Huazhu Fu
Learning to screen Glaucoma like the ophthalmologists
Sep 23, 2022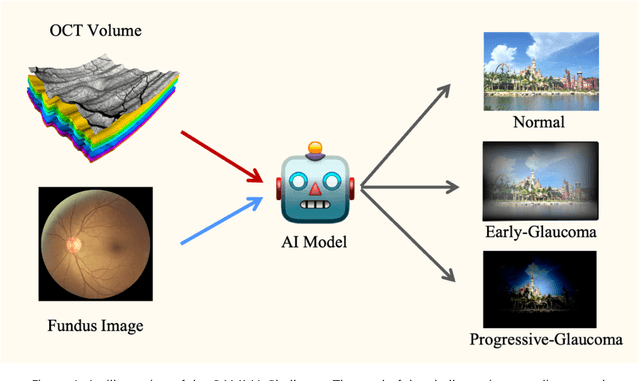
Abstract:GAMMA Challenge is organized to encourage the AI models to screen the glaucoma from a combination of 2D fundus image and 3D optical coherence tomography volume, like the ophthalmologists.
Unsupervised Domain Adaptation via Style-Aware Self-intermediate Domain
Sep 05, 2022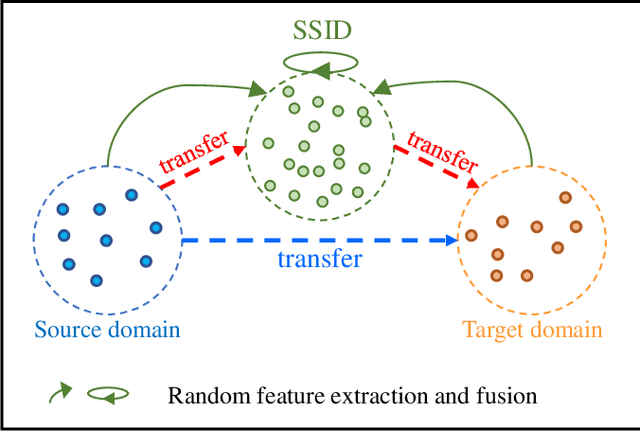
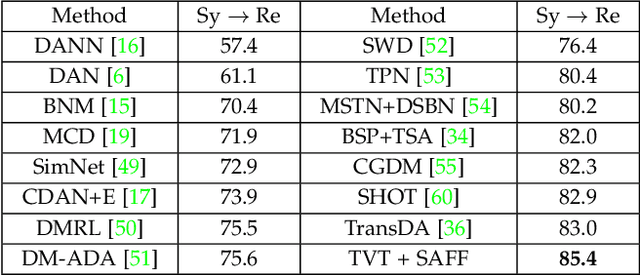
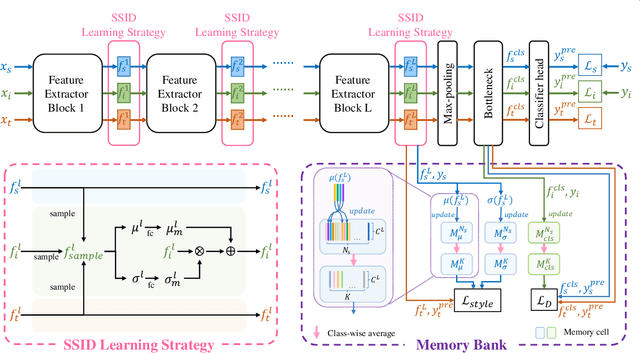
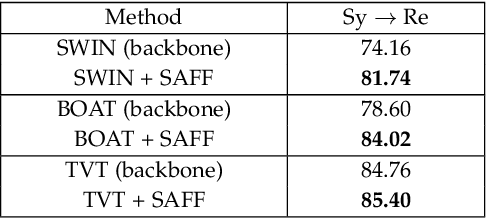
Abstract:Unsupervised domain adaptation (UDA) has attracted considerable attention, which transfers knowledge from a label-rich source domain to a related but unlabeled target domain. Reducing inter-domain differences has always been a crucial factor to improve performance in UDA, especially for tasks where there is a large gap between source and target domains. To this end, we propose a novel style-aware feature fusion method (SAFF) to bridge the large domain gap and transfer knowledge while alleviating the loss of class-discriminative information. Inspired by the human transitive inference and learning ability, a novel style-aware self-intermediate domain (SSID) is investigated to link two seemingly unrelated concepts through a series of intermediate auxiliary synthesized concepts. Specifically, we propose a novel learning strategy of SSID, which selects samples from both source and target domains as anchors, and then randomly fuses the object and style features of these anchors to generate labeled and style-rich intermediate auxiliary features for knowledge transfer. Moreover, we design an external memory bank to store and update specified labeled features to obtain stable class features and class-wise style features. Based on the proposed memory bank, the intra- and inter-domain loss functions are designed to improve the class recognition ability and feature compatibility, respectively. Meanwhile, we simulate the rich latent feature space of SSID by infinite sampling and the convergence of the loss function by mathematical theory. Finally, we conduct comprehensive experiments on commonly used domain adaptive benchmarks to evaluate the proposed SAFF, and the experimental results show that the proposed SAFF can be easily combined with different backbone networks and obtain better performance as a plug-in-plug-out module.
Dataset and Evaluation algorithm design for GOALS Challenge
Jul 29, 2022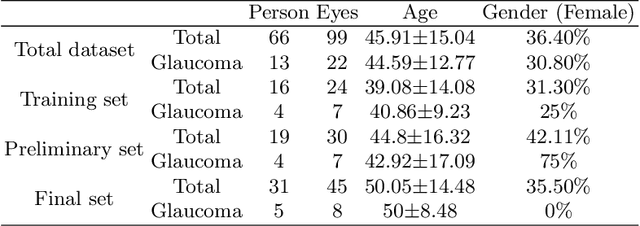
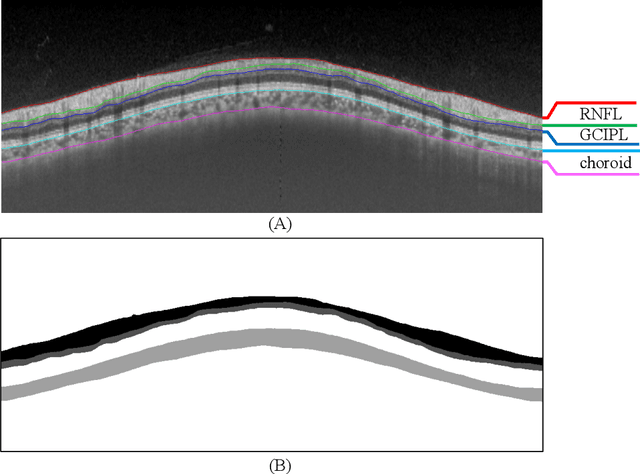
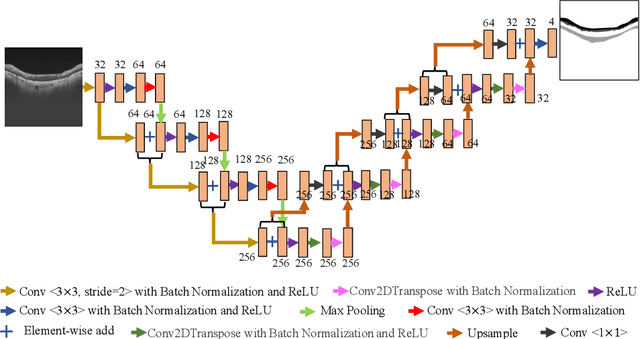
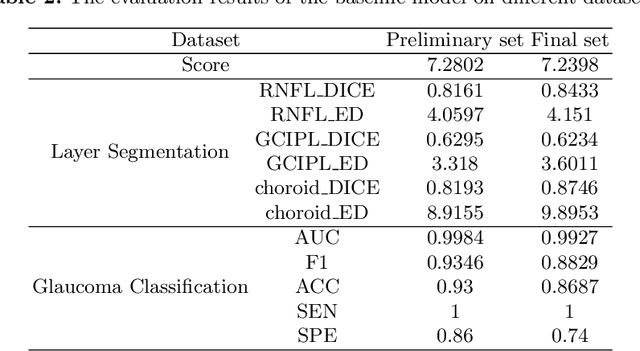
Abstract:Glaucoma causes irreversible vision loss due to damage to the optic nerve, and there is no cure for glaucoma.OCT imaging modality is an essential technique for assessing glaucomatous damage since it aids in quantifying fundus structures. To promote the research of AI technology in the field of OCT-assisted diagnosis of glaucoma, we held a Glaucoma OCT Analysis and Layer Segmentation (GOALS) Challenge in conjunction with the International Conference on Medical Image Computing and Computer Assisted Intervention (MICCAI) 2022 to provide data and corresponding annotations for researchers studying layer segmentation from OCT images and the classification of glaucoma. This paper describes the released 300 circumpapillary OCT images, the baselines of the two sub-tasks, and the evaluation methodology. The GOALS Challenge is accessible at https://aistudio.baidu.com/aistudio/competition/detail/230.
A New Dataset and A Baseline Model for Breast Lesion Detection in Ultrasound Videos
Jul 01, 2022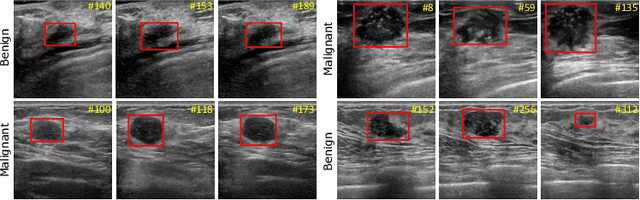
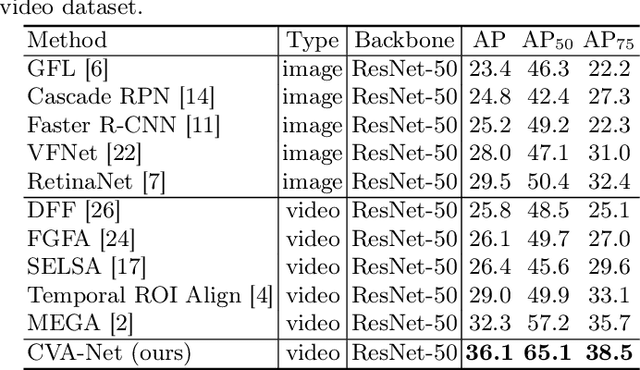
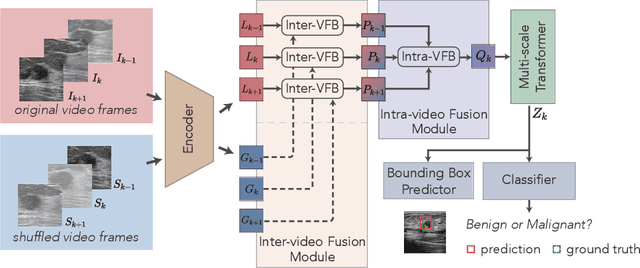
Abstract:Breast lesion detection in ultrasound is critical for breast cancer diagnosis. Existing methods mainly rely on individual 2D ultrasound images or combine unlabeled video and labeled 2D images to train models for breast lesion detection. In this paper, we first collect and annotate an ultrasound video dataset (188 videos) for breast lesion detection. Moreover, we propose a clip-level and video-level feature aggregated network (CVA-Net) for addressing breast lesion detection in ultrasound videos by aggregating video-level lesion classification features and clip-level temporal features. The clip-level temporal features encode local temporal information of ordered video frames and global temporal information of shuffled video frames. In our CVA-Net, an inter-video fusion module is devised to fuse local features from original video frames and global features from shuffled video frames, and an intra-video fusion module is devised to learn the temporal information among adjacent video frames. Moreover, we learn video-level features to classify the breast lesions of the original video as benign or malignant lesions to further enhance the final breast lesion detection performance in ultrasound videos. Experimental results on our annotated dataset demonstrate that our CVA-Net clearly outperforms state-of-the-art methods. The corresponding code and dataset are publicly available at \url{https://github.com/jhl-Det/CVA-Net}.
* 11 pages, 4 figures
TBraTS: Trusted Brain Tumor Segmentation
Jun 30, 2022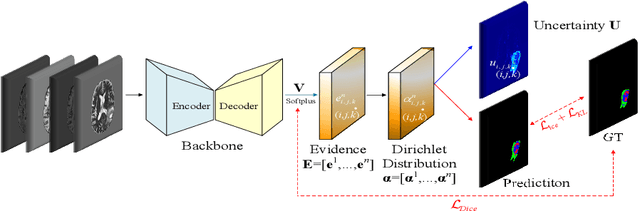
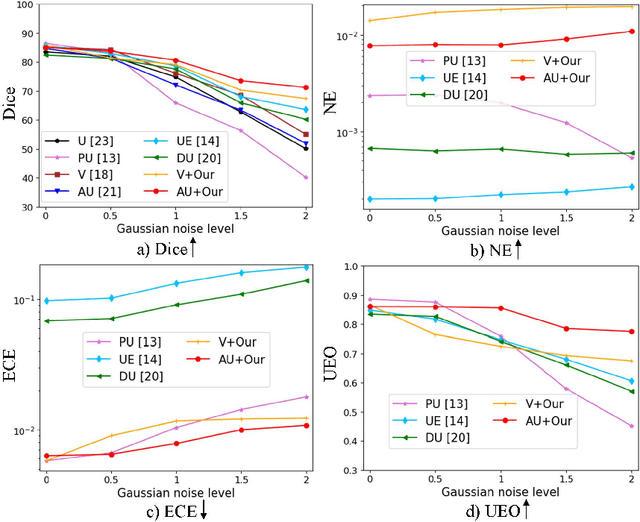
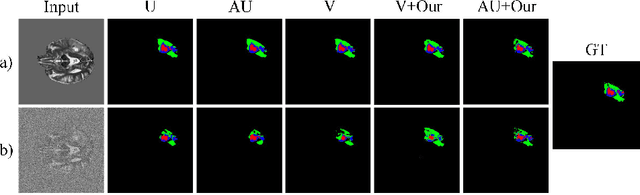
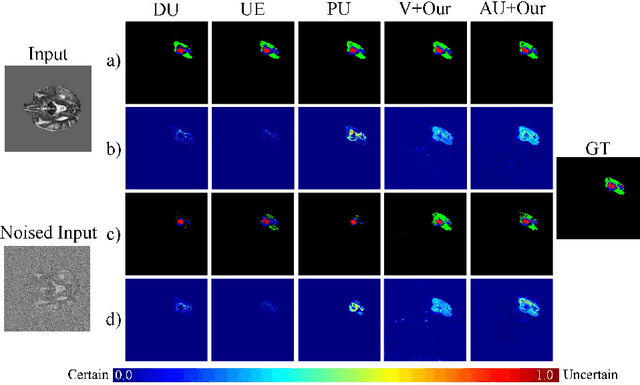
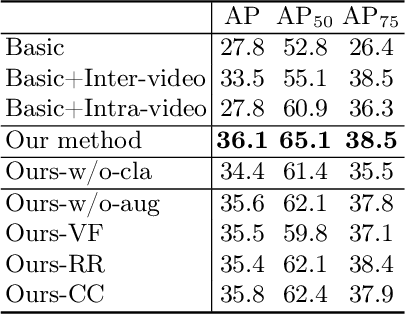
Abstract:Despite recent improvements in the accuracy of brain tumor segmentation, the results still exhibit low levels of confidence and robustness. Uncertainty estimation is one effective way to change this situation, as it provides a measure of confidence in the segmentation results. In this paper, we propose a trusted brain tumor segmentation network which can generate robust segmentation results and reliable uncertainty estimations without excessive computational burden and modification of the backbone network. In our method, uncertainty is modeled explicitly using subjective logic theory, which treats the predictions of backbone neural network as subjective opinions by parameterizing the class probabilities of the segmentation as a Dirichlet distribution. Meanwhile, the trusted segmentation framework learns the function that gathers reliable evidence from the feature leading to the final segmentation results. Overall, our unified trusted segmentation framework endows the model with reliability and robustness to out-of-distribution samples. To evaluate the effectiveness of our model in robustness and reliability, qualitative and quantitative experiments are conducted on the BraTS 2019 dataset.
Structure-consistent Restoration Network for Cataract Fundus Image Enhancement
Jun 09, 2022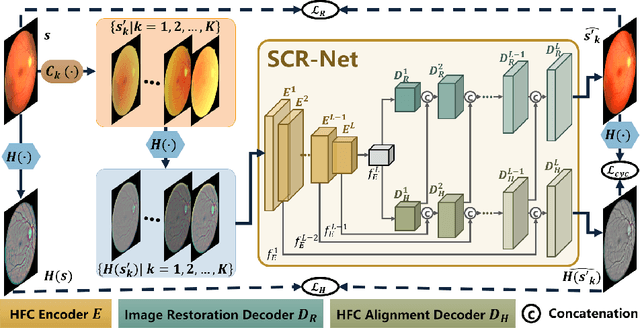
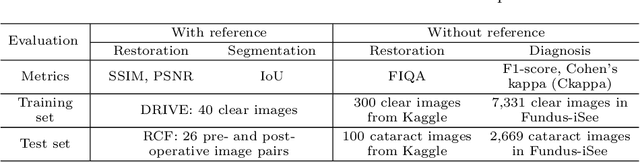
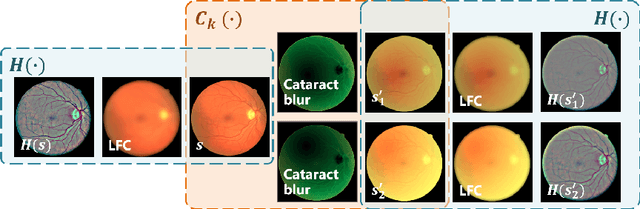
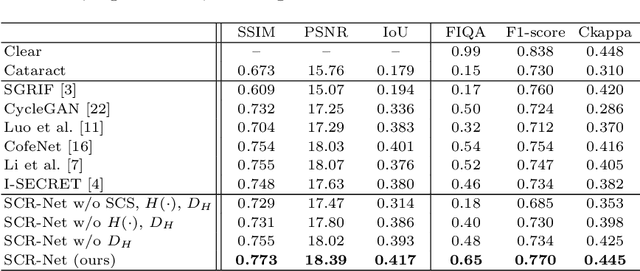
Abstract:Fundus photography is a routine examination in clinics to diagnose and monitor ocular diseases. However, for cataract patients, the fundus image always suffers quality degradation caused by the clouding lens. The degradation prevents reliable diagnosis by ophthalmologists or computer-aided systems. To improve the certainty in clinical diagnosis, restoration algorithms have been proposed to enhance the quality of fundus images. Unfortunately, challenges remain in the deployment of these algorithms, such as collecting sufficient training data and preserving retinal structures. In this paper, to circumvent the strict deployment requirement, a structure-consistent restoration network (SCR-Net) for cataract fundus images is developed from synthesized data that shares an identical structure. A cataract simulation model is firstly designed to collect synthesized cataract sets (SCS) formed by cataract fundus images sharing identical structures. Then high-frequency components (HFCs) are extracted from the SCS to constrain structure consistency such that the structure preservation in SCR-Net is enforced. The experiments demonstrate the effectiveness of SCR-Net in the comparison with state-of-the-art methods and the follow-up clinical applications. The code is available at https://github.com/liamheng/ArcNet-Medical-Image-Enhancement.
GCoNet+: A Stronger Group Collaborative Co-Salient Object Detector
Jun 01, 2022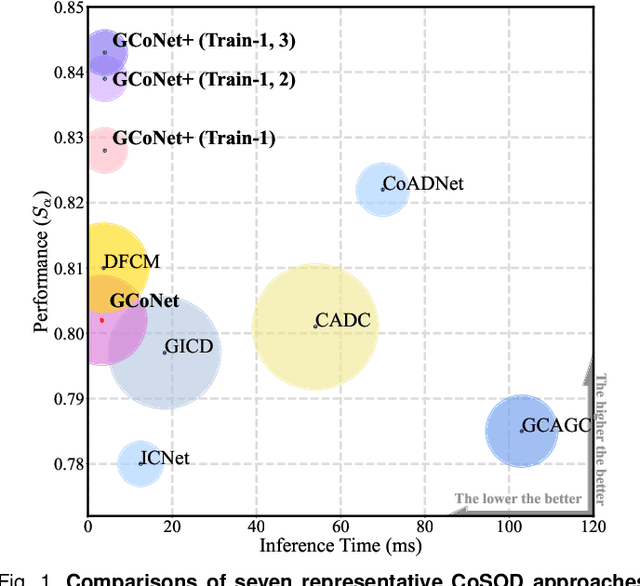
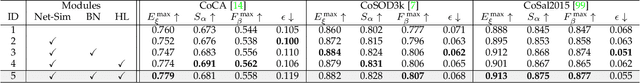

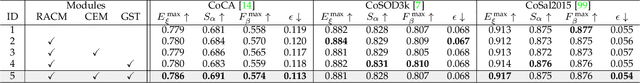
Abstract:In this paper, we present a novel end-to-end group collaborative learning network, termed GCoNet+, which can effectively and efficiently (250 fps) identify co-salient objects in natural scenes. The proposed GCoNet+ achieves the new state-of-the-art performance for co-salient object detection (CoSOD) through mining consensus representations based on the following two essential criteria: 1) intra-group compactness to better formulate the consistency among co-salient objects by capturing their inherent shared attributes using our novel group affinity module (GAM); 2) inter-group separability to effectively suppress the influence of noisy objects on the output by introducing our new group collaborating module (GCM) conditioning on the inconsistent consensus. To further improve the accuracy, we design a series of simple yet effective components as follows: i) a recurrent auxiliary classification module (RACM) promoting the model learning at the semantic level; ii) a confidence enhancement module (CEM) helping the model to improve the quality of the final predictions; and iii) a group-based symmetric triplet (GST) loss guiding the model to learn more discriminative features. Extensive experiments on three challenging benchmarks, i.e., CoCA, CoSOD3k, and CoSal2015, demonstrate that our GCoNet+ outperforms the existing 12 cutting-edge models. Code has been released at https://github.com/ZhengPeng7/GCoNet_plus.
Progressive Multi-scale Consistent Network for Multi-class Fundus Lesion Segmentation
May 31, 2022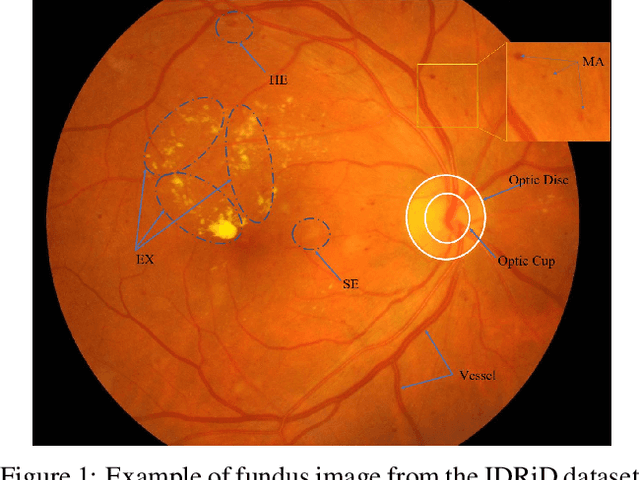
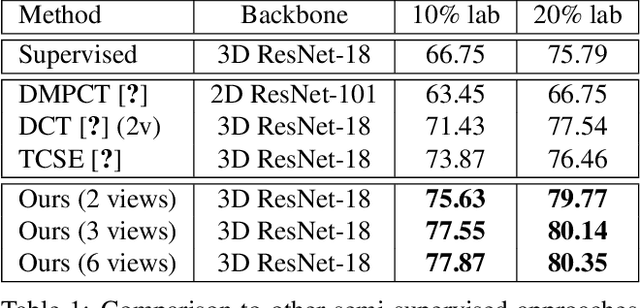
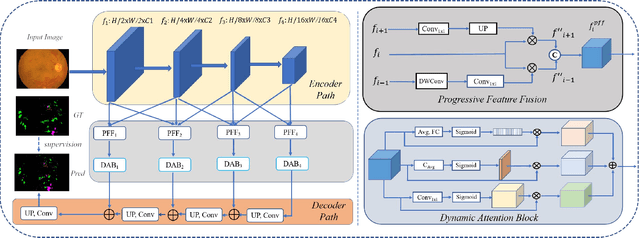
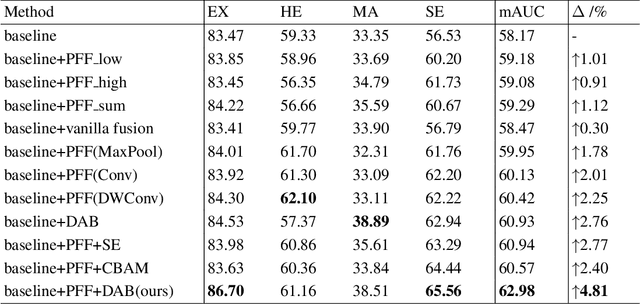
Abstract:Effectively integrating multi-scale information is of considerable significance for the challenging multi-class segmentation of fundus lesions because different lesions vary significantly in scales and shapes. Several methods have been proposed to successfully handle the multi-scale object segmentation. However, two issues are not considered in previous studies. The first is the lack of interaction between adjacent feature levels, and this will lead to the deviation of high-level features from low-level features and the loss of detailed cues. The second is the conflict between the low-level and high-level features, this occurs because they learn different scales of features, thereby confusing the model and decreasing the accuracy of the final prediction. In this paper, we propose a progressive multi-scale consistent network (PMCNet) that integrates the proposed progressive feature fusion (PFF) block and dynamic attention block (DAB) to address the aforementioned issues. Specifically, PFF block progressively integrates multi-scale features from adjacent encoding layers, facilitating feature learning of each layer by aggregating fine-grained details and high-level semantics. As features at different scales should be consistent, DAB is designed to dynamically learn the attentive cues from the fused features at different scales, thus aiming to smooth the essential conflicts existing in multi-scale features. The two proposed PFF and DAB blocks can be integrated with the off-the-shelf backbone networks to address the two issues of multi-scale and feature inconsistency in the multi-class segmentation of fundus lesions, which will produce better feature representation in the feature space. Experimental results on three public datasets indicate that the proposed method is more effective than recent state-of-the-art methods.
Structure Unbiased Adversarial Model for Medical Image Segmentation
May 26, 2022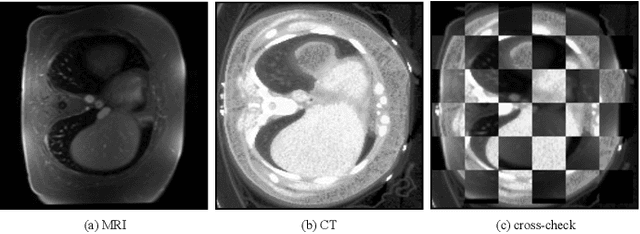
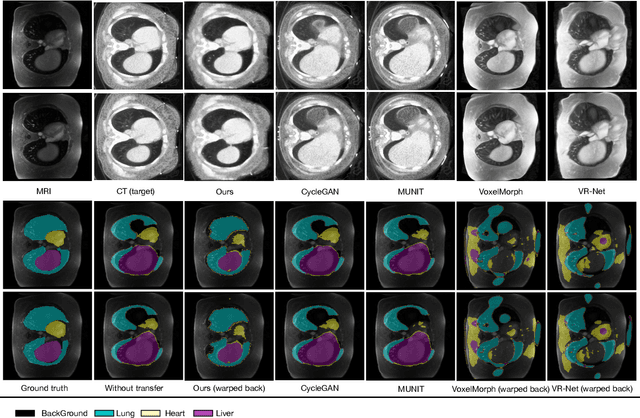
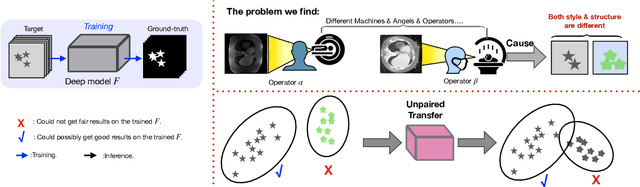
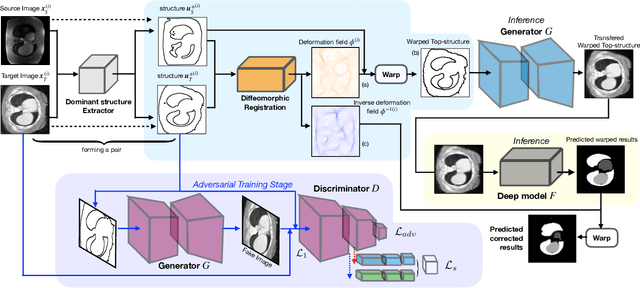
Abstract:Generative models have been widely proposed in image recognition to generate more images where the distribution is similar to that of the real images. It often introduces a discriminator network to discriminate original real data and generated data. However, such discriminator often considers the distribution of the data and did not pay enough attention to the intrinsic gap due to structure. In this paper, we reformulate a new image to image translation problem to reduce structural gap, in addition to the typical intensity distribution gap. We further propose a simple yet important Structure Unbiased Adversarial Model for Medical Image Segmentation (SUAM) with learnable inverse structural deformation for medical image segmentation. It consists of a structure extractor, an attention diffeomorphic registration and a structure \& intensity distribution rendering module. The structure extractor aims to extract the dominant structure of the input image. The attention diffeomorphic registration is proposed to reduce the structure gap with an inverse deformation field to warp the prediction masks back to their original form. The structure rendering module is to render the deformed structure to an image with targeted intensity distribution. We apply the proposed SUAM on both optical coherence tomography (OCT), magnetic resonance imaging (MRI) and computerized tomography (CT) data. Experimental results show that the proposed method has the capability to transfer both intensity and structure distributions.
Trusted Multi-View Classification with Dynamic Evidential Fusion
Apr 27, 2022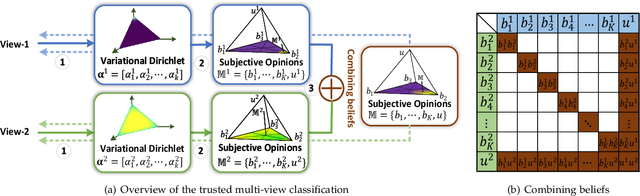
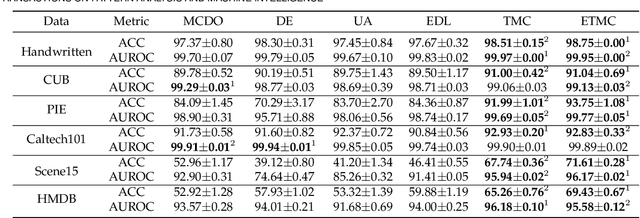
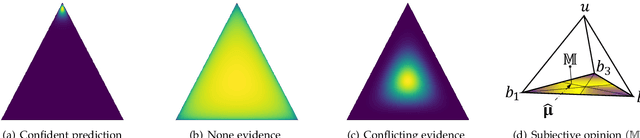
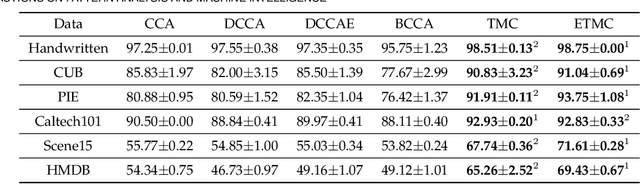
Abstract:Existing multi-view classification algorithms focus on promoting accuracy by exploiting different views, typically integrating them into common representations for follow-up tasks. Although effective, it is also crucial to ensure the reliability of both the multi-view integration and the final decision, especially for noisy, corrupted and out-of-distribution data. Dynamically assessing the trustworthiness of each view for different samples could provide reliable integration. This can be achieved through uncertainty estimation. With this in mind, we propose a novel multi-view classification algorithm, termed trusted multi-view classification (TMC), providing a new paradigm for multi-view learning by dynamically integrating different views at an evidence level. The proposed TMC can promote classification reliability by considering evidence from each view. Specifically, we introduce the variational Dirichlet to characterize the distribution of the class probabilities, parameterized with evidence from different views and integrated with the Dempster-Shafer theory. The unified learning framework induces accurate uncertainty and accordingly endows the model with both reliability and robustness against possible noise or corruption. Both theoretical and experimental results validate the effectiveness of the proposed model in accuracy, robustness and trustworthiness.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge