Junyu Chen
ConFit v3: Improving Resume-Job Matching with LLM-based Re-Ranking
May 10, 2026Abstract:A reliable resume-job matching system helps a company find suitable candidates from a pool of resumes and helps a job seeker find relevant jobs from a list of job posts. While recent advances in embedding-based methods such as ConFit and ConFit v2 can efficiently retrieve candidates at scale, the lack of controllability and explainability limits their real-world adaptations. LLM-based re-rankers can address these limitations through reasoning, but existing training recipes are developed on short-document benchmarks and do not account for noise in real-world recruiting data. In this work, we first conduct a systematic analysis over the LLM re-ranker training pipeline for person-job fit, covering inference algorithm design, RL algorithm selection, data processing, and SFT distillation. We find that using multi-pass re-ranking, training with listwise RL objectives, removing noisy samples, and distilling from a stronger LLM before RL significantly improves re-ranking performance. We then aggregate these findings to train ConFit v3 with Qwen3-8B and Qwen3-32B on real-world person-job fit datasets, and find significant improvements over existing best person-job fit systems as well as strong LLMs such as GPT-5 and Claude Opus-4.5. We hope our findings provide useful insights for future research on adapting LLM-based re-rankers to person-job fit systems.
CuraLight: Debate-Guided Data Curation for LLM-Centered Traffic Signal Control
Apr 07, 2026Abstract:Traffic signal control (TSC) is a core component of intelligent transportation systems (ITS), aiming to reduce congestion, emissions, and travel time. Recent approaches based on reinforcement learning (RL) and large language models (LLMs) have improved adaptivity, but still suffer from limited interpretability, insufficient interaction data, and weak generalization to heterogeneous intersections. This paper proposes CuraLight, an LLM-centered framework where an RL agent assists the fine-tuning of an LLM-based traffic signal controller. The RL agent explores traffic environments and generates high-quality interaction trajectories, which are converted into prompt-response pairs for imitation fine-tuning. A multi-LLM ensemble deliberation system further evaluates candidate signal timing actions through structured debate, providing preference-aware supervision signals for training. Experiments conducted in SUMO across heterogeneous real-world networks from Jinan, Hangzhou, and Yizhuang demonstrate that CuraLight consistently outperforms state-of-the-art baselines, reducing average travel time by 5.34 percent, average queue length by 5.14 percent, and average waiting time by 7.02 percent. The results highlight the effectiveness of combining RL-assisted exploration with deliberation-based data curation for scalable and interpretable traffic signal control.
Solving a Nonlinear Blind Inverse Problem for Tagged MRI with Physics and Deep Generative Priors
Mar 01, 2026Abstract:Tagged MRI enables tracking internal tissue motion non-invasively. It encodes motion by modulating anatomy with periodic tags, which deform along with tissue. However, the entanglement between anatomy, tags and motion poses significant challenges for post-processing. The existence of tags and imaging blur hinders downstream tasks such as segmenting anatomy. Tag fading, due to T1-relaxation, disrupts the brightness constancy assumption for motion tracking. For decades, these challenges have been handled in isolation and sub-optimally. In contrast, we introduce a blind and nonlinear inverse framework for tagged MRI that, for the first time, unifies these tasks: anatomical image recovery, high-resolution cine image synthesis, and motion estimation. At its core, the synergy of MR physics and generative priors enables us to blindly estimate the unknown forward imaging models and high-resolution underlying anatomy, while simultaneously tracking 3D diffeomorphic Lagrangian motion over time. Experiments on tagged brain MRI demonstrate that our approach yields high-resolution anatomy images, cine images, and more accurate motion than specialized methods.
Transformer-based CoVaR: Systemic Risk in Textual Information
Feb 13, 2026Abstract:Conditional Value-at-Risk (CoVaR) quantifies systemic financial risk by measuring the loss quantile of one asset, conditional on another asset experiencing distress. We develop a Transformer-based methodology that integrates financial news articles directly with market data to improve CoVaR estimates. Unlike approaches that use predefined sentiment scores, our method incorporates raw text embeddings generated by a large language model (LLM). We prove explicit error bounds for our Transformer CoVaR estimator, showing that accurate CoVaR learning is possible even with small datasets. Using U.S. market returns and Reuters news items from 2006--2013, our out-of-sample results show that textual information impacts the CoVaR forecasts. With better predictive performance, we identify a pronounced negative dip during market stress periods across several equity assets when comparing the Transformer-based CoVaR to both the CoVaR without text and the CoVaR using traditional sentiment measures. Our results show that textual data can be used to effectively model systemic risk without requiring prohibitively large data sets.
BPDQ: Bit-Plane Decomposition Quantization on a Variable Grid for Large Language Models
Feb 04, 2026Abstract:Large language model (LLM) inference is often bounded by memory footprint and memory bandwidth in resource-constrained deployments, making quantization a fundamental technique for efficient serving. While post-training quantization (PTQ) maintains high fidelity at 4-bit, it deteriorates at 2-3 bits. Fundamentally, existing methods enforce a shape-invariant quantization grid (e.g., the fixed uniform intervals of UINT2) for each group, severely restricting the feasible set for error minimization. To address this, we propose Bit-Plane Decomposition Quantization (BPDQ), which constructs a variable quantization grid via bit-planes and scalar coefficients, and iteratively refines them using approximate second-order information while progressively compensating quantization errors to minimize output discrepancy. In the 2-bit regime, BPDQ enables serving Qwen2.5-72B on a single RTX 3090 with 83.85% GSM8K accuracy (vs. 90.83% at 16-bit). Moreover, we provide theoretical analysis showing that the variable grid expands the feasible set, and that the quantization process consistently aligns with the optimization objective in Hessian-induced geometry. Code: github.com/KingdalfGoodman/BPDQ.
Deep classifier kriging for probabilistic spatial prediction of air quality index
Dec 29, 2025Abstract:Accurate spatial interpolation of the air quality index (AQI), computed from concentrations of multiple air pollutants, is essential for regulatory decision-making, yet AQI fields are inherently non-Gaussian and often exhibit complex nonlinear spatial structure. Classical spatial prediction methods such as kriging are linear and rely on Gaussian assumptions, which limits their ability to capture these features and to provide reliable predictive distributions. In this study, we propose \textit{deep classifier kriging} (DCK), a flexible, distribution-free deep learning framework for estimating full predictive distribution functions for univariate and bivariate spatial processes, together with a \textit{data fusion} mechanism that enables modeling of non-collocated bivariate processes and integration of heterogeneous air pollution data sources. Through extensive simulation experiments, we show that DCK consistently outperforms conventional approaches in predictive accuracy and uncertainty quantification. We further apply DCK to probabilistic spatial prediction of AQI by fusing sparse but high-quality station observations with spatially continuous yet biased auxiliary model outputs, yielding spatially resolved predictive distributions that support downstream tasks such as exceedance and extreme-event probability estimation for regulatory risk assessment and policy formulation.
Retrieval Augmented Generation-Enhanced Distributed LLM Agents for Generalizable Traffic Signal Control with Emergency Vehicles
Oct 30, 2025



Abstract:With increasing urban traffic complexity, Traffic Signal Control (TSC) is essential for optimizing traffic flow and improving road safety. Large Language Models (LLMs) emerge as promising approaches for TSC. However, they are prone to hallucinations in emergencies, leading to unreliable decisions that may cause substantial delays for emergency vehicles. Moreover, diverse intersection types present substantial challenges for traffic state encoding and cross-intersection training, limiting generalization across heterogeneous intersections. Therefore, this paper proposes Retrieval Augmented Generation (RAG)-enhanced distributed LLM agents with Emergency response for Generalizable TSC (REG-TSC). Firstly, this paper presents an emergency-aware reasoning framework, which dynamically adjusts reasoning depth based on the emergency scenario and is equipped with a novel Reviewer-based Emergency RAG (RERAG) to distill specific knowledge and guidance from historical cases, enhancing the reliability and rationality of agents' emergency decisions. Secondly, this paper designs a type-agnostic traffic representation and proposes a Reward-guided Reinforced Refinement (R3) for heterogeneous intersections. R3 adaptively samples training experience from diverse intersections with environment feedback-based priority and fine-tunes LLM agents with a designed reward-weighted likelihood loss, guiding REG-TSC toward high-reward policies across heterogeneous intersections. On three real-world road networks with 17 to 177 heterogeneous intersections, extensive experiments show that REG-TSC reduces travel time by 42.00%, queue length by 62.31%, and emergency vehicle waiting time by 83.16%, outperforming other state-of-the-art methods.
Pretraining Deformable Image Registration Networks with Random Images
May 30, 2025Abstract:Recent advances in deep learning-based medical image registration have shown that training deep neural networks~(DNNs) does not necessarily require medical images. Previous work showed that DNNs trained on randomly generated images with carefully designed noise and contrast properties can still generalize well to unseen medical data. Building on this insight, we propose using registration between random images as a proxy task for pretraining a foundation model for image registration. Empirical results show that our pretraining strategy improves registration accuracy, reduces the amount of domain-specific data needed to achieve competitive performance, and accelerates convergence during downstream training, thereby enhancing computational efficiency.
Beyond the LUMIR challenge: The pathway to foundational registration models
May 30, 2025



Abstract:Medical image challenges have played a transformative role in advancing the field, catalyzing algorithmic innovation and establishing new performance standards across diverse clinical applications. Image registration, a foundational task in neuroimaging pipelines, has similarly benefited from the Learn2Reg initiative. Building on this foundation, we introduce the Large-scale Unsupervised Brain MRI Image Registration (LUMIR) challenge, a next-generation benchmark designed to assess and advance unsupervised brain MRI registration. Distinct from prior challenges that leveraged anatomical label maps for supervision, LUMIR removes this dependency by providing over 4,000 preprocessed T1-weighted brain MRIs for training without any label maps, encouraging biologically plausible deformation modeling through self-supervision. In addition to evaluating performance on 590 held-out test subjects, LUMIR introduces a rigorous suite of zero-shot generalization tasks, spanning out-of-domain imaging modalities (e.g., FLAIR, T2-weighted, T2*-weighted), disease populations (e.g., Alzheimer's disease), acquisition protocols (e.g., 9.4T MRI), and species (e.g., macaque brains). A total of 1,158 subjects and over 4,000 image pairs were included for evaluation. Performance was assessed using both segmentation-based metrics (Dice coefficient, 95th percentile Hausdorff distance) and landmark-based registration accuracy (target registration error). Across both in-domain and zero-shot tasks, deep learning-based methods consistently achieved state-of-the-art accuracy while producing anatomically plausible deformation fields. The top-performing deep learning-based models demonstrated diffeomorphic properties and inverse consistency, outperforming several leading optimization-based methods, and showing strong robustness to most domain shifts, the exception being a drop in performance on out-of-domain contrasts.
Deep learning-derived arterial input function
May 30, 2025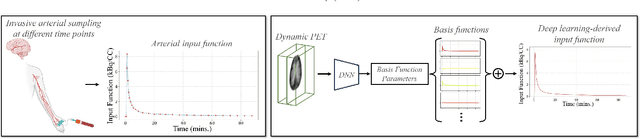
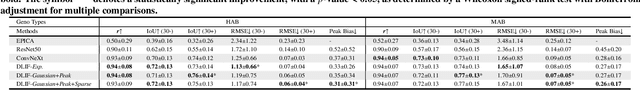
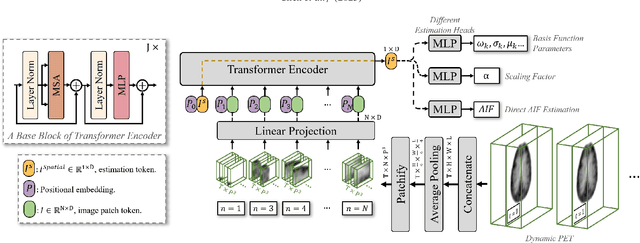
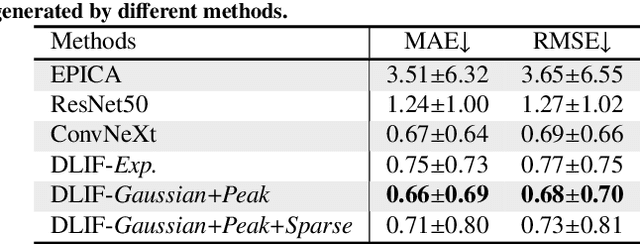
Abstract:Dynamic positron emission tomography (PET) imaging combined with radiotracer kinetic modeling is a powerful technique for visualizing biological processes in the brain, offering valuable insights into brain functions and neurological disorders such as Alzheimer's and Parkinson's diseases. Accurate kinetic modeling relies heavily on the use of a metabolite-corrected arterial input function (AIF), which typically requires invasive and labor-intensive arterial blood sampling. While alternative non-invasive approaches have been proposed, they often compromise accuracy or still necessitate at least one invasive blood sampling. In this study, we present the deep learning-derived arterial input function (DLIF), a deep learning framework capable of estimating a metabolite-corrected AIF directly from dynamic PET image sequences without any blood sampling. We validated DLIF using existing dynamic PET patient data. We compared DLIF and resulting parametric maps against ground truth measurements. Our evaluation shows that DLIF achieves accurate and robust AIF estimation. By leveraging deep learning's ability to capture complex temporal dynamics and incorporating prior knowledge of typical AIF shapes through basis functions, DLIF provides a rapid, accurate, and entirely non-invasive alternative to traditional AIF measurement methods.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge