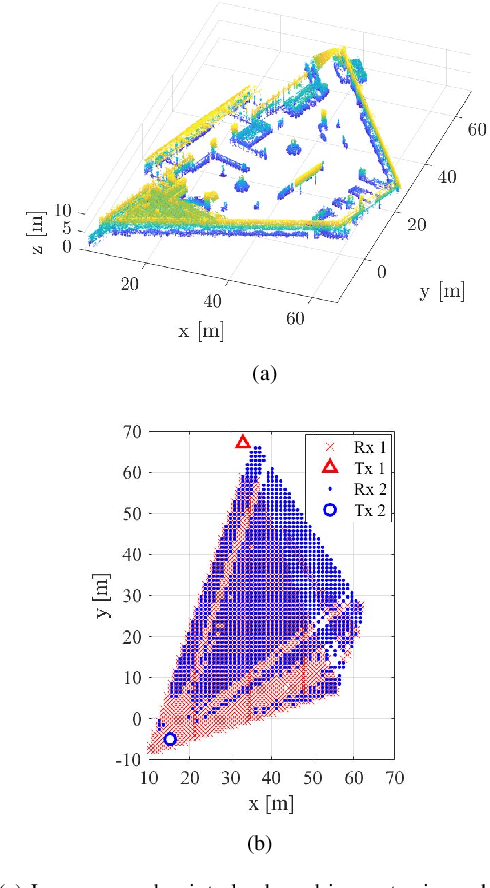
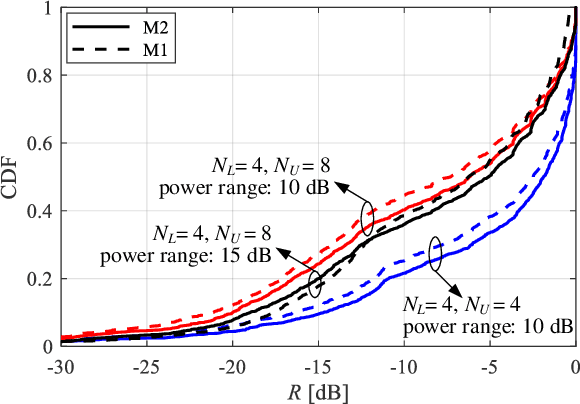
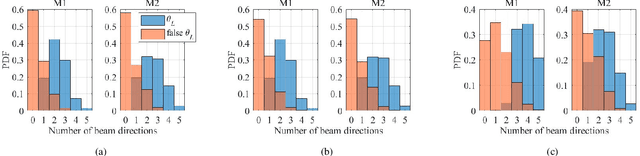
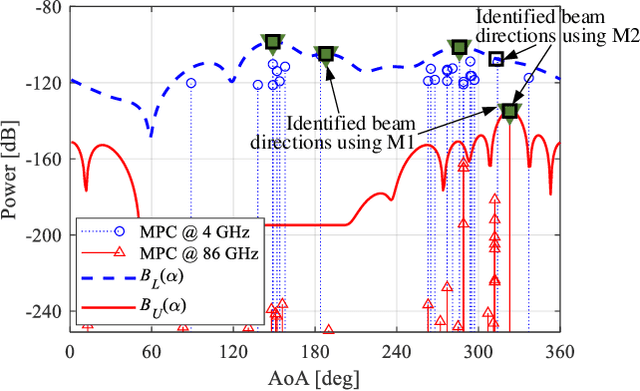
Abstract:The rollout of millimeter-wave (mmWave) cellular network enables us to realize the full potential of 5G/6G with vastly improved throughput and ultra-low latency. MmWave communication relies on highly directional transmission, which significantly increase the training overhead for fine beam alignment. The concept of using out-of-band spatial information to aid mmWave beam search is developed when multi-band systems operating in parallel. The feasibility of leveraging low-band channel information for coarse estimation of high-band beam directions strongly depends on the spatial congruence between two frequency bands. In this paper, we try to provide insights into the answers of two important questions. First, how similar is the power angular spectra (PAS) of radio channels between two well-separated frequency bands? Then, what is the impact of practical system configurations on spatial channel similarity? Specifically, the beam direction-based metric is proposed to measure the power loss and number of false directions if out-of-band spatial information is used instead of in-band information. This metric is more practical and useful than comparing normalized PAS directly. Point cloud ray-tracing and measurement results across multiple frequency bands and environments show that the degree of spatial similarity of beamformed channels is related to antenna beamwidth, frequency gap, and radio link conditions.
Omni-directional Pathloss Measurement Based on Virtual Antenna Array with Directional Antennas
Aug 07, 2022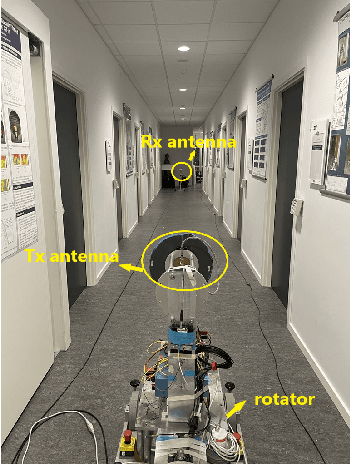
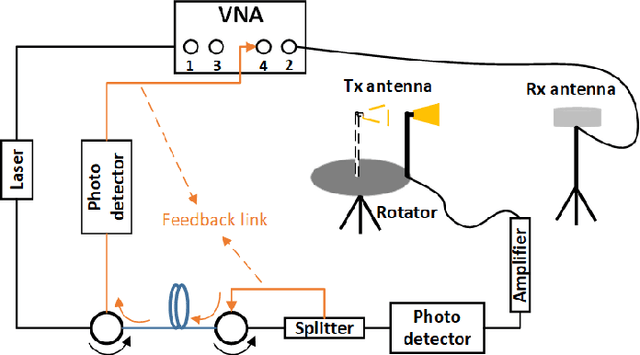

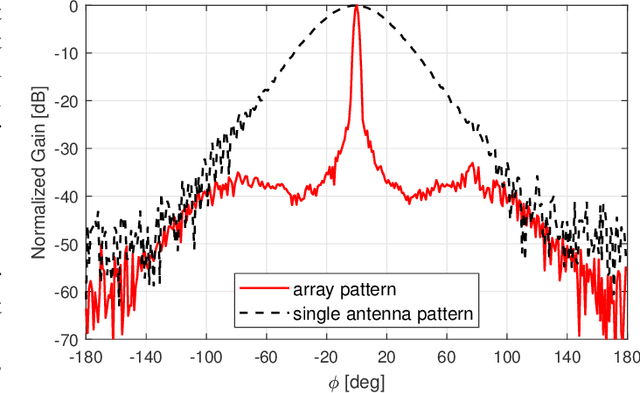
Abstract:Omni-directional pathloss, which refers to the pathloss when omni-directional antennas are used at the link ends, are essential for system design and evaluation. In the millimeter-wave (mm-Wave) and beyond bands, high gain directional antennas are widely used for channel measurements due to the significant signal attenuation. Conventional methods for omni-directional pathloss estimation are based on directional scanning sounding (DSS) system, i.e., a single directional antenna placed at the center of a rotator capturing signals from different rotation angles. The omni-directional pathloss is obtained by either summing up all the powers above the noise level or just summing up the powers of detected propagation paths. However, both methods are problematic with relatively wide main beams and high side-lobes provided by the directional antennas. In this letter, directional antenna based virtual antenna array (VAA) system is implemented for omni-directional pathloss estimation. The VAA scheme uses the same measurement system as the DSS, yet it offers high angular resolution (i.e. narrow main beam) and low side-lobes, which is essential for achieving accurate multipath detection in the power angular delay profiles (PADPs) and thereby obtaining accurate omni-directional pathloss. A measurement campaign was designed and conducted in an indoor corridor at 28-30 GHz to verify the effectiveness of the proposed method.
IDET: Iterative Difference-Enhanced Transformers for High-Quality Change Detection
Jul 15, 2022
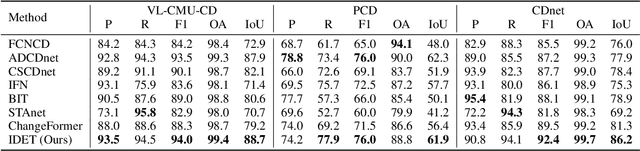
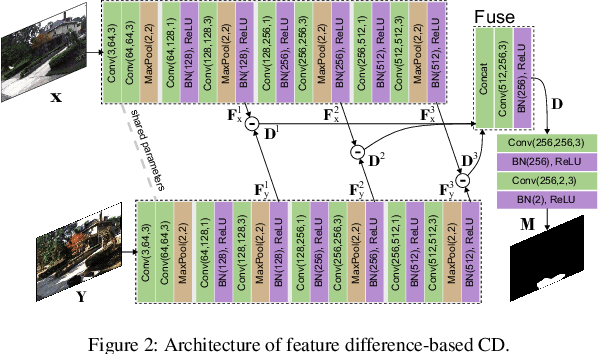
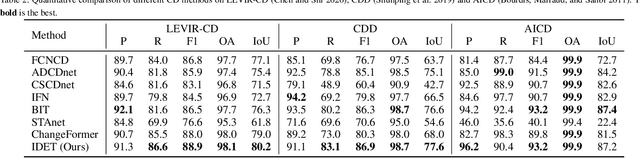
Abstract:Change detection (CD) aims to detect change regions within an image pair captured at different times, playing a significant role for diverse real-world applications. Nevertheless, most of existing works focus on designing advanced network architectures to map the feature difference to the final change map while ignoring the influence of the quality of the feature difference. In this paper, we study the CD from a new perspective, i.e., how to optimize the feature difference to highlight changes and suppress unchanged regions, and propose a novel module denoted as iterative difference-enhanced transformers (IDET). IDET contains three transformers: two transformers for extracting the long-range information of the two images and one transformer for enhancing the feature difference. In contrast to the previous transformers, the third transformer takes the outputs of the first two transformers to guide the enhancement of the feature difference iteratively. To achieve more effective refinement, we further propose the multi-scale IDET-based change detection that uses multi-scale representations of the images for multiple feature difference refinements and proposes a coarse-to-fine fusion strategy to combine all refinements. Our final CD method outperforms seven state-of-the-art methods on six large-scale datasets under diverse application scenarios, which demonstrates the importance of feature difference enhancements and the effectiveness of IDET.
Feature and Instance Joint Selection: A Reinforcement Learning Perspective
May 12, 2022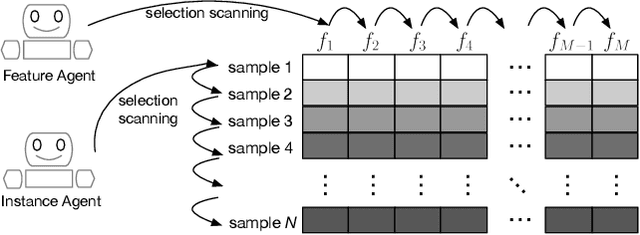
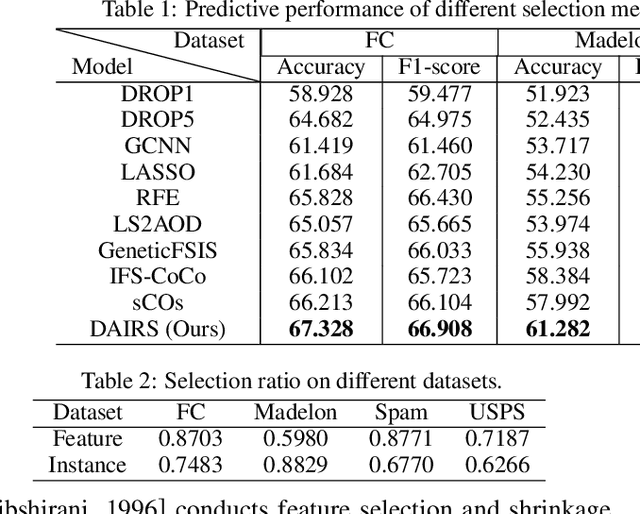
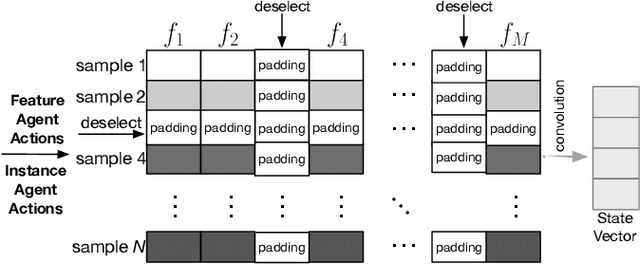
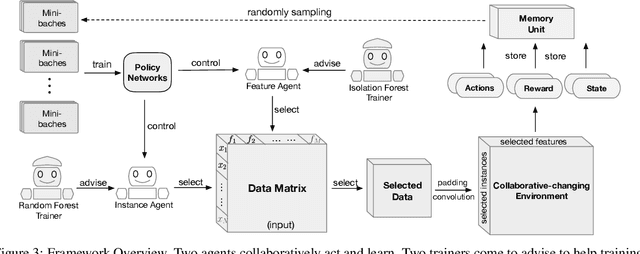
Abstract:Feature selection and instance selection are two important techniques of data processing. However, such selections have mostly been studied separately, while existing work towards the joint selection conducts feature/instance selection coarsely; thus neglecting the latent fine-grained interaction between feature space and instance space. To address this challenge, we propose a reinforcement learning solution to accomplish the joint selection task and simultaneously capture the interaction between the selection of each feature and each instance. In particular, a sequential-scanning mechanism is designed as action strategy of agents, and a collaborative-changing environment is used to enhance agent collaboration. In addition, an interactive paradigm introduces prior selection knowledge to help agents for more efficient exploration. Finally, extensive experiments on real-world datasets have demonstrated improved performances.
TransHER: Translating Knowledge Graph Embedding with Hyper-Ellipsoidal Restriction
Apr 27, 2022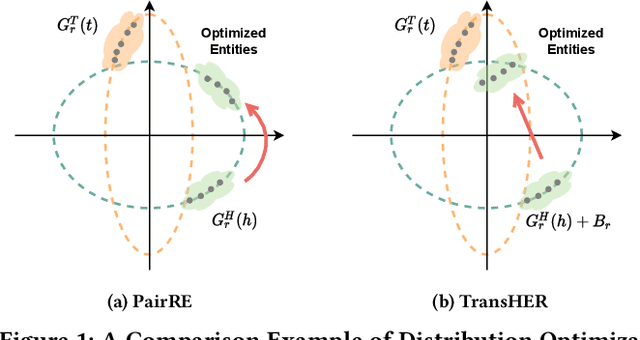
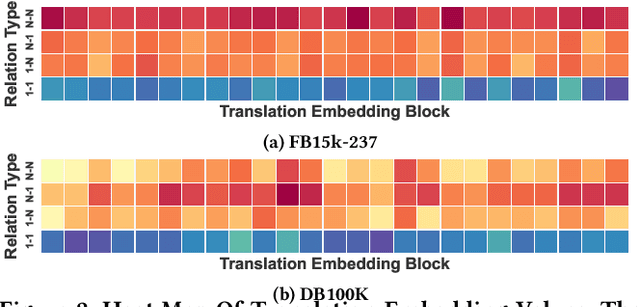
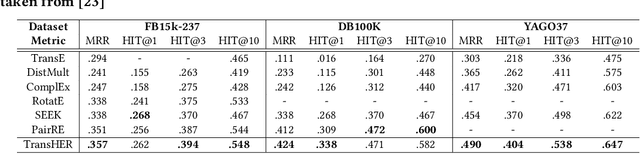
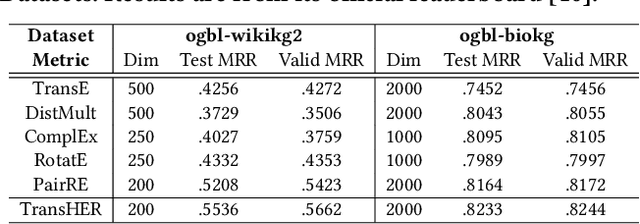

Abstract:Knowledge graph embedding methods are important for knowledge graph completion (link prediction) due to their robust performance and efficiency on large-magnitude datasets. One state-of-the-art method, PairRE, leverages two separate vectors for relations to model complex relations (i.e., 1-to-N, N-to-1, and N-to-N) in knowledge graphs. However, such a method strictly restricts entities on the hyper-ellipsoid surface and thus limits the optimization of entity distribution, which largely hinders the performance of knowledge graph completion. To address this problem, we propose a novel score function TransHER, which leverages relation-specific translations between head and tail entities restricted on separate hyper-ellipsoids. Specifically, given a triplet, our model first maps entities onto two separate hyper-ellipsoids and then conducts a relation-specific translation on one of them. The relation-specific translation provides TransHER with more direct guidance in optimization and the ability to learn semantic characteristics of entities with complex relations. Experimental results show that TransHER can achieve state-of-the-art performance and generalize to datasets in different domains and scales. All our code will be publicly available.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge