Siyuan Jiang
Semi-parametric Functional Classification via Path Signatures Logistic Regression
Jul 09, 2025Abstract:We propose Path Signatures Logistic Regression (PSLR), a semi-parametric framework for classifying vector-valued functional data with scalar covariates. Classical functional logistic regression models rely on linear assumptions and fixed basis expansions, which limit flexibility and degrade performance under irregular sampling. PSLR overcomes these issues by leveraging truncated path signatures to construct a finite-dimensional, basis-free representation that captures nonlinear and cross-channel dependencies. By embedding trajectories as time-augmented paths, PSLR extracts stable, geometry-aware features that are robust to sampling irregularity without requiring a common time grid, while still preserving subject-specific timing patterns. We establish theoretical guarantees for the existence and consistent estimation of the optimal truncation order, along with non-asymptotic risk bounds. Experiments on synthetic and real-world datasets show that PSLR outperforms traditional functional classifiers in accuracy, robustness, and interpretability, particularly under non-uniform sampling schemes. Our results highlight the practical and theoretical benefits of integrating rough path theory into modern functional data analysis.
DeepFRC: An End-to-End Deep Learning Model for Functional Registration and Classification
Jan 30, 2025



Abstract:Functional data analysis (FDA) is essential for analyzing continuous, high-dimensional data, yet existing methods often decouple functional registration and classification, limiting their efficiency and performance. We present DeepFRC, an end-to-end deep learning framework that unifies these tasks within a single model. Our approach incorporates an alignment module that learns time warping functions via elastic function registration and a learnable basis representation module for dimensionality reduction on aligned data. This integration enhances both alignment accuracy and predictive performance. Theoretical analysis establishes that DeepFRC achieves low misalignment and generalization error, while simulations elucidate the progression of registration, reconstruction, and classification during training. Experiments on real-world datasets demonstrate that DeepFRC consistently outperforms state-of-the-art methods, particularly in addressing complex registration challenges. Code is available at: https://github.com/Drivergo-93589/DeepFRC.
aiXcoder-7B: A Lightweight and Effective Large Language Model for Code Completion
Oct 17, 2024



Abstract:Large Language Models (LLMs) have been widely used in code completion, and researchers are focusing on scaling up LLMs to improve their accuracy. However, larger LLMs will increase the response time of code completion and decrease the developers' productivity. In this paper, we propose a lightweight and effective LLM for code completion named aiXcoder-7B. Compared to existing LLMs, aiXcoder-7B achieves higher code completion accuracy while having smaller scales (i.e., 7 billion parameters). We attribute the superiority of aiXcoder-7B to three key factors: (1) Multi-objective training. We employ three training objectives, one of which is our proposed Structured Fill-In-the-Middle (SFIM). SFIM considers the syntax structures in code and effectively improves the performance of LLMs for code. (2) Diverse data sampling strategies. They consider inter-file relationships and enhance the capability of LLMs in understanding cross-file contexts. (3) Extensive high-quality data. We establish a rigorous data collection pipeline and consume a total of 1.2 trillion unique tokens for training aiXcoder-7B. This vast volume of data enables aiXcoder-7B to learn a broad distribution of code. We evaluate aiXcoder-7B in five popular code completion benchmarks and a new benchmark collected by this paper. The results show that aiXcoder-7B outperforms the latest six LLMs with similar sizes and even surpasses four larger LLMs (e.g., StarCoder2-15B and CodeLlama-34B), positioning aiXcoder-7B as a lightweight and effective LLM for academia and industry. Finally, we summarize three valuable insights for helping practitioners train the next generations of LLMs for code. aiXcoder-7B has been open-souced and gained significant attention. As of the submission date, aiXcoder-7B has received 2,193 GitHub Stars.
Technique Report of CVPR 2024 PBDL Challenges
Jun 15, 2024



Abstract:The intersection of physics-based vision and deep learning presents an exciting frontier for advancing computer vision technologies. By leveraging the principles of physics to inform and enhance deep learning models, we can develop more robust and accurate vision systems. Physics-based vision aims to invert the processes to recover scene properties such as shape, reflectance, light distribution, and medium properties from images. In recent years, deep learning has shown promising improvements for various vision tasks, and when combined with physics-based vision, these approaches can enhance the robustness and accuracy of vision systems. This technical report summarizes the outcomes of the Physics-Based Vision Meets Deep Learning (PBDL) 2024 challenge, held in CVPR 2024 workshop. The challenge consisted of eight tracks, focusing on Low-Light Enhancement and Detection as well as High Dynamic Range (HDR) Imaging. This report details the objectives, methodologies, and results of each track, highlighting the top-performing solutions and their innovative approaches.
Deep RAW Image Super-Resolution. A NTIRE 2024 Challenge Survey
Apr 24, 2024



Abstract:This paper reviews the NTIRE 2024 RAW Image Super-Resolution Challenge, highlighting the proposed solutions and results. New methods for RAW Super-Resolution could be essential in modern Image Signal Processing (ISP) pipelines, however, this problem is not as explored as in the RGB domain. Th goal of this challenge is to upscale RAW Bayer images by 2x, considering unknown degradations such as noise and blur. In the challenge, a total of 230 participants registered, and 45 submitted results during thee challenge period. The performance of the top-5 submissions is reviewed and provided here as a gauge for the current state-of-the-art in RAW Image Super-Resolution.
Automatic nodule identification and differentiation in ultrasound videos to facilitate per-nodule examination
Oct 10, 2023



Abstract:Ultrasound is a vital diagnostic technique in health screening, with the advantages of non-invasive, cost-effective, and radiation free, and therefore is widely applied in the diagnosis of nodules. However, it relies heavily on the expertise and clinical experience of the sonographer. In ultrasound images, a single nodule might present heterogeneous appearances in different cross-sectional views which makes it hard to perform per-nodule examination. Sonographers usually discriminate different nodules by examining the nodule features and the surrounding structures like gland and duct, which is cumbersome and time-consuming. To address this problem, we collected hundreds of breast ultrasound videos and built a nodule reidentification system that consists of two parts: an extractor based on the deep learning model that can extract feature vectors from the input video clips and a real-time clustering algorithm that automatically groups feature vectors by nodules. The system obtains satisfactory results and exhibits the capability to differentiate ultrasound videos. As far as we know, it's the first attempt to apply re-identification technique in the ultrasonic field.
Statement-based Memory for Neural Source Code Summarization
Jul 21, 2023Abstract:Source code summarization is the task of writing natural language descriptions of source code behavior. Code summarization underpins software documentation for programmers. Short descriptions of code help programmers understand the program quickly without having to read the code itself. Lately, neural source code summarization has emerged as the frontier of research into automated code summarization techniques. By far the most popular targets for summarization are program subroutines. The idea, in a nutshell, is to train an encoder-decoder neural architecture using large sets of examples of subroutines extracted from code repositories. The encoder represents the code and the decoder represents the summary. However, most current approaches attempt to treat the subroutine as a single unit. For example, by taking the entire subroutine as input to a Transformer or RNN-based encoder. But code behavior tends to depend on the flow from statement to statement. Normally dynamic analysis may shed light on this flow, but dynamic analysis on hundreds of thousands of examples in large datasets is not practical. In this paper, we present a statement-based memory encoder that learns the important elements of flow during training, leading to a statement-based subroutine representation without the need for dynamic analysis. We implement our encoder for code summarization and demonstrate a significant improvement over the state-of-the-art.
DSDP: A Blind Docking Strategy Accelerated by GPUs
Mar 16, 2023



Abstract:Virtual screening, including molecular docking, plays an essential role in drug discovery. Many traditional and machine-learning based methods are available to fulfil the docking task. The traditional docking methods are normally extensively time-consuming, and their performance in blind docking remains to be improved. Although the runtime of docking based on machine learning is significantly decreased, their accuracy is still limited. In this study, we take the advantage of both traditional and machine-learning based methods, and present a method Deep Site and Docking Pose (DSDP) to improve the performance of blind docking. For the traditional blind docking, the entire protein is covered by a cube, and the initial positions of ligands are randomly generated in the cube. In contract, DSDP can predict the binding site of proteins and provide an accurate searching space and initial positions for the further conformational sampling. The docking task of DSDP makes use of the score function and a similar but modified searching strategy of AutoDock Vina, accelerated by implementation in GPUs. We systematically compare its performance with the state-of-the-art methods, including Autodock Vina, GNINA, QuickVina, SMINA, and DiffDock. DSDP reaches a 29.8% top-1 success rate (RMSD < 2 {\AA}) on an unbiased and challenging test dataset with 1.2 s wall-clock computational time per system. Its performances on DUD-E dataset and the time-split PDBBind dataset used in EquiBind, TankBind, and DiffDock are also effective, presenting a 57.2% and 41.8% top-1 success rate with 0.8 s and 1.0 s per system, respectively.
Automatically Generating Commit Messages from Diffs using Neural Machine Translation
Aug 30, 2017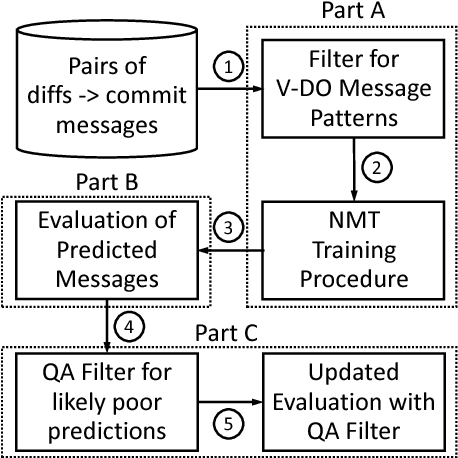
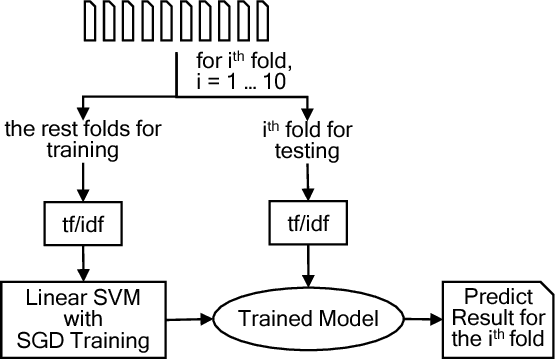
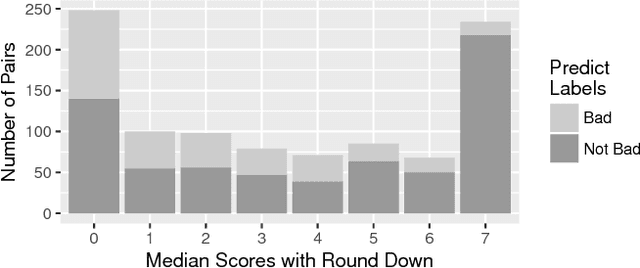
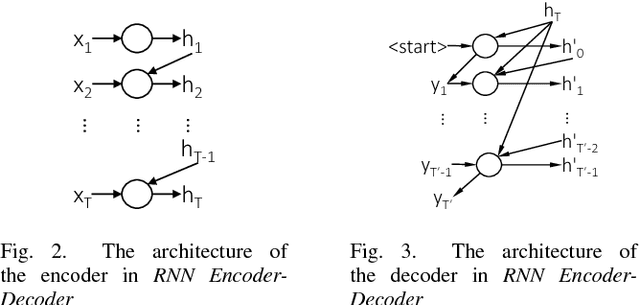
Abstract:Commit messages are a valuable resource in comprehension of software evolution, since they provide a record of changes such as feature additions and bug repairs. Unfortunately, programmers often neglect to write good commit messages. Different techniques have been proposed to help programmers by automatically writing these messages. These techniques are effective at describing what changed, but are often verbose and lack context for understanding the rationale behind a change. In contrast, humans write messages that are short and summarize the high level rationale. In this paper, we adapt Neural Machine Translation (NMT) to automatically "translate" diffs into commit messages. We trained an NMT algorithm using a corpus of diffs and human-written commit messages from the top 1k Github projects. We designed a filter to help ensure that we only trained the algorithm on higher-quality commit messages. Our evaluation uncovered a pattern in which the messages we generate tend to be either very high or very low quality. Therefore, we created a quality-assurance filter to detect cases in which we are unable to produce good messages, and return a warning instead.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge