Jingbo Zhou
LoReC: Rethinking Large Language Models for Graph Data Analysis
Apr 20, 2026Abstract:The advent of Large Language Models (LLMs) has fundamentally reshaped the way we interact with graphs, giving rise to a new paradigm called GraphLLM. As revealed in recent studies, graph learning can benefit from LLMs. However, we observe limited benefits when we directly utilize LLMs to make predictions for graph-related tasks within GraphLLM paradigm, which even yields suboptimal results compared to conventional GNN-based approaches. Through in-depth analysis, we find this failure can be attributed to LLMs' limited capability for processing graph data and their tendency to overlook graph information. To address this issue, we propose LoReC (Look, Remember, and Contrast), a novel plug-and-play method for GraphLLM paradigm, which enhances LLM's understanding of graph data through three stages: (1) Look: redistributing attention to graph; (2) Remember: re-injecting graph information into the Feed-Forward Network (FFN); (3) Contrast: rectifying the vanilla logits produced in the decoding process. Extensive experiments demonstrate that LoReC brings notable improvements over current GraphLLM methods and outperforms GNN-based approaches across diverse datasets. The implementation is available at https://github.com/Git-King-Zhan/LoReC.
MAT-Cell: A Multi-Agent Tree-Structured Reasoning Framework for Batch-Level Single-Cell Annotation
Apr 07, 2026Abstract:Automated cellular reasoning faces a core dichotomy: supervised methods fall into the Reference Trap and fail to generalize to out-of-distribution cell states, while large language models (LLMs), without grounded biological priors, suffer from a Signal-to-Noise Paradox that produces spurious associations. We propose MAT-Cell, a neuro-symbolic reasoning framework that reframes single-cell analysis from black-box classification into constructive, verifiable proof generation. MAT-Cell injects symbolic constraints through adaptive Retrieval-Augmented Generation (RAG) to ground neural reasoning in biological axioms and reduce transcriptomic noise. It further employs a dialectic verification process with homogeneous rebuttal agents to audit and prune reasoning paths, forming syllogistic derivation trees that enforce logical consistency.Across large-scale and cross-species benchmarks, MAT-Cell significantly outperforms state-of-the-art (SOTA) models and maintains robust per-formance in challenging scenarios where baselinemethods severely degrade. Code is available at https://gith ub.com/jiangliu91/MAT-Cell-A-Mul ti-Agent-Tree-Structured-Reasoni ng-Framework-for-Batch-Level-Sin gle-Cell-Annotation.
Regressor-guided Diffusion Model for De Novo Peptide Sequencing with Explicit Mass Control
Feb 23, 2026Abstract:The discovery of novel proteins relies on sensitive protein identification, for which de novo peptide sequencing (DNPS) from mass spectra is a crucial approach. While deep learning has advanced DNPS, existing models inadequately enforce the fundamental mass consistency constraint, that a predicted peptide's mass must match the experimental measured precursor mass. Previous DNPS methods often treat this critical information as a simple input feature or use it in post-processing, leading to numerous implausible predictions that do not adhere to this fundamental physical property. To address this limitation, we introduce DiffuNovo, a novel regressor-guided diffusion model for de novo peptide sequencing that provides explicit peptide-level mass control. Our approach integrates the mass constraint at two critical stages: during training, a novel peptide-level mass loss guides model optimization, while at inference, regressor-based guidance from gradient-based updates in the latent space steers the generation to compel the predicted peptide adheres to the mass constraint. Comprehensive evaluations on established benchmarks demonstrate that DiffuNovo surpasses state-of-the-art methods in DNPS accuracy. Additionally, as the first DNPS model to employ a diffusion model as its core backbone, DiffuNovo leverages the powerful controllability of diffusion architecture and achieves a significant reduction in mass error, thereby producing much more physically plausible peptides. These innovations represent a substantial advancement toward robust and broadly applicable DNPS. The source code is available in the supplementary material.
VecFormer: Towards Efficient and Generalizable Graph Transformer with Graph Token Attention
Feb 23, 2026Abstract:Graph Transformer has demonstrated impressive capabilities in the field of graph representation learning. However, existing approaches face two critical challenges: (1) most models suffer from exponentially increasing computational complexity, making it difficult to scale to large graphs; (2) attention mechanisms based on node-level operations limit the flexibility of the model and result in poor generalization performance in out-of-distribution (OOD) scenarios. To address these issues, we propose \textbf{VecFormer} (the \textbf{Vec}tor Quantized Graph Trans\textbf{former}), an efficient and highly generalizable model for node classification, particularly under OOD settings. VecFormer adopts a two-stage training paradigm. In the first stage, two codebooks are used to reconstruct the node features and the graph structure, aiming to learn the rich semantic \texttt{Graph Codes}. In the second stage, attention mechanisms are performed at the \texttt{Graph Token} level based on the transformed cross codebook, reducing computational complexity while enhancing the model's generalization capability. Extensive experiments on datasets of various sizes demonstrate that VecFormer outperforms the existing Graph Transformer in both performance and speed.
Unrewarded Exploration in Large Language Models Reveals Latent Learning from Psychology
Jan 30, 2026Abstract:Latent learning, classically theorized by Tolman, shows that biological agents (e.g., rats) can acquire internal representations of their environment without rewards, enabling rapid adaptation once rewards are introduced. In contrast, from a cognitive science perspective, reward learning remains overly dependent on external feedback, limiting flexibility and generalization. Although recent advances in the reasoning capabilities of large language models (LLMs), such as OpenAI-o1 and DeepSeek-R1, mark a significant breakthrough, these models still rely primarily on reward-centric reinforcement learning paradigms. Whether and how the well-established phenomenon of latent learning in psychology can inform or emerge within LLMs' training remains largely unexplored. In this work, we present novel findings from our experiments that LLMs also exhibit the latent learning dynamics. During an initial phase of unrewarded exploration, LLMs display modest performance improvements, as this phase allows LLMs to organize task-relevant knowledge without being constrained by reward-driven biases, and performance is further enhanced once rewards are introduced. LLMs post-trained under this two-stage exploration regime ultimately achieve higher competence than those post-trained with reward-based reinforcement learning throughout. Beyond these empirical observations, we also provide theoretical analyses for our experiments explaining why unrewarded exploration yields performance gains, offering a mechanistic account of these dynamics. Specifically, we conducted extensive experiments across multiple model families and diverse task domains to establish the existence of the latent learning dynamics in LLMs.
OmegaUse: Building a General-Purpose GUI Agent for Autonomous Task Execution
Jan 28, 2026Abstract:Graphical User Interface (GUI) agents show great potential for enabling foundation models to complete real-world tasks, revolutionizing human-computer interaction and improving human productivity. In this report, we present OmegaUse, a general-purpose GUI agent model for autonomous task execution on both mobile and desktop platforms, supporting computer-use and phone-use scenarios. Building an effective GUI agent model relies on two factors: (1) high-quality data and (2) effective training methods. To address these, we introduce a carefully engineered data-construction pipeline and a decoupled training paradigm. For data construction, we leverage rigorously curated open-source datasets and introduce a novel automated synthesis framework that integrates bottom-up autonomous exploration with top-down taxonomy-guided generation to create high-fidelity synthetic data. For training, to better leverage these data, we adopt a two-stage strategy: Supervised Fine-Tuning (SFT) to establish fundamental interaction syntax, followed by Group Relative Policy Optimization (GRPO) to improve spatial grounding and sequential planning. To balance computational efficiency with agentic reasoning capacity, OmegaUse is built on a Mixture-of-Experts (MoE) backbone. To evaluate cross-terminal capabilities in an offline setting, we introduce OS-Nav, a benchmark suite spanning multiple operating systems: ChiM-Nav, targeting Chinese Android mobile environments, and Ubu-Nav, focusing on routine desktop interactions on Ubuntu. Extensive experiments show that OmegaUse is highly competitive across established GUI benchmarks, achieving a state-of-the-art (SOTA) score of 96.3% on ScreenSpot-V2 and a leading 79.1% step success rate on AndroidControl. OmegaUse also performs strongly on OS-Nav, reaching 74.24% step success on ChiM-Nav and 55.9% average success on Ubu-Nav.
MergeDNA: Context-aware Genome Modeling with Dynamic Tokenization through Token Merging
Nov 17, 2025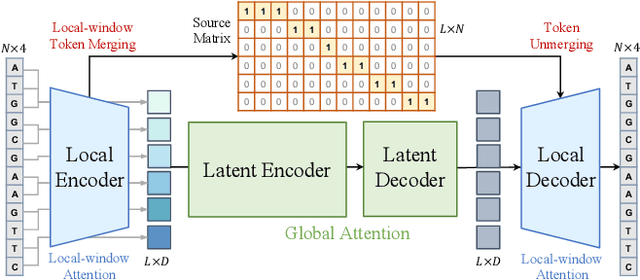
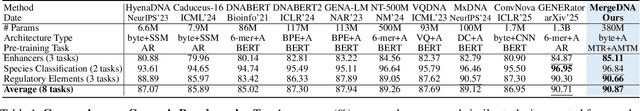
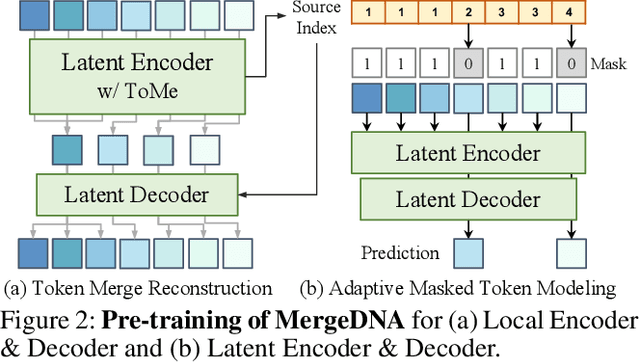
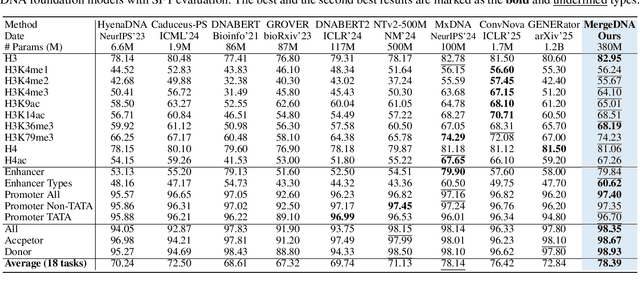
Abstract:Modeling genomic sequences faces two unsolved challenges: the information density varies widely across different regions, while there is no clearly defined minimum vocabulary unit. Relying on either four primitive bases or independently designed DNA tokenizers, existing approaches with naive masked language modeling pre-training often fail to adapt to the varying complexities of genomic sequences. Leveraging Token Merging techniques, this paper introduces a hierarchical architecture that jointly optimizes a dynamic genomic tokenizer and latent Transformers with context-aware pre-training tasks. As for network structures, the tokenization module automatically chunks adjacent bases into words by stacking multiple layers of the differentiable token merging blocks with local-window constraints, then a Latent Encoder captures the global context of these merged words by full-attention blocks. Symmetrically employing a Latent Decoder and a Local Decoder, MergeDNA learns with two pre-training tasks: Merged Token Reconstruction simultaneously trains the dynamic tokenization module and adaptively filters important tokens, while Adaptive Masked Token Modeling learns to predict these filtered tokens to capture informative contents. Extensive experiments show that MergeDNA achieves superior performance on three popular DNA benchmarks and several multi-omics tasks with fine-tuning or zero-shot evaluation, outperforming typical tokenization methods and large-scale DNA foundation models.
PRESCRIBE: Predicting Single-Cell Responses with Bayesian Estimation
Oct 09, 2025



Abstract:In single-cell perturbation prediction, a central task is to forecast the effects of perturbing a gene unseen in the training data. The efficacy of such predictions depends on two factors: (1) the similarity of the target gene to those covered in the training data, which informs model (epistemic) uncertainty, and (2) the quality of the corresponding training data, which reflects data (aleatoric) uncertainty. Both factors are critical for determining the reliability of a prediction, particularly as gene perturbation is an inherently stochastic biochemical process. In this paper, we propose PRESCRIBE (PREdicting Single-Cell Response wIth Bayesian Estimation), a multivariate deep evidential regression framework designed to measure both sources of uncertainty jointly. Our analysis demonstrates that PRESCRIBE effectively estimates a confidence score for each prediction, which strongly correlates with its empirical accuracy. This capability enables the filtering of untrustworthy results, and in our experiments, it achieves steady accuracy improvements of over 3% compared to comparable baselines.
AAPO: Enhance the Reasoning Capabilities of LLMs with Advantage Momentum
May 20, 2025Abstract:Reinforcement learning (RL) has emerged as an effective approach for enhancing the reasoning capabilities of large language models (LLMs), especially in scenarios where supervised fine-tuning (SFT) falls short due to limited chain-of-thought (CoT) data. Among RL-based post-training methods, group relative advantage estimation, as exemplified by Group Relative Policy Optimization (GRPO), has attracted considerable attention for eliminating the dependency on the value model, thereby simplifying training compared to traditional approaches like Proximal Policy Optimization (PPO). However, we observe that exsiting group relative advantage estimation method still suffers from training inefficiencies, particularly when the estimated advantage approaches zero. To address this limitation, we propose Advantage-Augmented Policy Optimization (AAPO), a novel RL algorithm that optimizes the cross-entropy (CE) loss using advantages enhanced through a momentum-based estimation scheme. This approach effectively mitigates the inefficiencies associated with group relative advantage estimation. Experimental results on multiple mathematical reasoning benchmarks demonstrate the superior performance of AAPO.
GRAPE: Heterogeneous Graph Representation Learning for Genetic Perturbation with Coding and Non-Coding Biotype
May 06, 2025



Abstract:Predicting genetic perturbations enables the identification of potentially crucial genes prior to wet-lab experiments, significantly improving overall experimental efficiency. Since genes are the foundation of cellular life, building gene regulatory networks (GRN) is essential to understand and predict the effects of genetic perturbations. However, current methods fail to fully leverage gene-related information, and solely rely on simple evaluation metrics to construct coarse-grained GRN. More importantly, they ignore functional differences between biotypes, limiting the ability to capture potential gene interactions. In this work, we leverage pre-trained large language model and DNA sequence model to extract features from gene descriptions and DNA sequence data, respectively, which serve as the initialization for gene representations. Additionally, we introduce gene biotype information for the first time in genetic perturbation, simulating the distinct roles of genes with different biotypes in regulating cellular processes, while capturing implicit gene relationships through graph structure learning (GSL). We propose GRAPE, a heterogeneous graph neural network (HGNN) that leverages gene representations initialized with features from descriptions and sequences, models the distinct roles of genes with different biotypes, and dynamically refines the GRN through GSL. The results on publicly available datasets show that our method achieves state-of-the-art performance.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge