Fengtao Zhou
A Decade-Scale Benchmark Evaluating LLMs' Clinical Practice Guidelines Detection and Adherence in Multi-turn Conversations
Mar 26, 2026Abstract:Clinical practice guidelines (CPGs) play a pivotal role in ensuring evidence-based decision-making and improving patient outcomes. While Large Language Models (LLMs) are increasingly deployed in healthcare scenarios, it is unclear to which extend LLMs could identify and adhere to CPGs during conversations. To address this gap, we introduce CPGBench, an automated framework benchmarking the clinical guideline detection and adherence capabilities of LLMs in multi-turn conversations. We collect 3,418 CPG documents from 9 countries/regions and 2 international organizations published in the last decade spanning across 24 specialties. From these documents, we extract 32,155 clinical recommendations with corresponding publication institute, date, country, specialty, recommendation strength, evidence level, etc. One multi-turn conversation is generated for each recommendation accordingly to evaluate the detection and adherence capabilities of 8 leading LLMs. We find that the 71.1%-89.6% recommendations can be correctly detected, while only 3.6%-29.7% corresponding titles can be correctly referenced, revealing the gap between knowing the guideline contents and where they come from. The adherence rates range from 21.8% to 63.2% in different models, indicating a large gap between knowing the guidelines and being able to apply them. To confirm the validity of our automatic analysis, we further conduct a comprehensive human evaluation involving 56 clinicians from different specialties. To our knowledge, CPGBench is the first benchmark systematically revealing which clinical recommendations LLMs fail to detect or adhere to during conversations. Given that each clinical recommendation may affect a large population and that clinical applications are inherently safety critical, addressing these gaps is crucial for the safe and responsible deployment of LLMs in real world clinical practice.
A Deployment-Friendly Foundational Framework for Efficient Computational Pathology
Feb 15, 2026Abstract:Pathology foundation models (PFMs) have enabled robust generalization in computational pathology through large-scale datasets and expansive architectures, but their substantial computational cost, particularly for gigapixel whole slide images, limits clinical accessibility and scalability. Here, we present LitePath, a deployment-friendly foundational framework designed to mitigate model over-parameterization and patch level redundancy. LitePath integrates LiteFM, a compact model distilled from three large PFMs (Virchow2, H-Optimus-1 and UNI2) using 190 million patches, and the Adaptive Patch Selector (APS), a lightweight component for task-specific patch selection. The framework reduces model parameters by 28x and lowers FLOPs by 403.5x relative to Virchow2, enabling deployment on low-power edge hardware such as the NVIDIA Jetson Orin Nano Super. On this device, LitePath processes 208 slides per hour, 104.5x faster than Virchow2, and consumes 0.36 kWh per 3,000 slides, 171x lower than Virchow2 on an RTX3090 GPU. We validated accuracy using 37 cohorts across four organs and 26 tasks (26 internal, 9 external, and 2 prospective), comprising 15,672 slides from 9,808 patients disjoint from the pretraining data. LitePath ranks second among 19 evaluated models and outperforms larger models including H-Optimus-1, mSTAR, UNI2 and GPFM, while retaining 99.71% of the AUC of Virchow2 on average. To quantify the balance between accuracy and efficiency, we propose the Deployability Score (D-Score), defined as the weighted geometric mean of normalized AUC and normalized FLOP, where LitePath achieves the highest value, surpassing Virchow2 by 10.64%. These results demonstrate that LitePath enables rapid, cost-effective and energy-efficient pathology image analysis on accessible hardware while maintaining accuracy comparable to state-of-the-art PFMs and reducing the carbon footprint of AI deployment.
MambaMIL+: Modeling Long-Term Contextual Patterns for Gigapixel Whole Slide Image
Dec 19, 2025



Abstract:Whole-slide images (WSIs) are an important data modality in computational pathology, yet their gigapixel resolution and lack of fine-grained annotations challenge conventional deep learning models. Multiple instance learning (MIL) offers a solution by treating each WSI as a bag of patch-level instances, but effectively modeling ultra-long sequences with rich spatial context remains difficult. Recently, Mamba has emerged as a promising alternative for long sequence learning, scaling linearly to thousands of tokens. However, despite its efficiency, it still suffers from limited spatial context modeling and memory decay, constraining its effectiveness to WSI analysis. To address these limitations, we propose MambaMIL+, a new MIL framework that explicitly integrates spatial context while maintaining long-range dependency modeling without memory forgetting. Specifically, MambaMIL+ introduces 1) overlapping scanning, which restructures the patch sequence to embed spatial continuity and instance correlations; 2) a selective stripe position encoder (S2PE) that encodes positional information while mitigating the biases of fixed scanning orders; and 3) a contextual token selection (CTS) mechanism, which leverages supervisory knowledge to dynamically enlarge the contextual memory for stable long-range modeling. Extensive experiments on 20 benchmarks across diagnostic classification, molecular prediction, and survival analysis demonstrate that MambaMIL+ consistently achieves state-of-the-art performance under three feature extractors (ResNet-50, PLIP, and CONCH), highlighting its effectiveness and robustness for large-scale computational pathology
LLM-driven Knowledge Enhancement for Multimodal Cancer Survival Prediction
Dec 16, 2025Abstract:Current multimodal survival prediction methods typically rely on pathology images (WSIs) and genomic data, both of which are high-dimensional and redundant, making it difficult to extract discriminative features from them and align different modalities. Moreover, using a simple survival follow-up label is insufficient to supervise such a complex task. To address these challenges, we propose KEMM, an LLM-driven Knowledge-Enhanced Multimodal Model for cancer survival prediction, which integrates expert reports and prognostic background knowledge. 1) Expert reports, provided by pathologists on a case-by-case basis and refined by large language model (LLM), offer succinct and clinically focused diagnostic statements. This information may typically suggest different survival outcomes. 2) Prognostic background knowledge (PBK), generated concisely by LLM, provides valuable prognostic background knowledge on different cancer types, which also enhances survival prediction. To leverage these knowledge, we introduce the knowledge-enhanced cross-modal (KECM) attention module. KECM can effectively guide the network to focus on discriminative and survival-relevant features from highly redundant modalities. Extensive experiments on five datasets demonstrate that KEMM achieves state-of-the-art performance. The code will be released upon acceptance.
A Clinical-grade Universal Foundation Model for Intraoperative Pathology
Oct 06, 2025Abstract:Intraoperative pathology is pivotal to precision surgery, yet its clinical impact is constrained by diagnostic complexity and the limited availability of high-quality frozen-section data. While computational pathology has made significant strides, the lack of large-scale, prospective validation has impeded its routine adoption in surgical workflows. Here, we introduce CRISP, a clinical-grade foundation model developed on over 100,000 frozen sections from eight medical centers, specifically designed to provide Clinical-grade Robust Intraoperative Support for Pathology (CRISP). CRISP was comprehensively evaluated on more than 15,000 intraoperative slides across nearly 100 retrospective diagnostic tasks, including benign-malignant discrimination, key intraoperative decision-making, and pan-cancer detection, etc. The model demonstrated robust generalization across diverse institutions, tumor types, and anatomical sites-including previously unseen sites and rare cancers. In a prospective cohort of over 2,000 patients, CRISP sustained high diagnostic accuracy under real-world conditions, directly informing surgical decisions in 92.6% of cases. Human-AI collaboration further reduced diagnostic workload by 35%, avoided 105 ancillary tests and enhanced detection of micrometastases with 87.5% accuracy. Together, these findings position CRISP as a clinical-grade paradigm for AI-driven intraoperative pathology, bridging computational advances with surgical precision and accelerating the translation of artificial intelligence into routine clinical practice.
Generative AI for Misalignment-Resistant Virtual Staining to Accelerate Histopathology Workflows
Sep 17, 2025Abstract:Accurate histopathological diagnosis often requires multiple differently stained tissue sections, a process that is time-consuming, labor-intensive, and environmentally taxing due to the use of multiple chemical stains. Recently, virtual staining has emerged as a promising alternative that is faster, tissue-conserving, and environmentally friendly. However, existing virtual staining methods face significant challenges in clinical applications, primarily due to their reliance on well-aligned paired data. Obtaining such data is inherently difficult because chemical staining processes can distort tissue structures, and a single tissue section cannot undergo multiple staining procedures without damage or loss of information. As a result, most available virtual staining datasets are either unpaired or roughly paired, making it difficult for existing methods to achieve accurate pixel-level supervision. To address this challenge, we propose a robust virtual staining framework featuring cascaded registration mechanisms to resolve spatial mismatches between generated outputs and their corresponding ground truth. Experimental results demonstrate that our method significantly outperforms state-of-the-art models across five datasets, achieving an average improvement of 3.2% on internal datasets and 10.1% on external datasets. Moreover, in datasets with substantial misalignment, our approach achieves a remarkable 23.8% improvement in peak signal-to-noise ratio compared to baseline models. The exceptional robustness of the proposed method across diverse datasets simplifies the data acquisition process for virtual staining and offers new insights for advancing its development.
A Multimodal Foundation Model to Enhance Generalizability and Data Efficiency for Pan-cancer Prognosis Prediction
Sep 16, 2025Abstract:Multimodal data provides heterogeneous information for a holistic understanding of the tumor microenvironment. However, existing AI models often struggle to harness the rich information within multimodal data and extract poorly generalizable representations. Here we present MICE (Multimodal data Integration via Collaborative Experts), a multimodal foundation model that effectively integrates pathology images, clinical reports, and genomics data for precise pan-cancer prognosis prediction. Instead of conventional multi-expert modules, MICE employs multiple functionally diverse experts to comprehensively capture both cross-cancer and cancer-specific insights. Leveraging data from 11,799 patients across 30 cancer types, we enhanced MICE's generalizability by coupling contrastive and supervised learning. MICE outperformed both unimodal and state-of-the-art multi-expert-based multimodal models, demonstrating substantial improvements in C-index ranging from 3.8% to 11.2% on internal cohorts and 5.8% to 8.8% on independent cohorts, respectively. Moreover, it exhibited remarkable data efficiency across diverse clinical scenarios. With its enhanced generalizability and data efficiency, MICE establishes an effective and scalable foundation for pan-cancer prognosis prediction, holding strong potential to personalize tailored therapies and improve treatment outcomes.
A Versatile Pathology Co-pilot via Reasoning Enhanced Multimodal Large Language Model
Jul 23, 2025



Abstract:Multimodal large language models (MLLMs) have emerged as powerful tools for computational pathology, offering unprecedented opportunities to integrate pathological images with language context for comprehensive diagnostic analysis. These models hold particular promise for automating complex tasks that traditionally require expert interpretation of pathologists. However, current MLLM approaches in pathology demonstrate significantly constrained reasoning capabilities, primarily due to their reliance on expensive chain-of-thought annotations. Additionally, existing methods remain limited to simplex application of visual question answering (VQA) at region-of-interest (ROI) level, failing to address the full spectrum of diagnostic needs such as ROI classification, detection, segmentation, whole-slide-image (WSI) classification and VQA in clinical practice. In this study, we present SmartPath-R1, a versatile MLLM capable of simultaneously addressing both ROI-level and WSI-level tasks while demonstrating robust pathological reasoning capability. Our framework combines scale-dependent supervised fine-tuning and task-aware reinforcement fine-tuning, which circumvents the requirement for chain-of-thought supervision by leveraging the intrinsic knowledge within MLLM. Furthermore, SmartPath-R1 integrates multiscale and multitask analysis through a mixture-of-experts mechanism, enabling dynamic processing for diverse tasks. We curate a large-scale dataset comprising 2.3M ROI samples and 188K WSI samples for training and evaluation. Extensive experiments across 72 tasks validate the effectiveness and superiority of the proposed approach. This work represents a significant step toward developing versatile, reasoning-enhanced AI systems for precision pathology.
Genome-Anchored Foundation Model Embeddings Improve Molecular Prediction from Histology Images
Jun 24, 2025

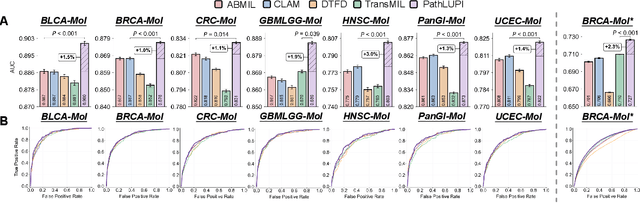

Abstract:Precision oncology requires accurate molecular insights, yet obtaining these directly from genomics is costly and time-consuming for broad clinical use. Predicting complex molecular features and patient prognosis directly from routine whole-slide images (WSI) remains a major challenge for current deep learning methods. Here we introduce PathLUPI, which uses transcriptomic privileged information during training to extract genome-anchored histological embeddings, enabling effective molecular prediction using only WSIs at inference. Through extensive evaluation across 49 molecular oncology tasks using 11,257 cases among 20 cohorts, PathLUPI demonstrated superior performance compared to conventional methods trained solely on WSIs. Crucially, it achieves AUC $\geq$ 0.80 in 14 of the biomarker prediction and molecular subtyping tasks and C-index $\geq$ 0.70 in survival cohorts of 5 major cancer types. Moreover, PathLUPI embeddings reveal distinct cellular morphological signatures associated with specific genotypes and related biological pathways within WSIs. By effectively encoding molecular context to refine WSI representations, PathLUPI overcomes a key limitation of existing models and offers a novel strategy to bridge molecular insights with routine pathology workflows for wider clinical application.
PathBench: A comprehensive comparison benchmark for pathology foundation models towards precision oncology
May 26, 2025



Abstract:The emergence of pathology foundation models has revolutionized computational histopathology, enabling highly accurate, generalized whole-slide image analysis for improved cancer diagnosis, and prognosis assessment. While these models show remarkable potential across cancer diagnostics and prognostics, their clinical translation faces critical challenges including variability in optimal model across cancer types, potential data leakage in evaluation, and lack of standardized benchmarks. Without rigorous, unbiased evaluation, even the most advanced PFMs risk remaining confined to research settings, delaying their life-saving applications. Existing benchmarking efforts remain limited by narrow cancer-type focus, potential pretraining data overlaps, or incomplete task coverage. We present PathBench, the first comprehensive benchmark addressing these gaps through: multi-center in-hourse datasets spanning common cancers with rigorous leakage prevention, evaluation across the full clinical spectrum from diagnosis to prognosis, and an automated leaderboard system for continuous model assessment. Our framework incorporates large-scale data, enabling objective comparison of PFMs while reflecting real-world clinical complexity. All evaluation data comes from private medical providers, with strict exclusion of any pretraining usage to avoid data leakage risks. We have collected 15,888 WSIs from 8,549 patients across 10 hospitals, encompassing over 64 diagnosis and prognosis tasks. Currently, our evaluation of 19 PFMs shows that Virchow2 and H-Optimus-1 are the most effective models overall. This work provides researchers with a robust platform for model development and offers clinicians actionable insights into PFM performance across diverse clinical scenarios, ultimately accelerating the translation of these transformative technologies into routine pathology practice.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge