Deep Learning of Potential Outcomes
Oct 09, 2021Bernard Koch, Tim Sainburg, Pablo Geraldo, Song Jiang, Yizhou Sun, Jacob Gates Foster
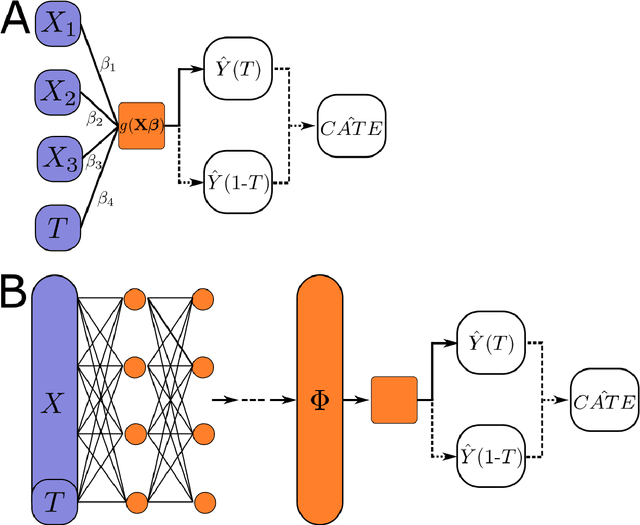
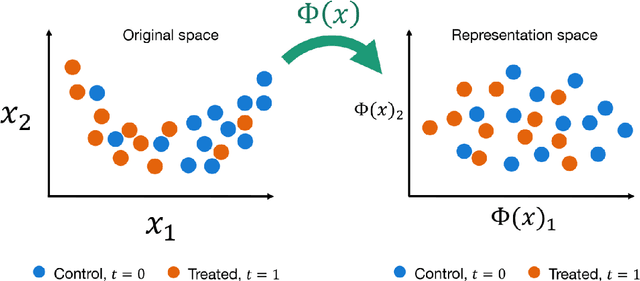
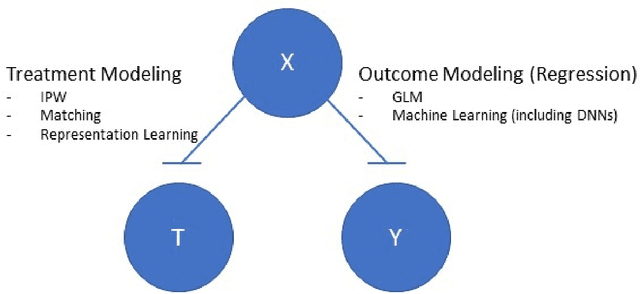
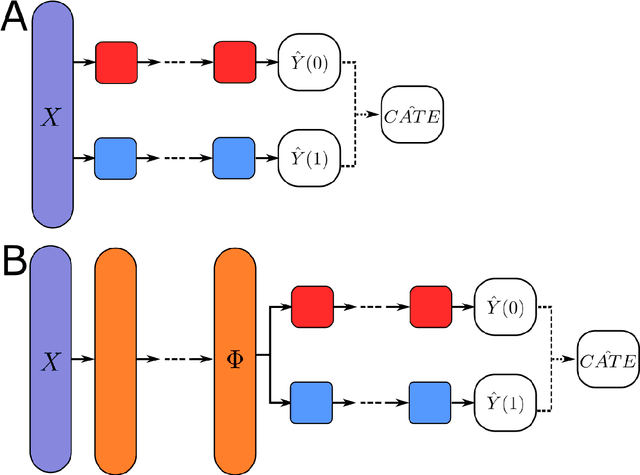
This review systematizes the emerging literature for causal inference using deep neural networks under the potential outcomes framework. It provides an intuitive introduction on how deep learning can be used to estimate/predict heterogeneous treatment effects and extend causal inference to settings where confounding is non-linear, time varying, or encoded in text, networks, and images. To maximize accessibility, we also introduce prerequisite concepts from causal inference and deep learning. The survey differs from other treatments of deep learning and causal inference in its sharp focus on observational causal estimation, its extended exposition of key algorithms, and its detailed tutorials for implementing, training, and selecting among deep estimators in Tensorflow 2 available at github.com/kochbj/Deep-Learning-for-Causal-Inference.
Relation-Guided Pre-Training for Open-Domain Question Answering
Sep 21, 2021Ziniu Hu, Yizhou Sun, Kai-Wei Chang
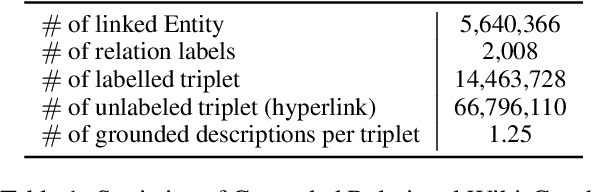
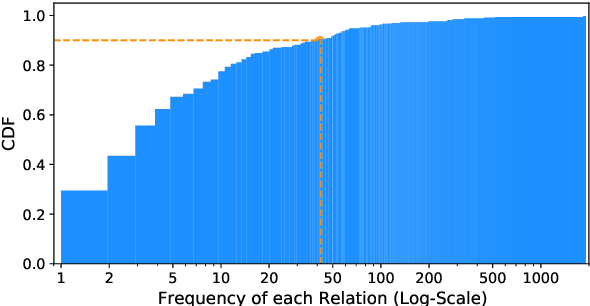

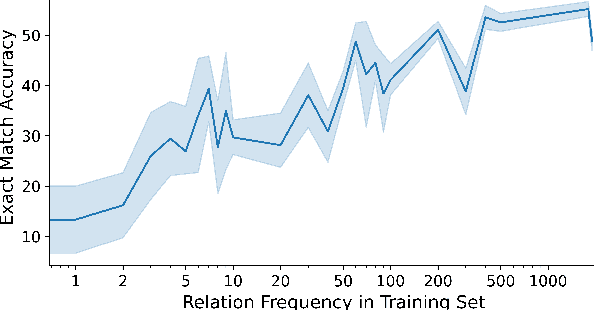
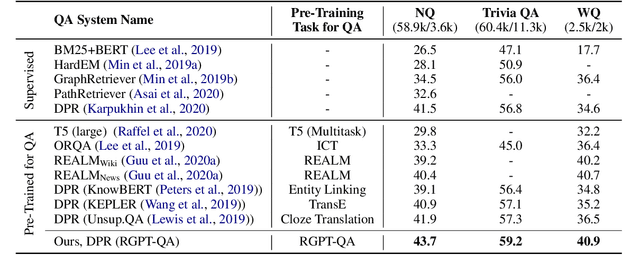
Answering complex open-domain questions requires understanding the latent relations between involving entities. However, we found that the existing QA datasets are extremely imbalanced in some types of relations, which hurts the generalization performance over questions with long-tail relations. To remedy this problem, in this paper, we propose a Relation-Guided Pre-Training (RGPT-QA) framework. We first generate a relational QA dataset covering a wide range of relations from both the Wikidata triplets and Wikipedia hyperlinks. We then pre-train a QA model to infer the latent relations from the question, and then conduct extractive QA to get the target answer entity. We demonstrate that by pretraining with propoed RGPT-QA techique, the popular open-domain QA model, Dense Passage Retriever (DPR), achieves 2.2%, 2.4%, and 6.3% absolute improvement in Exact Match accuracy on Natural Questions, TriviaQA, and WebQuestions. Particularly, we show that RGPT-QA improves significantly on questions with long-tail relations
Fuzzy Logic based Logical Query Answering on Knowledge Graph
Aug 05, 2021Xuelu Chen, Ziniu Hu, Yizhou Sun
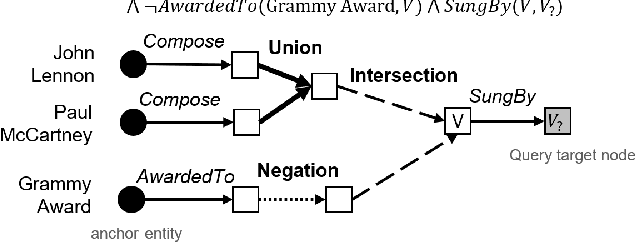

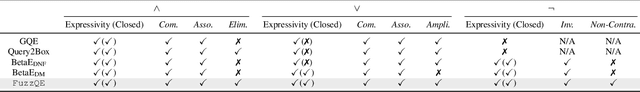
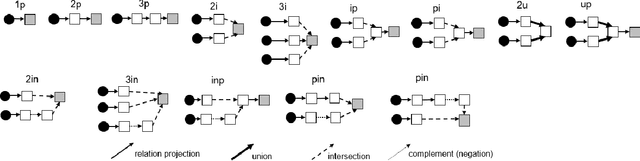
Answering complex First-Order Logical (FOL) queries on large-scale incomplete knowledge graphs (KGs) is an important yet challenging task. Recent advances embed logical queries and KG entities in the vector space and conduct query answering via dense similarity search. However, most of the designed logical operators in existing works do not satisfy the axiomatic system of classical logic. Moreover, these logical operators are parameterized so that they require a large number of complex FOL queries as training data, which are often arduous or even inaccessible to collect in most real-world KGs. In this paper, we present FuzzQE, a fuzzy logic based query embedding framework for answering FOL queries over KGs. FuzzQE follows fuzzy logic to define logical operators in a principled and learning free manner. Extensive experiments on two benchmark datasets demonstrate that FuzzQE achieves significantly better performance in answering FOL queries compared to the state-of-the-art methods. In addition, FuzzQE trained with only KG link prediction without any complex queries can achieve comparable performance with the systems trained with all FOL queries.
Clinical Named Entity Recognition using Contextualized Token Representations
Jun 23, 2021Yichao Zhou, Chelsea Ju, J. Harry Caufield, Kevin Shih, Calvin Chen, Yizhou Sun, Kai-Wei Chang, Peipei Ping, Wei Wang
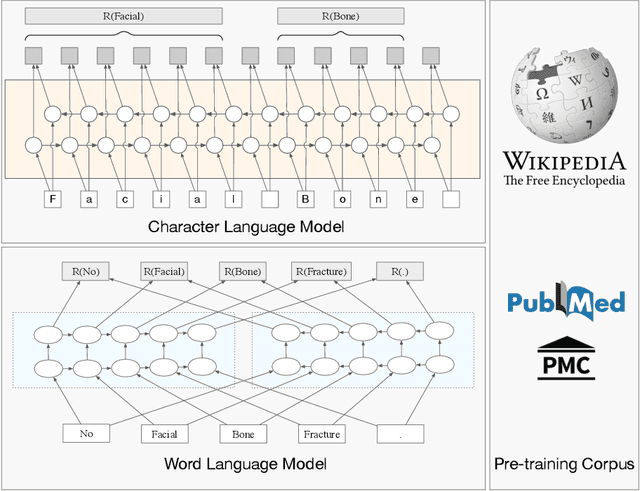
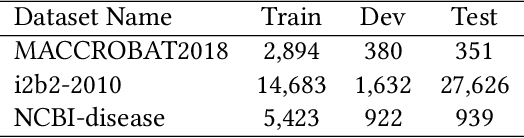
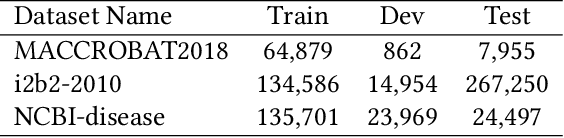

The clinical named entity recognition (CNER) task seeks to locate and classify clinical terminologies into predefined categories, such as diagnostic procedure, disease disorder, severity, medication, medication dosage, and sign symptom. CNER facilitates the study of side-effect on medications including identification of novel phenomena and human-focused information extraction. Existing approaches in extracting the entities of interests focus on using static word embeddings to represent each word. However, one word can have different interpretations that depend on the context of the sentences. Evidently, static word embeddings are insufficient to integrate the diverse interpretation of a word. To overcome this challenge, the technique of contextualized word embedding has been introduced to better capture the semantic meaning of each word based on its context. Two of these language models, ELMo and Flair, have been widely used in the field of Natural Language Processing to generate the contextualized word embeddings on domain-generic documents. However, these embeddings are usually too general to capture the proximity among vocabularies of specific domains. To facilitate various downstream applications using clinical case reports (CCRs), we pre-train two deep contextualized language models, Clinical Embeddings from Language Model (C-ELMo) and Clinical Contextual String Embeddings (C-Flair) using the clinical-related corpus from the PubMed Central. Explicit experiments show that our models gain dramatic improvements compared to both static word embeddings and domain-generic language models.
Universal Representation Learning of Knowledge Bases by Jointly Embedding Instances and Ontological Concepts
Mar 15, 2021Junheng Hao, Muhao Chen, Wenchao Yu, Yizhou Sun, Wei Wang
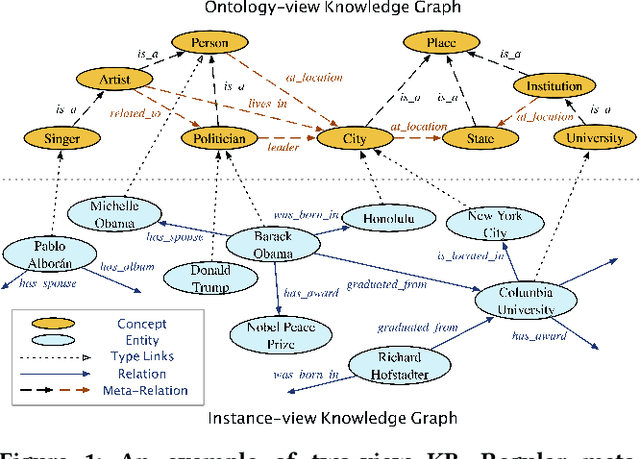
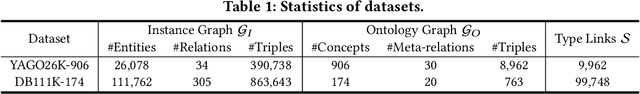
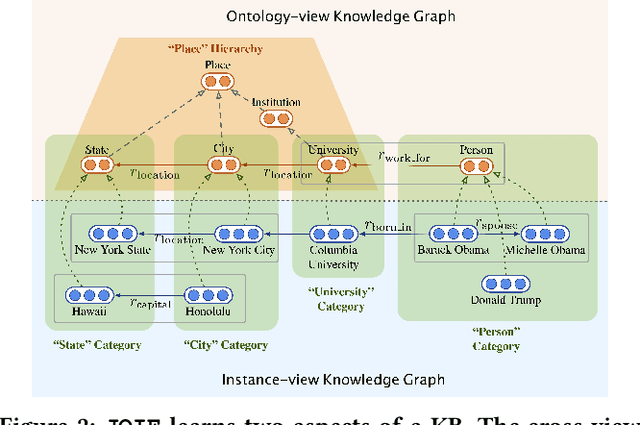

Many large-scale knowledge bases simultaneously represent two views of knowledge graphs (KGs): an ontology view for abstract and commonsense concepts, and an instance view for specific entities that are instantiated from ontological concepts. Existing KG embedding models, however, merely focus on representing one of the two views alone. In this paper, we propose a novel two-view KG embedding model, JOIE, with the goal to produce better knowledge embedding and enable new applications that rely on multi-view knowledge. JOIE employs both cross-view and intra-view modeling that learn on multiple facets of the knowledge base. The cross-view association model is learned to bridge the embeddings of ontological concepts and their corresponding instance-view entities. The intra-view models are trained to capture the structured knowledge of instance and ontology views in separate embedding spaces, with a hierarchy-aware encoding technique enabled for ontologies with hierarchies. We explore multiple representation techniques for the two model components and investigate with nine variants of JOIE. Our model is trained on large-scale knowledge bases that consist of massive instances and their corresponding ontological concepts connected via a (small) set of cross-view links. Experimental results on public datasets show that the best variant of JOIE significantly outperforms previous models on instance-view triple prediction task as well as ontology population on ontologyview KG. In addition, our model successfully extends the use of KG embeddings to entity typing with promising performance.
Bio-JOIE: Joint Representation Learning of Biological Knowledge Bases
Mar 07, 2021Junheng Hao, Chelsea Ju, Muhao Chen, Yizhou Sun, Carlo Zaniolo, Wei Wang
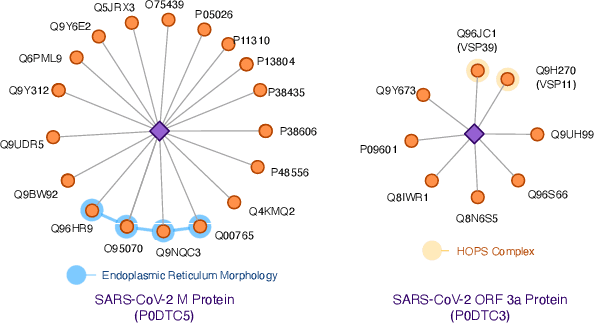
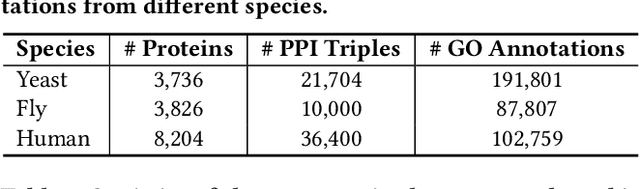
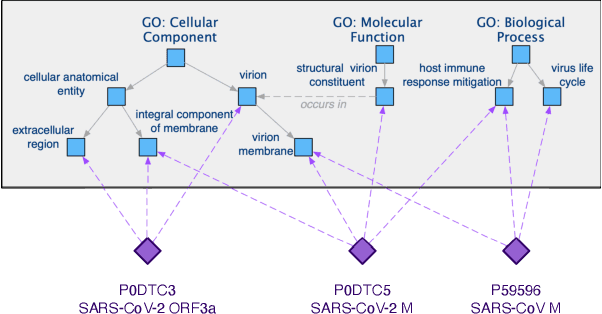
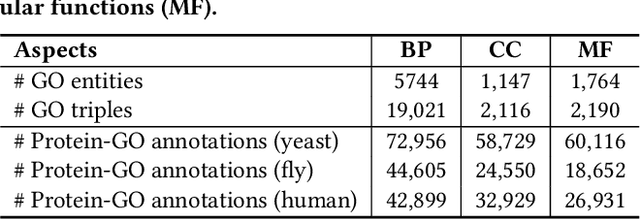
The widespread of Coronavirus has led to a worldwide pandemic with a high mortality rate. Currently, the knowledge accumulated from different studies about this virus is very limited. Leveraging a wide-range of biological knowledge, such as gene ontology and protein-protein interaction (PPI) networks from other closely related species presents a vital approach to infer the molecular impact of a new species. In this paper, we propose the transferred multi-relational embedding model Bio-JOIE to capture the knowledge of gene ontology and PPI networks, which demonstrates superb capability in modeling the SARS-CoV-2-human protein interactions. Bio-JOIE jointly trains two model components. The knowledge model encodes the relational facts from the protein and GO domains into separated embedding spaces, using a hierarchy-aware encoding technique employed for the GO terms. On top of that, the transfer model learns a non-linear transformation to transfer the knowledge of PPIs and gene ontology annotations across their embedding spaces. By leveraging only structured knowledge, Bio-JOIE significantly outperforms existing state-of-the-art methods in PPI type prediction on multiple species. Furthermore, we also demonstrate the potential of leveraging the learned representations on clustering proteins with enzymatic function into enzyme commission families. Finally, we show that Bio-JOIE can accurately identify PPIs between the SARS-CoV-2 proteins and human proteins, providing valuable insights for advancing research on this new disease.
* ACM BCB 2020, Best Student Paper
CREATe: Clinical Report Extraction and Annotation Technology
Feb 28, 2021Yichao Zhou, Wei-Ting Chen, Bowen Zhang, David Lee, J. Harry Caufield, Kai-Wei Chang, Yizhou Sun, Peipei Ping, Wei Wang
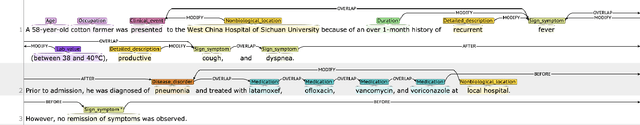
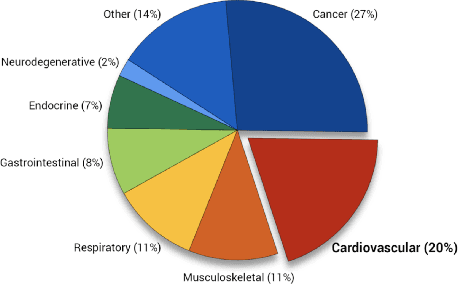
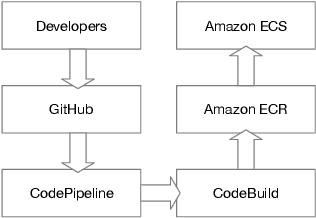
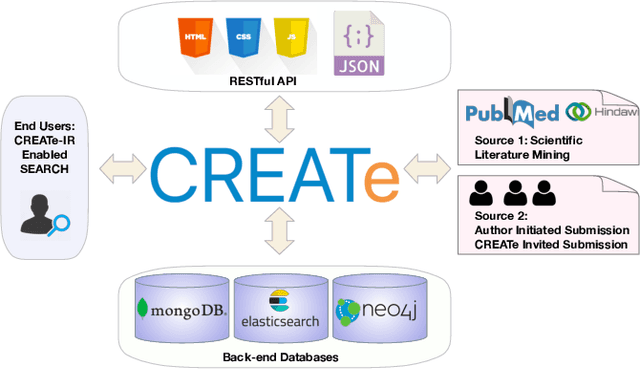

Clinical case reports are written descriptions of the unique aspects of a particular clinical case, playing an essential role in sharing clinical experiences about atypical disease phenotypes and new therapies. However, to our knowledge, there has been no attempt to develop an end-to-end system to annotate, index, or otherwise curate these reports. In this paper, we propose a novel computational resource platform, CREATe, for extracting, indexing, and querying the contents of clinical case reports. CREATe fosters an environment of sustainable resource support and discovery, enabling researchers to overcome the challenges of information science. An online video of the demonstration can be viewed at https://youtu.be/Q8owBQYTjDc.
Motif-Driven Contrastive Learning of Graph Representations
Dec 23, 2020Shichang Zhang, Ziniu Hu, Arjun Subramonian, Yizhou Sun
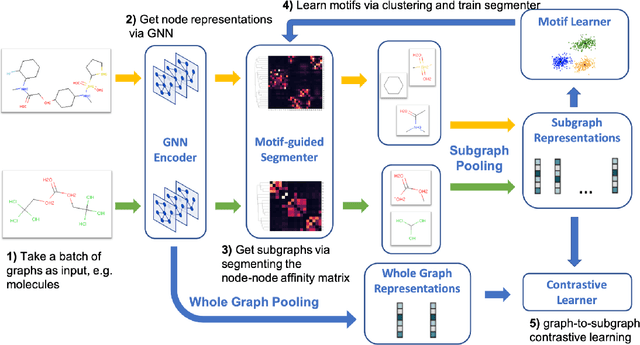
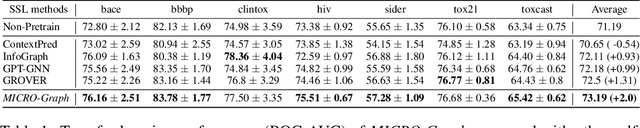
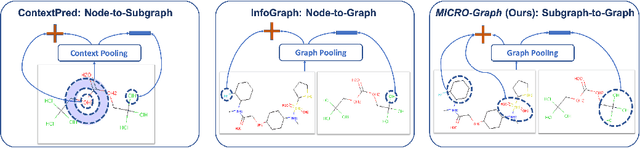
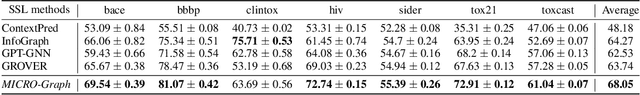
Graph motifs are significant subgraph patterns occurring frequently in graphs, and they play important roles in representing the whole graph characteristics. For example, in chemical domain, functional groups are motifs that can determine molecule properties. Mining and utilizing motifs, however, is a non-trivial task for large graph datasets. Traditional motif discovery approaches rely on exact counting or statistical estimation, which are hard to scale for large datasets with continuous and high-dimension features. In light of the significance and challenges of motif mining, we propose MICRO-Graph: a framework for MotIf-driven Contrastive leaRning Of Graph representations to: 1) pre-train Graph Neural Net-works (GNNs) in a self-supervised manner to automatically extract motifs from large graph datasets; 2) leverage learned motifs to guide the contrastive learning of graph representations, which further benefit various downstream tasks. Specifically, given a graph dataset, a motif learner cluster similar and significant subgraphs into corresponding motif slots. Based on the learned motifs, a motif-guided subgraph segmenter is trained to generate more informative subgraphs, which are used to conduct graph-to-subgraph contrastive learning of GNNs. By pre-training on ogbg-molhiv molecule dataset with our proposed MICRO-Graph, the pre-trained GNN model can enhance various chemical property prediction down-stream tasks with scarce label by 2.0%, which is significantly higher than other state-of-the-art self-supervised learning baselines.
Leveraging Meta-path Contexts for Classification in Heterogeneous Information Networks
Dec 18, 2020Xiang Li, Danhao Ding, Ben Kao, Yizhou Sun, Nikos Mamoulis
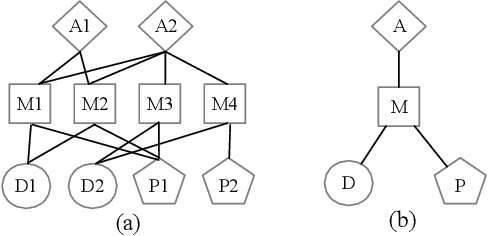
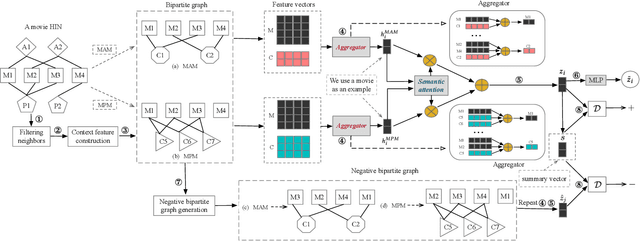
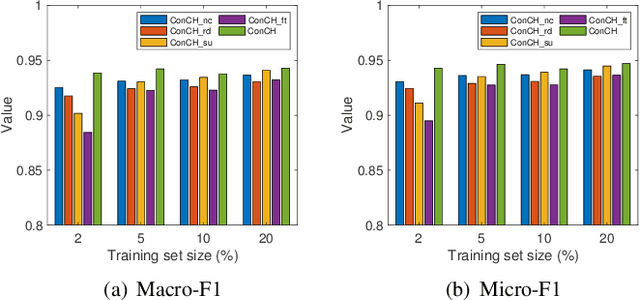
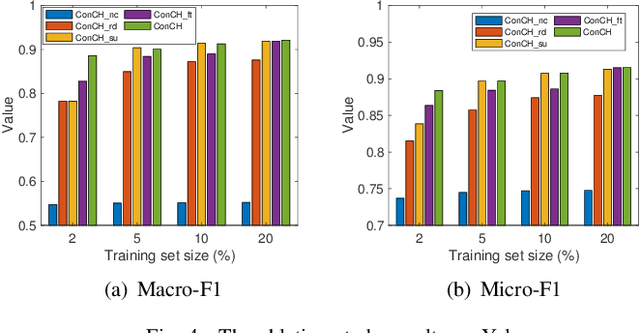
A heterogeneous information network (HIN) has as vertices objects of different types and as edges the relations between objects, which are also of various types. We study the problem of classifying objects in HINs. Most existing methods perform poorly when given scarce labeled objects as training sets, and methods that improve classification accuracy under such scenarios are often computationally expensive. To address these problems, we propose ConCH, a graph neural network model. ConCH formulates the classification problem as a multi-task learning problem that combines semi-supervised learning with self-supervised learning to learn from both labeled and unlabeled data. ConCH employs meta-paths, which are sequences of object types that capture semantic relationships between objects. Based on meta-paths, it considers two sources of information for an object x: (1) Meta-path-based neighbors of x are retrieved and ranked, and the top-k neighbors are retained. (2) The meta-path instances of x to its selected neighbors are used to derive meta-path-based contexts. ConCH utilizes the above information to co-derive object embeddings and context embeddings via graph convolution. It also uses the attention mechanism to fuse the embeddings of x generated from various meta-paths to obtain x's final embedding. We conduct extensive experiments to evaluate the performance of ConCH against other 14 classification methods. Our results show that ConCH is an effective and efficient method for HIN classification.
Clinical Temporal Relation Extraction with Probabilistic Soft Logic Regularization and Global Inference
Dec 16, 2020Yichao Zhou, Yu Yan, Rujun Han, J. Harry Caufield, Kai-Wei Chang, Yizhou Sun, Peipei Ping, Wei Wang
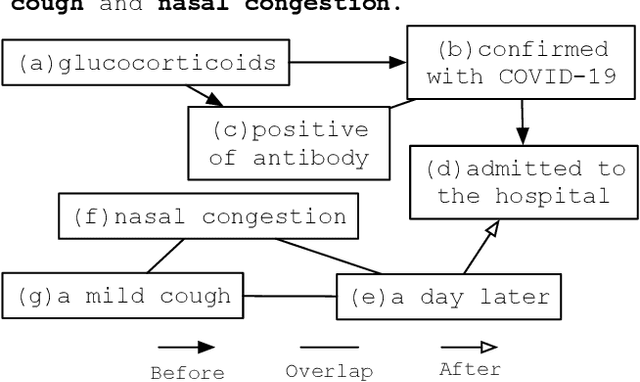
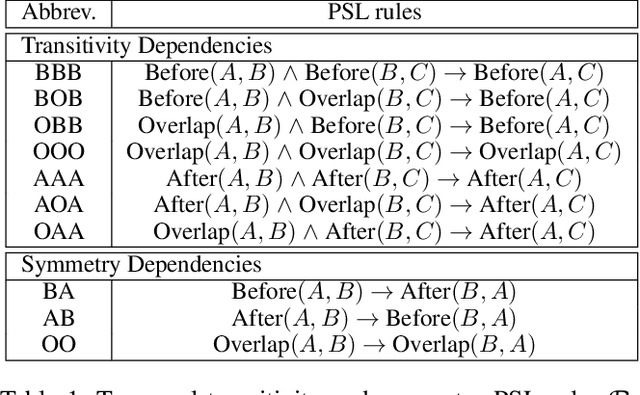
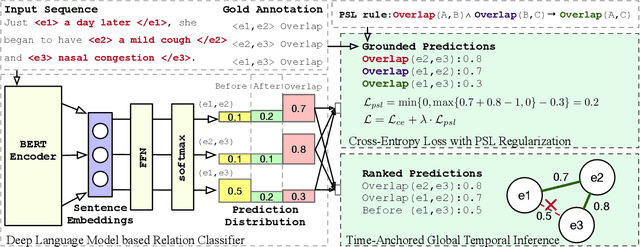
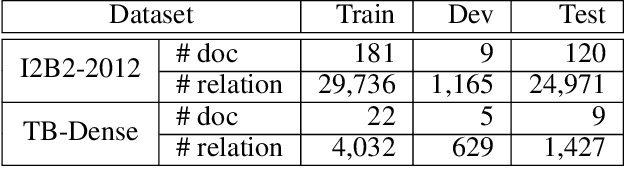
There has been a steady need in the medical community to precisely extract the temporal relations between clinical events. In particular, temporal information can facilitate a variety of downstream applications such as case report retrieval and medical question answering. Existing methods either require expensive feature engineering or are incapable of modeling the global relational dependencies among the events. In this paper, we propose a novel method, Clinical Temporal ReLation Exaction with Probabilistic Soft Logic Regularization and Global Inference (CTRL-PG) to tackle the problem at the document level. Extensive experiments on two benchmark datasets, I2B2-2012 and TB-Dense, demonstrate that CTRL-PG significantly outperforms baseline methods for temporal relation extraction.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge