Shuo Wang
State Key Laboratory of Management and Control for Complex Systems, Institute of Automation, Chinese Academy of Sciences, China
From Unaligned to Aligned: Scaling Multilingual LLMs with Multi-Way Parallel Corpora
May 20, 2025



Abstract:Continued pretraining and instruction tuning on large-scale multilingual data have proven to be effective in scaling large language models (LLMs) to low-resource languages. However, the unaligned nature of such data limits its ability to effectively capture cross-lingual semantics. In contrast, multi-way parallel data, where identical content is aligned across multiple languages, provides stronger cross-lingual consistency and offers greater potential for improving multilingual performance. In this paper, we introduce a large-scale, high-quality multi-way parallel corpus, TED2025, based on TED Talks. The corpus spans 113 languages, with up to 50 languages aligned in parallel, ensuring extensive multilingual coverage. Using this dataset, we investigate best practices for leveraging multi-way parallel data to enhance LLMs, including strategies for continued pretraining, instruction tuning, and the analysis of key influencing factors. Experiments on six multilingual benchmarks show that models trained on multiway parallel data consistently outperform those trained on unaligned multilingual data.
DD-Ranking: Rethinking the Evaluation of Dataset Distillation
May 19, 2025
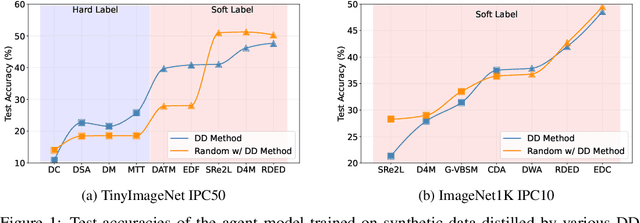
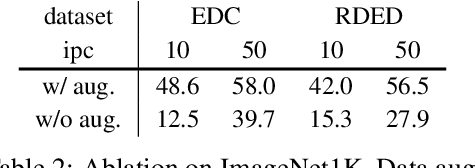
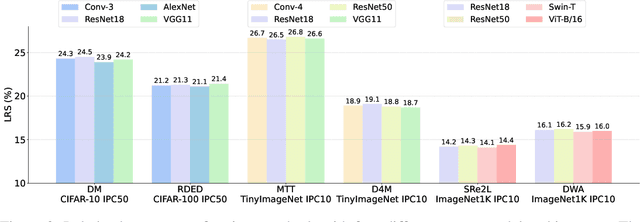
Abstract:In recent years, dataset distillation has provided a reliable solution for data compression, where models trained on the resulting smaller synthetic datasets achieve performance comparable to those trained on the original datasets. To further improve the performance of synthetic datasets, various training pipelines and optimization objectives have been proposed, greatly advancing the field of dataset distillation. Recent decoupled dataset distillation methods introduce soft labels and stronger data augmentation during the post-evaluation phase and scale dataset distillation up to larger datasets (e.g., ImageNet-1K). However, this raises a question: Is accuracy still a reliable metric to fairly evaluate dataset distillation methods? Our empirical findings suggest that the performance improvements of these methods often stem from additional techniques rather than the inherent quality of the images themselves, with even randomly sampled images achieving superior results. Such misaligned evaluation settings severely hinder the development of DD. Therefore, we propose DD-Ranking, a unified evaluation framework, along with new general evaluation metrics to uncover the true performance improvements achieved by different methods. By refocusing on the actual information enhancement of distilled datasets, DD-Ranking provides a more comprehensive and fair evaluation standard for future research advancements.
Aux-Think: Exploring Reasoning Strategies for Data-Efficient Vision-Language Navigation
May 17, 2025Abstract:Vision-Language Navigation (VLN) is a critical task for developing embodied agents that can follow natural language instructions to navigate in complex real-world environments. Recent advances in VLN by large pretrained models have significantly improved generalization and instruction grounding compared to traditional approaches. However, the role of reasoning strategies in navigation-an action-centric, long-horizon task-remains underexplored, despite Chain-of-Thought (CoT) reasoning's demonstrated success in static tasks like visual question answering. To address this gap, we conduct the first systematic evaluation of reasoning strategies for VLN, including No-Think (direct action prediction), Pre-Think (reason before action), and Post-Think (reason after action). Surprisingly, our findings reveal the Inference-time Reasoning Collapse issue, where inference-time reasoning degrades navigation accuracy, highlighting the challenges of integrating reasoning into VLN. Based on this insight, we propose Aux-Think, a framework that trains models to internalize structured reasoning patterns through CoT supervision, while inferring action directly without reasoning in online prediction. To support this framework, we release R2R-CoT-320k, the first Chain-of-Thought annotated dataset for VLN. Extensive experiments show that Aux-Think reduces training effort greatly and achieves the best performance under the same data scale.
Patient-Specific Dynamic Digital-Physical Twin for Coronary Intervention Training: An Integrated Mixed Reality Approach
May 16, 2025Abstract:Background and Objective: Precise preoperative planning and effective physician training for coronary interventions are increasingly important. Despite advances in medical imaging technologies, transforming static or limited dynamic imaging data into comprehensive dynamic cardiac models remains challenging. Existing training systems lack accurate simulation of cardiac physiological dynamics. This study develops a comprehensive dynamic cardiac model research framework based on 4D-CTA, integrating digital twin technology, computer vision, and physical model manufacturing to provide precise, personalized tools for interventional cardiology. Methods: Using 4D-CTA data from a 60-year-old female with three-vessel coronary stenosis, we segmented cardiac chambers and coronary arteries, constructed dynamic models, and implemented skeletal skinning weight computation to simulate vessel deformation across 20 cardiac phases. Transparent vascular physical models were manufactured using medical-grade silicone. We developed cardiac output analysis and virtual angiography systems, implemented guidewire 3D reconstruction using binocular stereo vision, and evaluated the system through angiography validation and CABG training applications. Results: Morphological consistency between virtual and real angiography reached 80.9%. Dice similarity coefficients for guidewire motion ranged from 0.741-0.812, with mean trajectory errors below 1.1 mm. The transparent model demonstrated advantages in CABG training, allowing direct visualization while simulating beating heart challenges. Conclusion: Our patient-specific digital-physical twin approach effectively reproduces both anatomical structures and dynamic characteristics of coronary vasculature, offering a dynamic environment with visual and tactile feedback valuable for education and clinical planning.
VTLA: Vision-Tactile-Language-Action Model with Preference Learning for Insertion Manipulation
May 14, 2025Abstract:While vision-language models have advanced significantly, their application in language-conditioned robotic manipulation is still underexplored, especially for contact-rich tasks that extend beyond visually dominant pick-and-place scenarios. To bridge this gap, we introduce Vision-Tactile-Language-Action model, a novel framework that enables robust policy generation in contact-intensive scenarios by effectively integrating visual and tactile inputs through cross-modal language grounding. A low-cost, multi-modal dataset has been constructed in a simulation environment, containing vision-tactile-action-instruction pairs specifically designed for the fingertip insertion task. Furthermore, we introduce Direct Preference Optimization (DPO) to offer regression-like supervision for the VTLA model, effectively bridging the gap between classification-based next token prediction loss and continuous robotic tasks. Experimental results show that the VTLA model outperforms traditional imitation learning methods (e.g., diffusion policies) and existing multi-modal baselines (TLA/VLA), achieving over 90% success rates on unseen peg shapes. Finally, we conduct real-world peg-in-hole experiments to demonstrate the exceptional Sim2Real performance of the proposed VTLA model. For supplementary videos and results, please visit our project website: https://sites.google.com/view/vtla
Endo-CLIP: Progressive Self-Supervised Pre-training on Raw Colonoscopy Records
May 14, 2025



Abstract:Pre-training on image-text colonoscopy records offers substantial potential for improving endoscopic image analysis, but faces challenges including non-informative background images, complex medical terminology, and ambiguous multi-lesion descriptions. We introduce Endo-CLIP, a novel self-supervised framework that enhances Contrastive Language-Image Pre-training (CLIP) for this domain. Endo-CLIP's three-stage framework--cleansing, attunement, and unification--addresses these challenges by (1) removing background frames, (2) leveraging large language models to extract clinical attributes for fine-grained contrastive learning, and (3) employing patient-level cross-attention to resolve multi-polyp ambiguities. Extensive experiments demonstrate that Endo-CLIP significantly outperforms state-of-the-art pre-training methods in zero-shot and few-shot polyp detection and classification, paving the way for more accurate and clinically relevant endoscopic analysis.
CLTP: Contrastive Language-Tactile Pre-training for 3D Contact Geometry Understanding
May 13, 2025Abstract:Recent advancements in integrating tactile sensing with vision-language models (VLMs) have demonstrated remarkable potential for robotic multimodal perception. However, existing tactile descriptions remain limited to superficial attributes like texture, neglecting critical contact states essential for robotic manipulation. To bridge this gap, we propose CLTP, an intuitive and effective language tactile pretraining framework that aligns tactile 3D point clouds with natural language in various contact scenarios, thus enabling contact-state-aware tactile language understanding for contact-rich manipulation tasks. We first collect a novel dataset of 50k+ tactile 3D point cloud-language pairs, where descriptions explicitly capture multidimensional contact states (e.g., contact location, shape, and force) from the tactile sensor's perspective. CLTP leverages a pre-aligned and frozen vision-language feature space to bridge holistic textual and tactile modalities. Experiments validate its superiority in three downstream tasks: zero-shot 3D classification, contact state classification, and tactile 3D large language model (LLM) interaction. To the best of our knowledge, this is the first study to align tactile and language representations from the contact state perspective for manipulation tasks, providing great potential for tactile-language-action model learning. Code and datasets are open-sourced at https://sites.google.com/view/cltp/.
Avoid Recommending Out-of-Domain Items: Constrained Generative Recommendation with LLMs
May 06, 2025Abstract:Large Language Models (LLMs) have shown promise for generative recommender systems due to their transformative capabilities in user interaction. However, ensuring they do not recommend out-of-domain (OOD) items remains a challenge. We study two distinct methods to address this issue: RecLM-ret, a retrieval-based method, and RecLM-cgen, a constrained generation method. Both methods integrate seamlessly with existing LLMs to ensure in-domain recommendations. Comprehensive experiments on three recommendation datasets demonstrate that RecLM-cgen consistently outperforms RecLM-ret and existing LLM-based recommender models in accuracy while eliminating OOD recommendations, making it the preferred method for adoption. Additionally, RecLM-cgen maintains strong generalist capabilities and is a lightweight plug-and-play module for easy integration into LLMs, offering valuable practical benefits for the community. Source code is available at https://github.com/microsoft/RecAI
Dynamic Arthroscopic Navigation System for Anterior Cruciate Ligament Reconstruction Based on Multi-level Memory Architecture
Apr 28, 2025Abstract:This paper presents a dynamic arthroscopic navigation system based on multi-level memory architecture for anterior cruciate ligament (ACL) reconstruction surgery. The system extends our previously proposed markerless navigation method from static image matching to dynamic video sequence tracking. By integrating the Atkinson-Shiffrin memory model's three-level architecture (sensory memory, working memory, and long-term memory), our system maintains continuous tracking of the femoral condyle throughout the surgical procedure, providing stable navigation support even in complex situations involving viewpoint changes, instrument occlusion, and tissue deformation. Unlike existing methods, our system operates in real-time on standard arthroscopic equipment without requiring additional tracking hardware, achieving 25.3 FPS with a latency of only 39.5 ms, representing a 3.5-fold improvement over our previous static system. For extended sequences (1000 frames), the dynamic system maintained an error of 5.3 plus-minus 1.5 pixels, compared to the static system's 12.6 plus-minus 3.7 pixels - an improvement of approximately 45 percent. For medium-length sequences (500 frames) and short sequences (100 frames), the system achieved approximately 35 percent and 19 percent accuracy improvements, respectively. Experimental results demonstrate the system overcomes limitations of traditional static matching methods, providing new technical support for improving surgical precision in ACL reconstruction.
Innovative Integration of 4D Cardiovascular Reconstruction and Hologram: A New Visualization Tool for Coronary Artery Bypass Grafting Planning
Apr 28, 2025Abstract:Background: Coronary artery bypass grafting (CABG) planning requires advanced spatial visualization and consideration of coronary artery depth, calcification, and pericardial adhesions. Objective: To develop and evaluate a dynamic cardiovascular holographic visualization tool for preoperative CABG planning. Methods: Using 4D cardiac computed tomography angiography data from 14 CABG candidates, we developed a semi-automated workflow for time-resolved segmentation of cardiac structures, epicardial adipose tissue (EAT), and coronary arteries with calcium scoring. The workflow incorporated methods for cardiac segmentation, coronary calcification quantification, visualization of coronary depth within EAT, and pericardial adhesion assessment through motion analysis. Dynamic cardiovascular holograms were displayed using the Looking Glass platform. Thirteen cardiac surgeons evaluated the tool using a Likert scale. Additionally, pericardial adhesion scores from holograms of 21 patients (including seven undergoing secondary cardiac surgeries) were compared with intraoperative findings. Results: Surgeons rated the visualization tool highly for preoperative planning utility (mean Likert score: 4.57/5.0). Hologram-based pericardial adhesion scoring strongly correlated with intraoperative findings (r=0.786, P<0.001). Conclusion: This study establishes a visualization framework for CABG planning that produces clinically relevant dynamic holograms from patient-specific data, with clinical feedback confirming its effectiveness for preoperative planning.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge