Tao Xu
College of Intelligent Systems Science and Engineering, Harbin Engineering University, Harbin, China
Intelligent Computing: The Latest Advances, Challenges and Future
Nov 21, 2022Abstract:Computing is a critical driving force in the development of human civilization. In recent years, we have witnessed the emergence of intelligent computing, a new computing paradigm that is reshaping traditional computing and promoting digital revolution in the era of big data, artificial intelligence and internet-of-things with new computing theories, architectures, methods, systems, and applications. Intelligent computing has greatly broadened the scope of computing, extending it from traditional computing on data to increasingly diverse computing paradigms such as perceptual intelligence, cognitive intelligence, autonomous intelligence, and human-computer fusion intelligence. Intelligence and computing have undergone paths of different evolution and development for a long time but have become increasingly intertwined in recent years: intelligent computing is not only intelligence-oriented but also intelligence-driven. Such cross-fertilization has prompted the emergence and rapid advancement of intelligent computing. Intelligent computing is still in its infancy and an abundance of innovations in the theories, systems, and applications of intelligent computing are expected to occur soon. We present the first comprehensive survey of literature on intelligent computing, covering its theory fundamentals, the technological fusion of intelligence and computing, important applications, challenges, and future perspectives. We believe that this survey is highly timely and will provide a comprehensive reference and cast valuable insights into intelligent computing for academic and industrial researchers and practitioners.
3D-Aware Encoding for Style-based Neural Radiance Fields
Nov 12, 2022



Abstract:We tackle the task of NeRF inversion for style-based neural radiance fields, (e.g., StyleNeRF). In the task, we aim to learn an inversion function to project an input image to the latent space of a NeRF generator and then synthesize novel views of the original image based on the latent code. Compared with GAN inversion for 2D generative models, NeRF inversion not only needs to 1) preserve the identity of the input image, but also 2) ensure 3D consistency in generated novel views. This requires the latent code obtained from the single-view image to be invariant across multiple views. To address this new challenge, we propose a two-stage encoder for style-based NeRF inversion. In the first stage, we introduce a base encoder that converts the input image to a latent code. To ensure the latent code is view-invariant and is able to synthesize 3D consistent novel view images, we utilize identity contrastive learning to train the base encoder. Second, to better preserve the identity of the input image, we introduce a refining encoder to refine the latent code and add finer details to the output image. Importantly note that the novelty of this model lies in the design of its first-stage encoder which produces the closest latent code lying on the latent manifold and thus the refinement in the second stage would be close to the NeRF manifold. Through extensive experiments, we demonstrate that our proposed two-stage encoder qualitatively and quantitatively exhibits superiority over the existing encoders for inversion in both image reconstruction and novel-view rendering.
Exploiting Dynamic and Fine-grained Semantic Scope for Extreme Multi-label Text Classification
May 24, 2022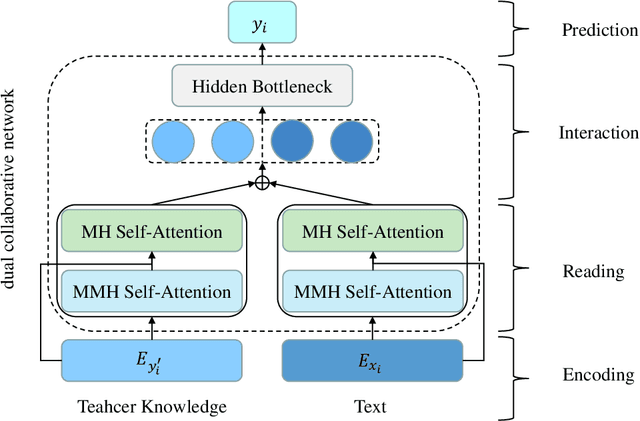

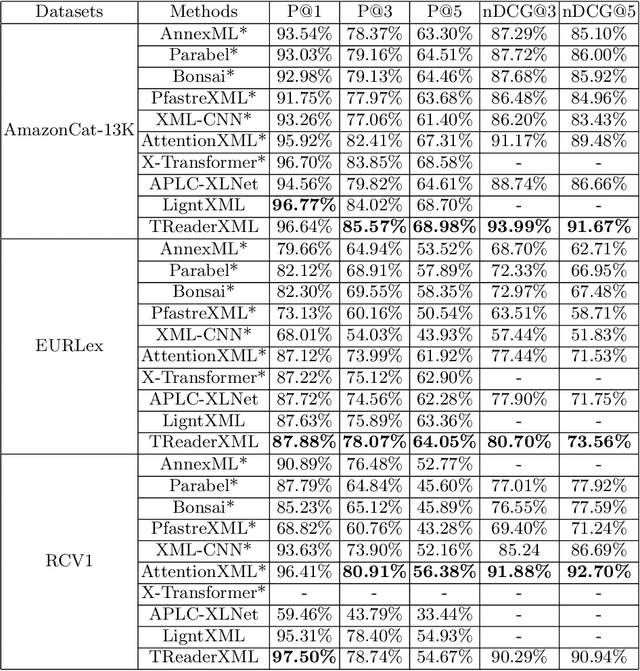
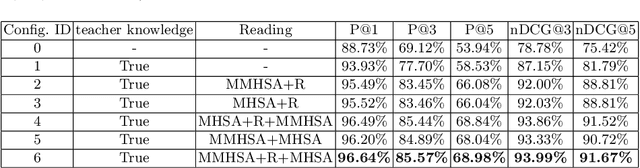
Abstract:Extreme multi-label text classification (XMTC) refers to the problem of tagging a given text with the most relevant subset of labels from a large label set. A majority of labels only have a few training instances due to large label dimensionality in XMTC. To solve this data sparsity issue, most existing XMTC methods take advantage of fixed label clusters obtained in early stage to balance performance on tail labels and head labels. However, such label clusters provide static and coarse-grained semantic scope for every text, which ignores distinct characteristics of different texts and has difficulties modelling accurate semantics scope for texts with tail labels. In this paper, we propose a novel framework TReaderXML for XMTC, which adopts dynamic and fine-grained semantic scope from teacher knowledge for individual text to optimize text conditional prior category semantic ranges. TReaderXML dynamically obtains teacher knowledge for each text by similar texts and hierarchical label information in training sets to release the ability of distinctly fine-grained label-oriented semantic scope. Then, TReaderXML benefits from a novel dual cooperative network that firstly learns features of a text and its corresponding label-oriented semantic scope by parallel Encoding Module and Reading Module, secondly embeds two parts by Interaction Module to regularize the text's representation by dynamic and fine-grained label-oriented semantic scope, and finally find target labels by Prediction Module. Experimental results on three XMTC benchmark datasets show that our method achieves new state-of-the-art results and especially performs well for severely imbalanced and sparse datasets.
Efficient conditioned face animation using frontally-viewed embedding
Mar 16, 2022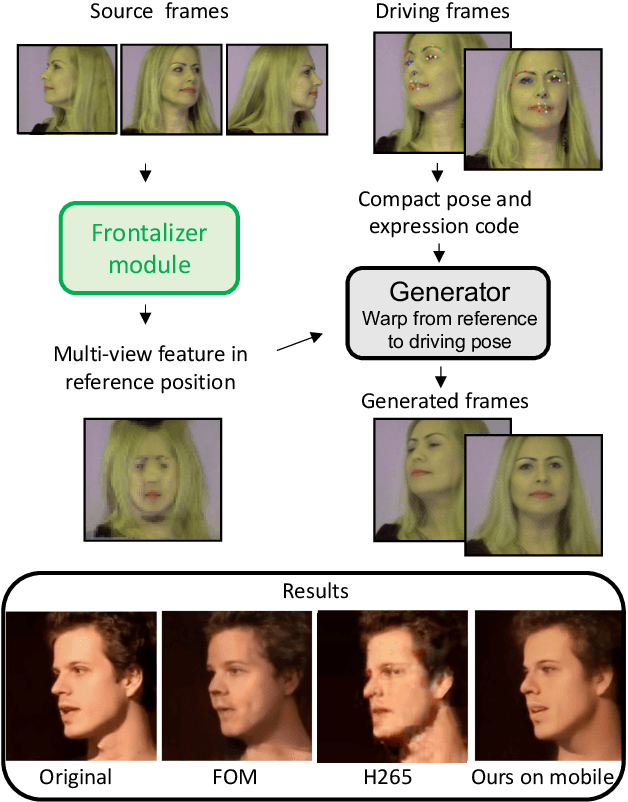
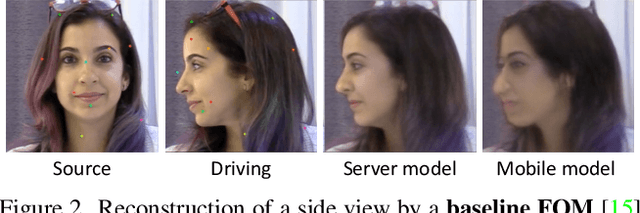


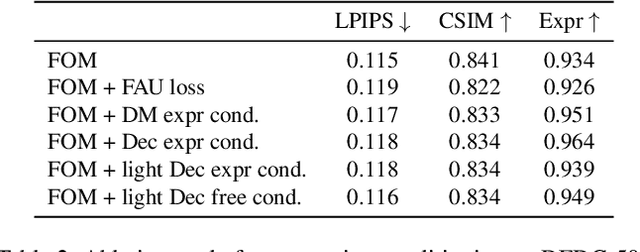
Abstract:As the quality of few shot facial animation from landmarks increases, new applications become possible, such as ultra low bandwidth video chat compression with a high degree of realism. However, there are some important challenges to tackle in order to improve the experience in real world conditions. In particular, the current approaches fail to represent profile views without distortions, while running in a low compute regime. We focus on this key problem by introducing a multi-frames embedding dubbed Frontalizer to improve profile views rendering. In addition to this core improvement, we explore the learning of a latent code conditioning generations along with landmarks to better convey facial expressions. Our dense models achieves 22% of improvement in perceptual quality and 73% reduction of landmark error over the first order model baseline on a subset of DFDC videos containing head movements. Declined with mobile architectures, our models outperform the previous state-of-the-art (improving perceptual quality by more than 16% and reducing landmark error by more than 47% on two datasets) while running on real time on iPhone 8 with very low bandwidth requirements.
DAGAM: A Domain Adversarial Graph Attention Model for Subject Independent EEG-Based Emotion Recognition
Feb 27, 2022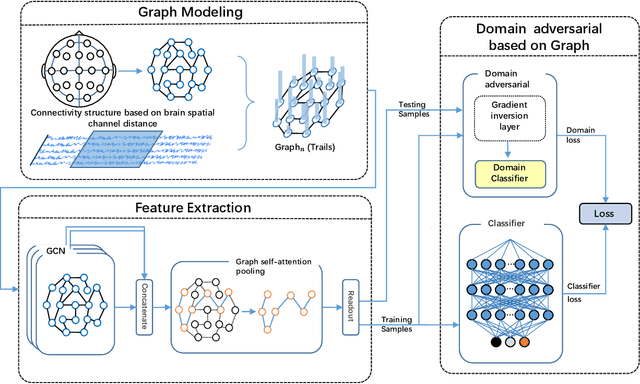
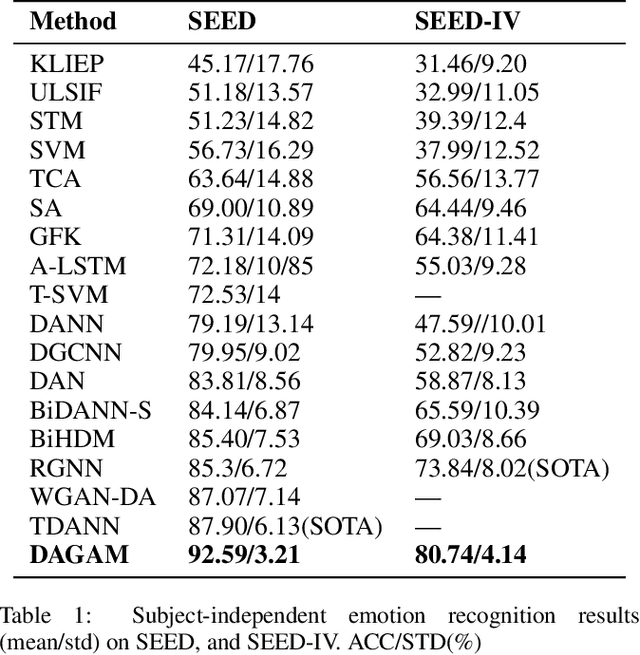
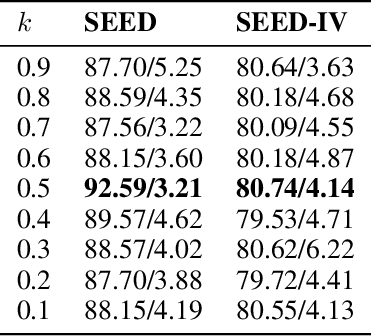
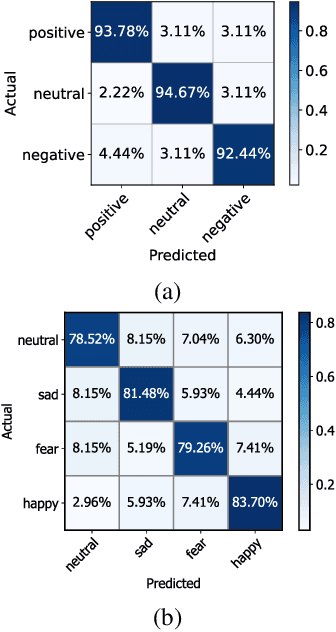
Abstract:One of the most significant challenges of EEG-based emotion recognition is the cross-subject EEG variations, leading to poor performance and generalizability. This paper proposes a novel EEG-based emotion recognition model called the domain adversarial graph attention model (DAGAM). The basic idea is to generate a graph to model multichannel EEG signals using biological topology. Graph theory can topologically describe and analyze relationships and mutual dependency between channels of EEG. Then, unlike other graph convolutional networks, self-attention pooling is applied to benefit salient EEG feature extraction from the graph, which effectively improves the performance. Finally, after graph pooling, the domain adversarial based on the graph is employed to identify and handle EEG variation across subjects, efficiently reaching good generalizability. We conduct extensive evaluations on two benchmark datasets (SEED and SEED IV) and obtain state-of-the-art results in subject-independent emotion recognition. Our model boosts the SEED accuracy to 92.59% (4.69% improvement) with the lowest standard deviation of 3.21% (2.92% decrements) and SEED IV accuracy to 80.74% (6.90% improvement) with the lowest standard deviation of 4.14% (3.88% decrements) respectively.
Text and Code Embeddings by Contrastive Pre-Training
Jan 24, 2022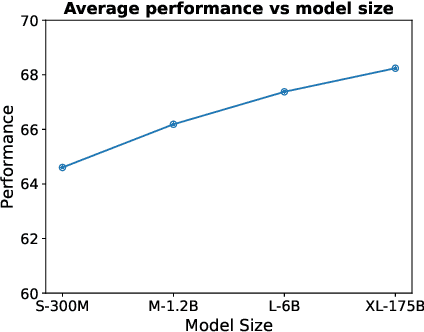
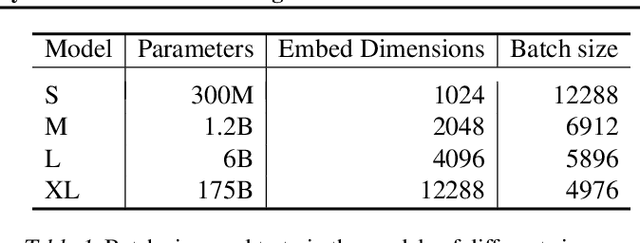
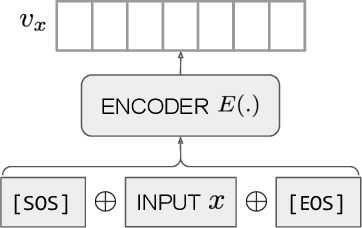
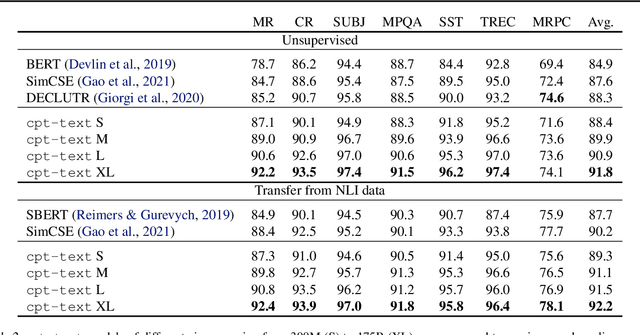
Abstract:Text embeddings are useful features in many applications such as semantic search and computing text similarity. Previous work typically trains models customized for different use cases, varying in dataset choice, training objective and model architecture. In this work, we show that contrastive pre-training on unsupervised data at scale leads to high quality vector representations of text and code. The same unsupervised text embeddings that achieve new state-of-the-art results in linear-probe classification also display impressive semantic search capabilities and sometimes even perform competitively with fine-tuned models. On linear-probe classification accuracy averaging over 7 tasks, our best unsupervised model achieves a relative improvement of 4% and 1.8% over previous best unsupervised and supervised text embedding models respectively. The same text embeddings when evaluated on large-scale semantic search attains a relative improvement of 23.4%, 14.7%, and 10.6% over previous best unsupervised methods on MSMARCO, Natural Questions and TriviaQA benchmarks, respectively. Similarly to text embeddings, we train code embedding models on (text, code) pairs, obtaining a 20.8% relative improvement over prior best work on code search.
Alleviating Noisy-label Effects in Image Classification via Probability Transition Matrix
Oct 19, 2021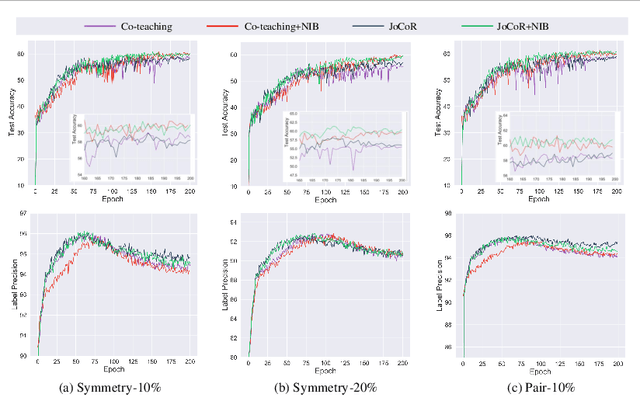
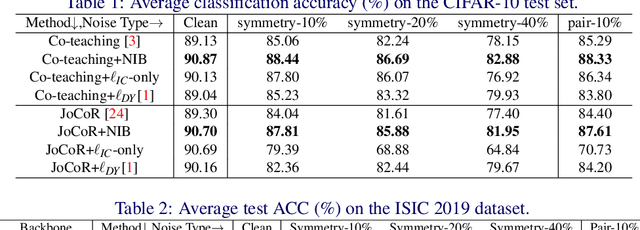
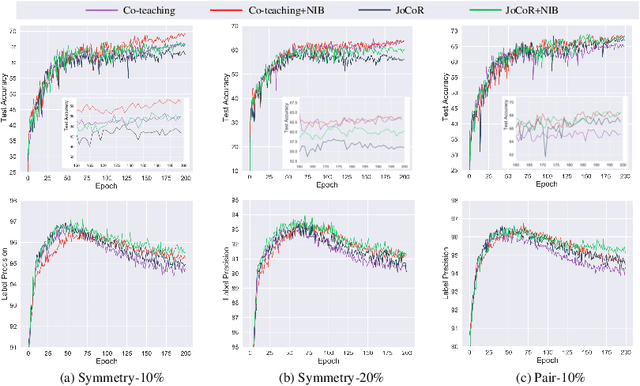
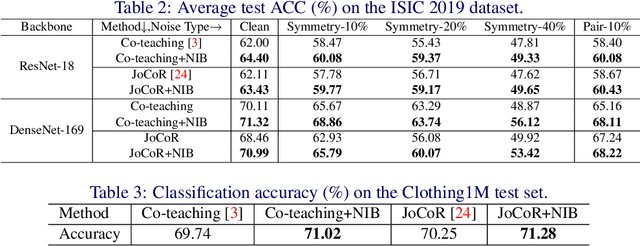
Abstract:Deep-learning-based image classification frameworks often suffer from the noisy label problem caused by the inter-observer variation. Recent studies employed learning-to-learn paradigms (e.g., Co-teaching and JoCoR) to filter the samples with noisy labels from the training set. However, most of them use a simple cross-entropy loss as the criterion for noisy label identification. The hard samples, which are beneficial for classifier learning, are often mistakenly treated as noises in such a setting since both the hard samples and ones with noisy labels lead to a relatively larger loss value than the easy cases. In this paper, we propose a plugin module, namely noise ignoring block (NIB), consisting of a probability transition matrix and an inter-class correlation (IC) loss, to separate the hard samples from the mislabeled ones, and further boost the accuracy of image classification network trained with noisy labels. Concretely, our IC loss is calculated as Kullback-Leibler divergence between the network prediction and the accumulative soft label generated by the probability transition matrix. Such that, with the lower value of IC loss, the hard cases can be easily distinguished from mislabeled cases. Extensive experiments are conducted on natural and medical image datasets (CIFAR-10 and ISIC 2019). The experimental results show that our NIB module consistently improves the performances of the state-of-the-art robust training methods.
Unsupervised Neural Machine Translation with Generative Language Models Only
Oct 11, 2021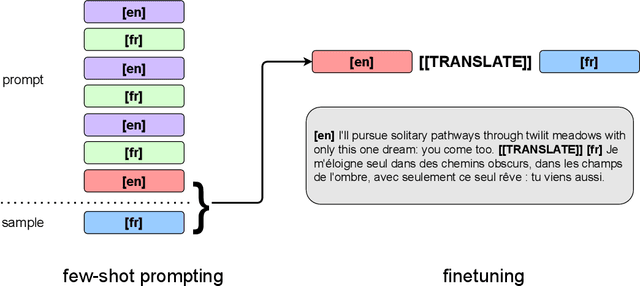
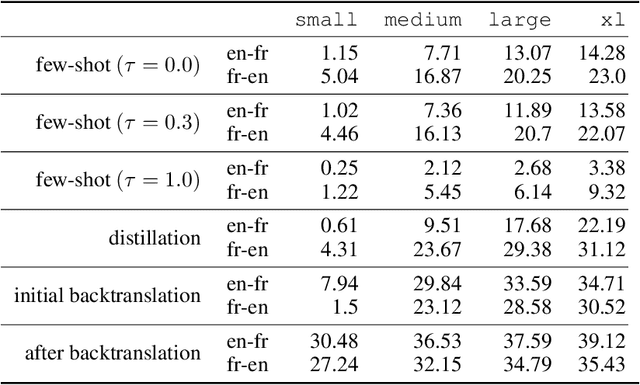
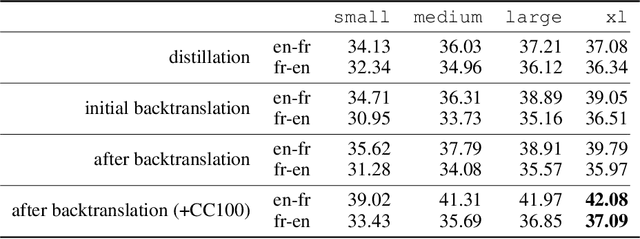
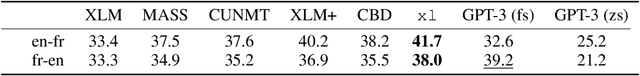
Abstract:We show how to derive state-of-the-art unsupervised neural machine translation systems from generatively pre-trained language models. Our method consists of three steps: few-shot amplification, distillation, and backtranslation. We first use the zero-shot translation ability of large pre-trained language models to generate translations for a small set of unlabeled sentences. We then amplify these zero-shot translations by using them as few-shot demonstrations for sampling a larger synthetic dataset. This dataset is distilled by discarding the few-shot demonstrations and then fine-tuning. During backtranslation, we repeatedly generate translations for a set of inputs and then fine-tune a single language model on both directions of the translation task at once, ensuring cycle-consistency by swapping the roles of gold monotext and generated translations when fine-tuning. By using our method to leverage GPT-3's zero-shot translation capability, we achieve a new state-of-the-art in unsupervised translation on the WMT14 English-French benchmark, attaining a BLEU score of 42.1.
Deep Image Synthesis from Intuitive User Input: A Review and Perspectives
Jul 09, 2021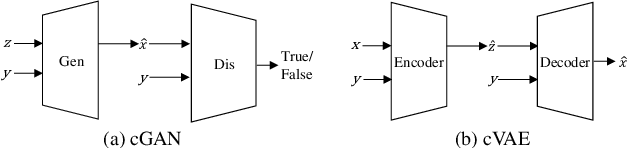
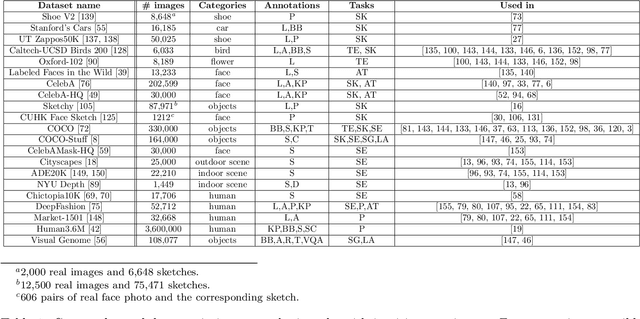

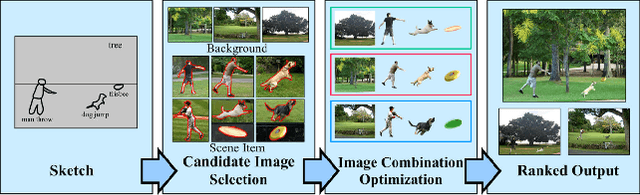
Abstract:In many applications of computer graphics, art and design, it is desirable for a user to provide intuitive non-image input, such as text, sketch, stroke, graph or layout, and have a computer system automatically generate photo-realistic images that adhere to the input content. While classic works that allow such automatic image content generation have followed a framework of image retrieval and composition, recent advances in deep generative models such as generative adversarial networks (GANs), variational autoencoders (VAEs), and flow-based methods have enabled more powerful and versatile image generation tasks. This paper reviews recent works for image synthesis given intuitive user input, covering advances in input versatility, image generation methodology, benchmark datasets, and evaluation metrics. This motivates new perspectives on input representation and interactivity, cross pollination between major image generation paradigms, and evaluation and comparison of generation methods.
* 26 pages, 7 figures, 1 table
FAIR1M: A Benchmark Dataset for Fine-grained Object Recognition in High-Resolution Remote Sensing Imagery
Mar 24, 2021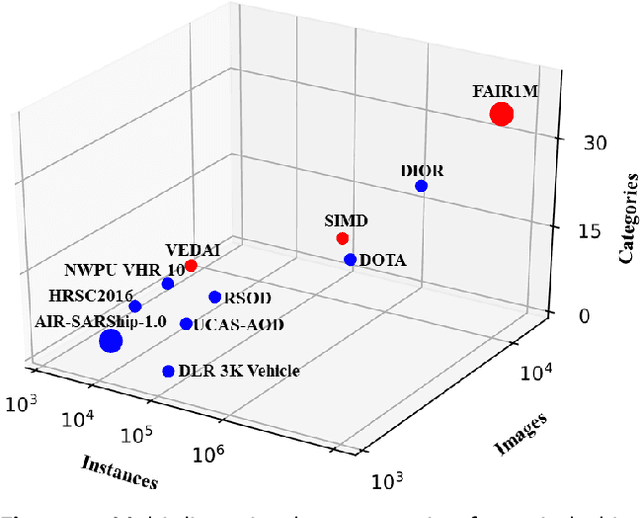
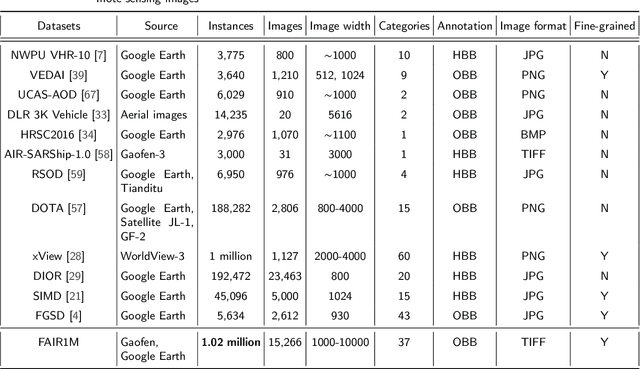
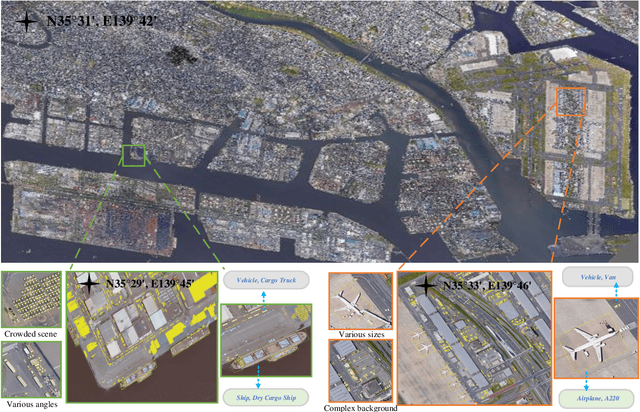
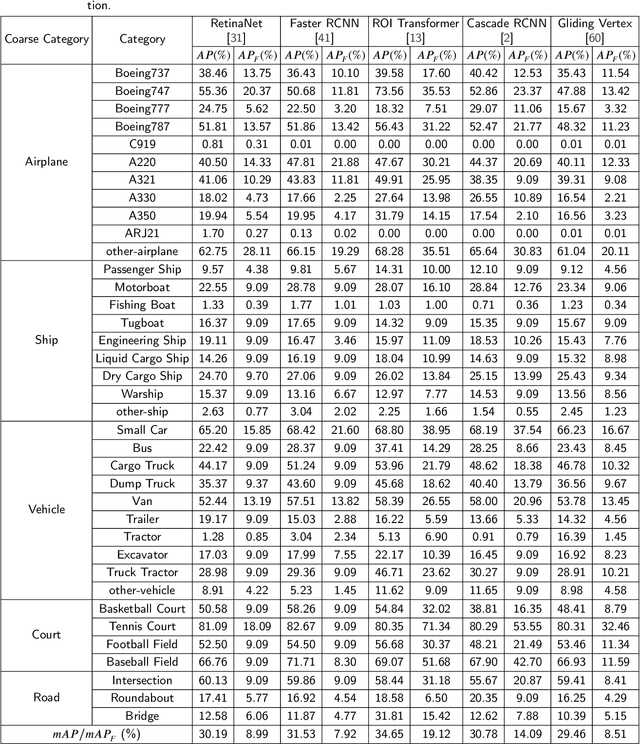
Abstract:With the rapid development of deep learning, many deep learning-based approaches have made great achievements in object detection task. It is generally known that deep learning is a data-driven method. Data directly impact the performance of object detectors to some extent. Although existing datasets have included common objects in remote sensing images, they still have some limitations in terms of scale, categories, and images. Therefore, there is a strong requirement for establishing a large-scale benchmark on object detection in high-resolution remote sensing images. In this paper, we propose a novel benchmark dataset with more than 1 million instances and more than 15,000 images for Fine-grAined object recognItion in high-Resolution remote sensing imagery which is named as FAIR1M. All objects in the FAIR1M dataset are annotated with respect to 5 categories and 37 sub-categories by oriented bounding boxes. Compared with existing detection datasets dedicated to object detection, the FAIR1M dataset has 4 particular characteristics: (1) it is much larger than other existing object detection datasets both in terms of the quantity of instances and the quantity of images, (2) it provides more rich fine-grained category information for objects in remote sensing images, (3) it contains geographic information such as latitude, longitude and resolution, (4) it provides better image quality owing to a careful data cleaning procedure. To establish a baseline for fine-grained object recognition, we propose a novel evaluation method and benchmark fine-grained object detection tasks and a visual classification task using several State-Of-The-Art (SOTA) deep learning-based models on our FAIR1M dataset. Experimental results strongly indicate that the FAIR1M dataset is closer to practical application and it is considerably more challenging than existing datasets.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge