Shwetak Patel
CoDaS: AI Co-Data-Scientist for Biomarker Discovery via Wearable Sensors
Apr 16, 2026Abstract:Scientific discovery in digital health requires converting continuous physiological signals from wearable devices into clinically actionable biomarkers. We introduce CoDaS (AI Co-Data-Scientist), a multi-agent system that structures biomarker discovery as an iterative process combining hypothesis generation, statistical analysis, adversarial validation, and literature-grounded reasoning with human oversight using large-scale wearable datasets. Across three cohorts totaling 9,279 participant-observations, CoDaS identified 41 candidate digital biomarkers for mental health and 25 for metabolic outcomes, each subjected to an internal validation battery spanning replication, stability, robustness, and discriminative power. Across two independent depression cohorts, CoDaS surfaced circadian instability-related features in both datasets, reflected in sleep duration variability (DWB, ρ= 0.252, p < 0.001) and sleep onset variability (GLOBEM, ρ= 0.126, p < 0.001). In a metabolic cohort, CoDaS derived a cardiovascular fitness index (steps/resting heart rate; ρ= -0.374, p < 0.001), and recovered established clinical associations, including the hepatic function ratio (AST/ALT; ρ= -0.375, p < 0.001), a known correlate of insulin resistance. Incorporating CoDaS-derived features alongside demographic variables led to modest but consistent improvements in predictive performance, with cross-validated ΔR^2 increases of 0.040 for depression and 0.021 for insulin resistance. These findings suggest that CoDaS enables systematic and traceable hypothesis generation and prioritization for biomarker discovery from large-scale wearable data.
Towards a Science of Scaling Agent Systems
Dec 17, 2025Abstract:Agents, language model-based systems that are capable of reasoning, planning, and acting are becoming the dominant paradigm for real-world AI applications. Despite this widespread adoption, the principles that determine their performance remain underexplored. We address this by deriving quantitative scaling principles for agent systems. We first formalize a definition for agentic evaluation and characterize scaling laws as the interplay between agent quantity, coordination structure, model capability, and task properties. We evaluate this across four benchmarks: Finance-Agent, BrowseComp-Plus, PlanCraft, and Workbench. With five canonical agent architectures (Single-Agent and four Multi-Agent Systems: Independent, Centralized, Decentralized, Hybrid), instantiated across three LLM families, we perform a controlled evaluation spanning 180 configurations. We derive a predictive model using coordination metrics, that achieves cross-validated R^2=0.524, enabling prediction on unseen task domains. We identify three effects: (1) a tool-coordination trade-off: under fixed computational budgets, tool-heavy tasks suffer disproportionately from multi-agent overhead. (2) a capability saturation: coordination yields diminishing or negative returns once single-agent baselines exceed ~45%. (3) topology-dependent error amplification: independent agents amplify errors 17.2x, while centralized coordination contains this to 4.4x. Centralized coordination improves performance by 80.8% on parallelizable tasks, while decentralized coordination excels on web navigation (+9.2% vs. +0.2%). Yet for sequential reasoning tasks, every multi-agent variants degraded performance by 39-70%. The framework predicts the optimal coordination strategy for 87% of held-out configurations. Out-of-sample validation on GPT-5.2, achieves MAE=0.071 and confirms four of five scaling principles generalize to unseen frontier models.
Smartphone monitoring of smiling as a behavioral proxy of well-being in everyday life
Dec 10, 2025Abstract:Subjective well-being is a cornerstone of individual and societal health, yet its scientific measurement has traditionally relied on self-report methods prone to recall bias and high participant burden. This has left a gap in our understanding of well-being as it is expressed in everyday life. We hypothesized that candid smiles captured during natural smartphone interactions could serve as a scalable, objective behavioral correlate of positive affect. To test this, we analyzed 405,448 video clips passively recorded from 233 consented participants over one week. Using a deep learning model to quantify smile intensity, we identified distinct diurnal and daily patterns. Daily patterns of smile intensity across the week showed strong correlation with national survey data on happiness (r=0.92), and diurnal rhythms documented close correspondence with established results from the day reconstruction method (r=0.80). Higher daily mean smile intensity was significantly associated with more physical activity (Beta coefficient = 0.043, 95% CI [0.001, 0.085]) and greater light exposure (Beta coefficient = 0.038, [0.013, 0.063]), whereas no significant effects were found for smartphone use. These findings suggest that passive smartphone sensing could serve as a powerful, ecologically valid methodology for studying the dynamics of affective behavior and open the door to understanding this behavior at a population scale.
ConvFill: Model Collaboration for Responsive Conversational Voice Agents
Nov 10, 2025Abstract:Deploying conversational voice agents with large language models faces a critical challenge: cloud-based foundation models provide deep reasoning and domain knowledge but introduce latency that disrupts natural conversation, while on-device models respond immediately but lack sophistication. We propose conversational infill, a task where a lightweight on-device model generates contextually appropriate dialogue while seamlessly incorporating streaming knowledge from a powerful backend model. This approach decouples response latency from model capability, enabling systems that feel responsive while accessing the full power of large-scale models. We present ConvFill, a 360M parameter model trained on synthetic multi-domain conversations. Evaluation across multiple backend models shows that conversational infill can be successfully learned, with ConvFill achieving accuracy improvements of 36-42% over standalone small models of the same size while consistently retaining sub-200ms response latencies. Our results demonstrate the promise of this approach for building on-device conversational agents that are both immediately responsive and knowledgeable.
SnappyMeal: Design and Longitudinal Evaluation of a Multimodal AI Food Logging Application
Nov 05, 2025Abstract:Food logging, both self-directed and prescribed, plays a critical role in uncovering correlations between diet, medical, fitness, and health outcomes. Through conversations with nutritional experts and individuals who practice dietary tracking, we find current logging methods, such as handwritten and app-based journaling, are inflexible and result in low adherence and potentially inaccurate nutritional summaries. These findings, corroborated by prior literature, emphasize the urgent need for improved food logging methods. In response, we propose SnappyMeal, an AI-powered dietary tracking system that leverages multimodal inputs to enable users to more flexibly log their food intake. SnappyMeal introduces goal-dependent follow-up questions to intelligently seek missing context from the user and information retrieval from user grocery receipts and nutritional databases to improve accuracy. We evaluate SnappyMeal through publicly available nutrition benchmarks and a multi-user, 3-week, in-the-wild deployment capturing over 500 logged food instances. Users strongly praised the multiple available input methods and reported a strong perceived accuracy. These insights suggest that multimodal AI systems can be leveraged to significantly improve dietary tracking flexibility and context-awareness, laying the groundwork for a new class of intelligent self-tracking applications.
Towards Autonomous Sustainability Assessment via Multimodal AI Agents
Jul 22, 2025



Abstract:Interest in sustainability information has surged in recent years. However, the data required for a life cycle assessment (LCA) that maps the materials and processes from product manufacturing to disposal into environmental impacts (EI) are often unavailable. Here we reimagine conventional LCA by introducing multimodal AI agents that emulate interactions between LCA experts and stakeholders like product managers and engineers to calculate the cradle-to-gate (production) carbon emissions of electronic devices. The AI agents iteratively generate a detailed life-cycle inventory leveraging a custom data abstraction and software tools that extract information from online text and images from repair communities and government certifications. This approach reduces weeks or months of expert time to under one minute and closes data availability gaps while yielding carbon footprint estimates within 19% of expert LCAs with zero proprietary data. Additionally, we develop a method to directly estimate EI by comparing an input to a cluster of products with similar descriptions and known carbon footprints. This runs in 3 ms on a laptop with a MAPE of 12.28% on electronic products. Further, we develop a data-driven method to generate emission factors. We use the properties of an unknown material to represent it as a weighted sum of emission factors for similar materials. Compared to human experts picking the closest LCA database entry, this improves MAPE by 120.26%. We analyze the data and compute scaling of this approach and discuss its implications for future LCA workflows.
SensorLM: Learning the Language of Wearable Sensors
Jun 10, 2025Abstract:We present SensorLM, a family of sensor-language foundation models that enable wearable sensor data understanding with natural language. Despite its pervasive nature, aligning and interpreting sensor data with language remains challenging due to the lack of paired, richly annotated sensor-text descriptions in uncurated, real-world wearable data. We introduce a hierarchical caption generation pipeline designed to capture statistical, structural, and semantic information from sensor data. This approach enabled the curation of the largest sensor-language dataset to date, comprising over 59.7 million hours of data from more than 103,000 people. Furthermore, SensorLM extends prominent multimodal pretraining architectures (e.g., CLIP, CoCa) and recovers them as specific variants within a generic architecture. Extensive experiments on real-world tasks in human activity analysis and healthcare verify the superior performance of SensorLM over state-of-the-art in zero-shot recognition, few-shot learning, and cross-modal retrieval. SensorLM also demonstrates intriguing capabilities including scaling behaviors, label efficiency, sensor captioning, and zero-shot generalization to unseen tasks.
RADAR: Benchmarking Language Models on Imperfect Tabular Data
Jun 09, 2025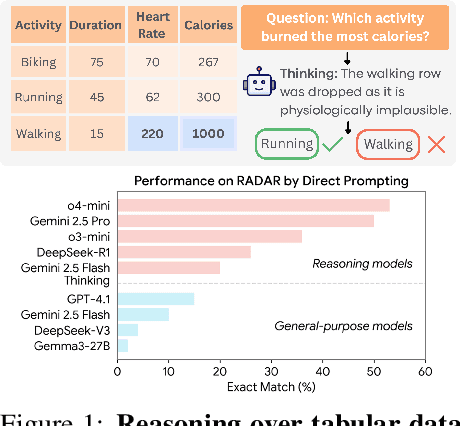
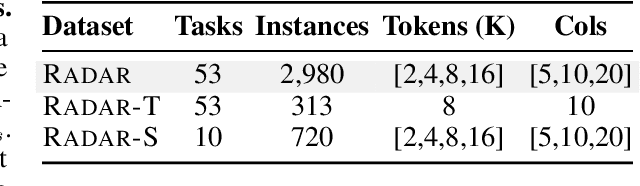
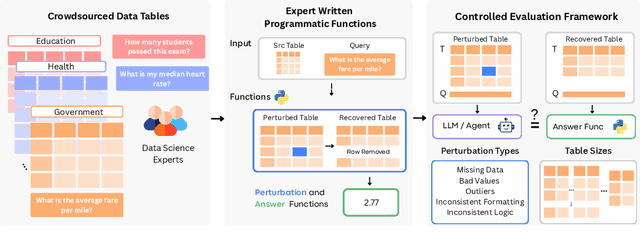
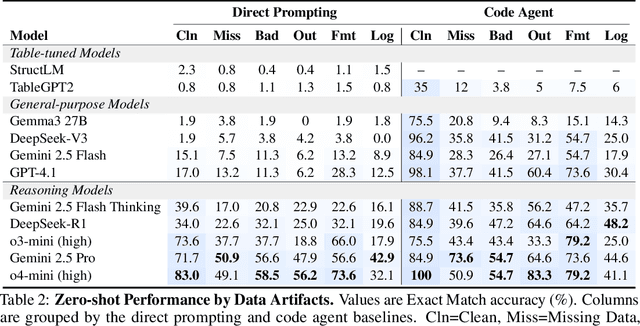
Abstract:Language models (LMs) are increasingly being deployed to perform autonomous data analyses. However, their data awareness -- the ability to recognize, reason over, and appropriately handle data artifacts such as missing values, outliers, and logical inconsistencies -- remains underexplored. These artifacts are especially common in real-world tabular data and, if mishandled, can significantly compromise the validity of analytical conclusions. To address this gap, we present RADAR, a benchmark for systematically evaluating data-aware reasoning on tabular data. We develop a framework to simulate data artifacts via programmatic perturbations to enable targeted evaluation of model behavior. RADAR comprises 2980 table query pairs, grounded in real-world data spanning 9 domains and 5 data artifact types. In addition to evaluating artifact handling, RADAR systematically varies table size to study how reasoning performance holds when increasing table size. Our evaluation reveals that, despite decent performance on tables without data artifacts, frontier models degrade significantly when data artifacts are introduced, exposing critical gaps in their capacity for robust, data-aware analysis. Designed to be flexible and extensible, RADAR supports diverse perturbation types and controllable table sizes, offering a valuable resource for advancing tabular reasoning.
LSM-2: Learning from Incomplete Wearable Sensor Data
Jun 05, 2025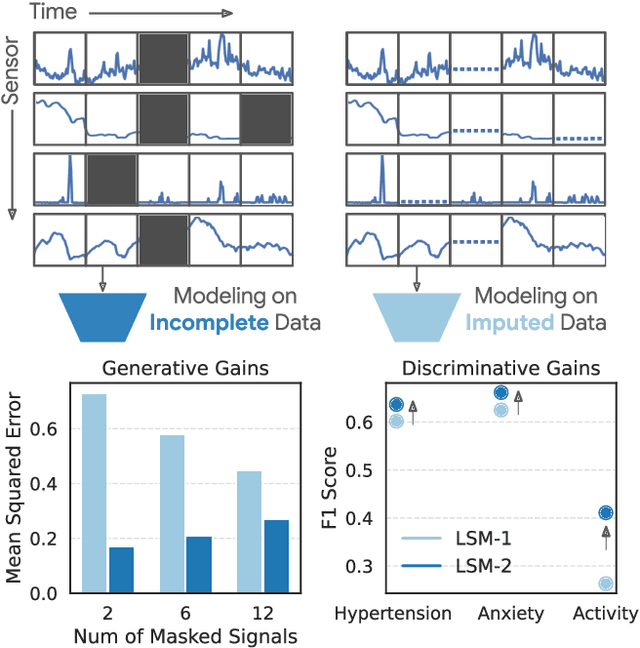
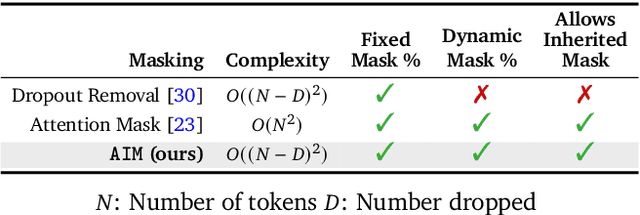
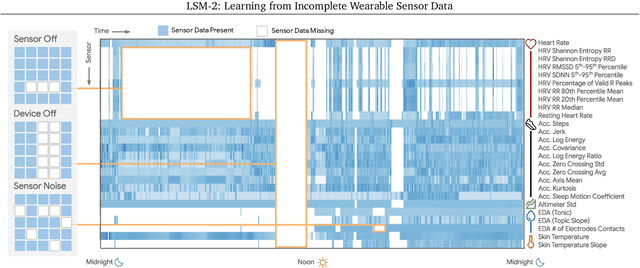
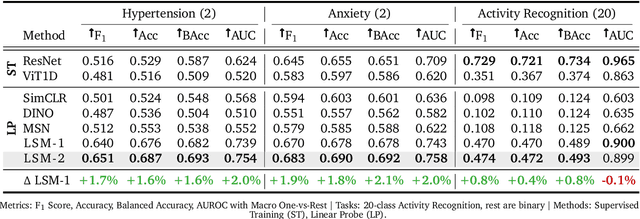
Abstract:Foundation models, a cornerstone of recent advancements in machine learning, have predominantly thrived on complete and well-structured data. Wearable sensor data frequently suffers from significant missingness, posing a substantial challenge for self-supervised learning (SSL) models that typically assume complete data inputs. This paper introduces the second generation of Large Sensor Model (LSM-2) with Adaptive and Inherited Masking (AIM), a novel SSL approach that learns robust representations directly from incomplete data without requiring explicit imputation. AIM's core novelty lies in its use of learnable mask tokens to model both existing ("inherited") and artificially introduced missingness, enabling it to robustly handle fragmented real-world data during inference. Pre-trained on an extensive dataset of 40M hours of day-long multimodal sensor data, our LSM-2 with AIM achieves the best performance across a diverse range of tasks, including classification, regression and generative modeling. Furthermore, LSM-2 with AIM exhibits superior scaling performance, and critically, maintains high performance even under targeted missingness scenarios, reflecting clinically coherent patterns, such as the diagnostic value of nighttime biosignals for hypertension prediction. This makes AIM a more reliable choice for real-world wearable data applications.
Insulin Resistance Prediction From Wearables and Routine Blood Biomarkers
Apr 30, 2025Abstract:Insulin resistance, a precursor to type 2 diabetes, is characterized by impaired insulin action in tissues. Current methods for measuring insulin resistance, while effective, are expensive, inaccessible, not widely available and hinder opportunities for early intervention. In this study, we remotely recruited the largest dataset to date across the US to study insulin resistance (N=1,165 participants, with median BMI=28 kg/m2, age=45 years, HbA1c=5.4%), incorporating wearable device time series data and blood biomarkers, including the ground-truth measure of insulin resistance, homeostatic model assessment for insulin resistance (HOMA-IR). We developed deep neural network models to predict insulin resistance based on readily available digital and blood biomarkers. Our results show that our models can predict insulin resistance by combining both wearable data and readily available blood biomarkers better than either of the two data sources separately (R2=0.5, auROC=0.80, Sensitivity=76%, and specificity 84%). The model showed 93% sensitivity and 95% adjusted specificity in obese and sedentary participants, a subpopulation most vulnerable to developing type 2 diabetes and who could benefit most from early intervention. Rigorous evaluation of model performance, including interpretability, and robustness, facilitates generalizability across larger cohorts, which is demonstrated by reproducing the prediction performance on an independent validation cohort (N=72 participants). Additionally, we demonstrated how the predicted insulin resistance can be integrated into a large language model agent to help understand and contextualize HOMA-IR values, facilitating interpretation and safe personalized recommendations. This work offers the potential for early detection of people at risk of type 2 diabetes and thereby facilitate earlier implementation of preventative strategies.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge