Jia Fu
SegRap2025: A Benchmark of Gross Tumor Volume and Lymph Node Clinical Target Volume Segmentation for Radiotherapy Planning of Nasopharyngeal Carcinoma
Jan 28, 2026Abstract:Accurate delineation of Gross Tumor Volume (GTV), Lymph Node Clinical Target Volume (LN CTV), and Organ-at-Risk (OAR) from Computed Tomography (CT) scans is essential for precise radiotherapy planning in Nasopharyngeal Carcinoma (NPC). Building upon SegRap2023, which focused on OAR and GTV segmentation using single-center paired non-contrast CT (ncCT) and contrast-enhanced CT (ceCT) scans, the SegRap2025 challenge aims to enhance the generalizability and robustness of segmentation models across imaging centers and modalities. SegRap2025 comprises two tasks: Task01 addresses GTV segmentation using paired CT from the SegRap2023 dataset, with an additional external testing set to evaluate cross-center generalization, and Task02 focuses on LN CTV segmentation using multi-center training data and an unseen external testing set, where each case contains paired CT scans or a single modality, emphasizing both cross-center and cross-modality robustness. This paper presents the challenge setup and provides a comprehensive analysis of the solutions submitted by ten participating teams. For GTV segmentation task, the top-performing models achieved average Dice Similarity Coefficient (DSC) of 74.61% and 56.79% on the internal and external testing cohorts, respectively. For LN CTV segmentation task, the highest average DSC values reached 60.24%, 60.50%, and 57.23% on paired CT, ceCT-only, and ncCT-only subsets, respectively. SegRap2025 establishes a large-scale multi-center, multi-modality benchmark for evaluating the generalization and robustness in radiotherapy target segmentation, providing valuable insights toward clinically applicable automated radiotherapy planning systems. The benchmark is available at: https://hilab-git.github.io/SegRap2025_Challenge.
DeepSynth-Eval: Objectively Evaluating Information Consolidation in Deep Survey Writing
Jan 07, 2026Abstract:The evolution of Large Language Models (LLMs) towards autonomous agents has catalyzed progress in Deep Research. While retrieval capabilities are well-benchmarked, the post-retrieval synthesis stage--where agents must digest massive amounts of context and consolidate fragmented evidence into coherent, long-form reports--remains under-evaluated due to the subjectivity of open-ended writing. To bridge this gap, we introduce DeepSynth-Eval, a benchmark designed to objectively evaluate information consolidation capabilities. We leverage high-quality survey papers as gold standards, reverse-engineering research requests and constructing "Oracle Contexts" from their bibliographies to isolate synthesis from retrieval noise. We propose a fine-grained evaluation protocol using General Checklists (for factual coverage) and Constraint Checklists (for structural organization), transforming subjective judgment into verifiable metrics. Experiments across 96 tasks reveal that synthesizing information from hundreds of references remains a significant challenge. Our results demonstrate that agentic plan-and-write workflows significantly outperform single-turn generation, effectively reducing hallucinations and improving adherence to complex structural constraints.
Klear-AgentForge: Forging Agentic Intelligence through Posttraining Scaling
Nov 08, 2025



Abstract:Despite the proliferation of powerful agentic models, the lack of critical post-training details hinders the development of strong counterparts in the open-source community. In this study, we present a comprehensive and fully open-source pipeline for training a high-performance agentic model for interacting with external tools and environments, named Klear-Qwen3-AgentForge, starting from the Qwen3-8B base model. We design effective supervised fine-tuning (SFT) with synthetic data followed by multi-turn reinforcement learning (RL) to unlock the potential for multiple diverse agentic tasks. We perform exclusive experiments on various agentic benchmarks in both tool use and coding domains. Klear-Qwen3-AgentForge-8B achieves state-of-the-art performance among LLMs of similar size and remains competitive with significantly larger models.
EReLiFM: Evidential Reliability-Aware Residual Flow Meta-Learning for Open-Set Domain Generalization under Noisy Labels
Oct 14, 2025



Abstract:Open-Set Domain Generalization (OSDG) aims to enable deep learning models to recognize unseen categories in new domains, which is crucial for real-world applications. Label noise hinders open-set domain generalization by corrupting source-domain knowledge, making it harder to recognize known classes and reject unseen ones. While existing methods address OSDG under Noisy Labels (OSDG-NL) using hyperbolic prototype-guided meta-learning, they struggle to bridge domain gaps, especially with limited clean labeled data. In this paper, we propose Evidential Reliability-Aware Residual Flow Meta-Learning (EReLiFM). We first introduce an unsupervised two-stage evidential loss clustering method to promote label reliability awareness. Then, we propose a residual flow matching mechanism that models structured domain- and category-conditioned residuals, enabling diverse and uncertainty-aware transfer paths beyond interpolation-based augmentation. During this meta-learning process, the model is optimized such that the update direction on the clean set maximizes the loss decrease on the noisy set, using pseudo labels derived from the most confident predicted class for supervision. Experimental results show that EReLiFM outperforms existing methods on OSDG-NL, achieving state-of-the-art performance. The source code is available at https://github.com/KPeng9510/ERELIFM.
THAT: Token-wise High-frequency Augmentation Transformer for Hyperspectral Pansharpening
Aug 11, 2025Abstract:Transformer-based methods have demonstrated strong potential in hyperspectral pansharpening by modeling long-range dependencies. However, their effectiveness is often limited by redundant token representations and a lack of multi-scale feature modeling. Hyperspectral images exhibit intrinsic spectral priors (e.g., abundance sparsity) and spatial priors (e.g., non-local similarity), which are critical for accurate reconstruction. From a spectral-spatial perspective, Vision Transformers (ViTs) face two major limitations: they struggle to preserve high-frequency components--such as material edges and texture transitions--and suffer from attention dispersion across redundant tokens. These issues stem from the global self-attention mechanism, which tends to dilute high-frequency signals and overlook localized details. To address these challenges, we propose the Token-wise High-frequency Augmentation Transformer (THAT), a novel framework designed to enhance hyperspectral pansharpening through improved high-frequency feature representation and token selection. Specifically, THAT introduces: (1) Pivotal Token Selective Attention (PTSA) to prioritize informative tokens and suppress redundancy; (2) a Multi-level Variance-aware Feed-forward Network (MVFN) to enhance high-frequency detail learning. Experiments on standard benchmarks show that THAT achieves state-of-the-art performance with improved reconstruction quality and efficiency. The source code is available at https://github.com/kailuo93/THAT.
RLEP: Reinforcement Learning with Experience Replay for LLM Reasoning
Jul 10, 2025



Abstract:Reinforcement learning (RL) for large language models is an energy-intensive endeavor: training can be unstable, and the policy may gradually drift away from its pretrained weights. We present \emph{RLEP}\, -- \,Reinforcement Learning with Experience rePlay\, -- \,a two-phase framework that first collects verified trajectories and then replays them during subsequent training. At every update step, the policy is optimized on mini-batches that blend newly generated rollouts with these replayed successes. By replaying high-quality examples, RLEP steers the model away from fruitless exploration, focuses learning on promising reasoning paths, and delivers both faster convergence and stronger final performance. On the Qwen2.5-Math-7B base model, RLEP reaches baseline peak accuracy with substantially fewer updates and ultimately surpasses it, improving accuracy on AIME-2024 from 38.2% to 39.9%, on AIME-2025 from 19.8% to 22.3%, and on AMC-2023 from 77.0% to 82.2%. Our code, datasets, and checkpoints are publicly available at https://github.com/Kwai-Klear/RLEP to facilitate reproducibility and further research.
DiffCAP: Diffusion-based Cumulative Adversarial Purification for Vision Language Models
Jun 04, 2025



Abstract:Vision Language Models (VLMs) have shown remarkable capabilities in multimodal understanding, yet their susceptibility to perturbations poses a significant threat to their reliability in real-world applications. Despite often being imperceptible to humans, these perturbations can drastically alter model outputs, leading to erroneous interpretations and decisions. This paper introduces DiffCAP, a novel diffusion-based purification strategy that can effectively neutralize adversarial corruptions in VLMs. We observe that adding minimal noise to an adversarially corrupted image significantly alters its latent embedding with respect to VLMs. Building on this insight, DiffCAP cumulatively injects random Gaussian noise into adversarially perturbed input data. This process continues until the embeddings of two consecutive noisy images reach a predefined similarity threshold, indicating a potential approach to neutralize the adversarial effect. Subsequently, a pretrained diffusion model is employed to denoise the stabilized image, recovering a clean representation suitable for the VLMs to produce an output. Through extensive experiments across six datasets with three VLMs under varying attack strengths in three task scenarios, we show that DiffCAP consistently outperforms existing defense techniques by a substantial margin. Notably, DiffCAP significantly reduces both hyperparameter tuning complexity and the required diffusion time, thereby accelerating the denoising process. Equipped with strong theoretical and empirical support, DiffCAP provides a robust and practical solution for securely deploying VLMs in adversarial environments.
Advances in Automated Fetal Brain MRI Segmentation and Biometry: Insights from the FeTA 2024 Challenge
May 05, 2025


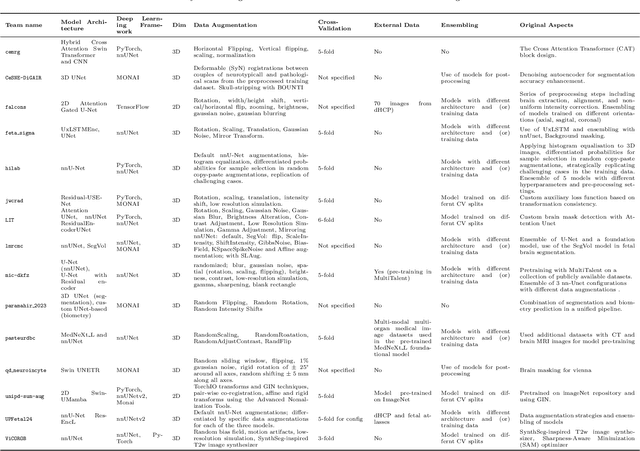
Abstract:Accurate fetal brain tissue segmentation and biometric analysis are essential for studying brain development in utero. The FeTA Challenge 2024 advanced automated fetal brain MRI analysis by introducing biometry prediction as a new task alongside tissue segmentation. For the first time, our diverse multi-centric test set included data from a new low-field (0.55T) MRI dataset. Evaluation metrics were also expanded to include the topology-specific Euler characteristic difference (ED). Sixteen teams submitted segmentation methods, most of which performed consistently across both high- and low-field scans. However, longitudinal trends indicate that segmentation accuracy may be reaching a plateau, with results now approaching inter-rater variability. The ED metric uncovered topological differences that were missed by conventional metrics, while the low-field dataset achieved the highest segmentation scores, highlighting the potential of affordable imaging systems when paired with high-quality reconstruction. Seven teams participated in the biometry task, but most methods failed to outperform a simple baseline that predicted measurements based solely on gestational age, underscoring the challenge of extracting reliable biometric estimates from image data alone. Domain shift analysis identified image quality as the most significant factor affecting model generalization, with super-resolution pipelines also playing a substantial role. Other factors, such as gestational age, pathology, and acquisition site, had smaller, though still measurable, effects. Overall, FeTA 2024 offers a comprehensive benchmark for multi-class segmentation and biometry estimation in fetal brain MRI, underscoring the need for data-centric approaches, improved topological evaluation, and greater dataset diversity to enable clinically robust and generalizable AI tools.
CLISC: Bridging clip and sam by enhanced cam for unsupervised brain tumor segmentation
Jan 27, 2025



Abstract:Brain tumor segmentation is important for diagnosis of the tumor, and current deep-learning methods rely on a large set of annotated images for training, with high annotation costs. Unsupervised segmentation is promising to avoid human annotations while the performance is often limited. In this study, we present a novel unsupervised segmentation approach that leverages the capabilities of foundation models, and it consists of three main steps: (1) A vision-language model (i.e., CLIP) is employed to obtain image-level pseudo-labels for training a classification network. Class Activation Mapping (CAM) is then employed to extract Regions of Interest (ROIs), where an adaptive masking-based data augmentation is used to enhance ROI identification.(2) The ROIs are used to generate bounding box and point prompts for the Segment Anything Model (SAM) to obtain segmentation pseudo-labels. (3) A 3D segmentation network is trained with the SAM-derived pseudo-labels, where low-quality pseudo-labels are filtered out in a self-learning process based on the similarity between the SAM's output and the network's prediction. Evaluation on the BraTS2020 dataset demonstrates that our approach obtained an average Dice Similarity Score (DSC) of 85.60%, outperforming five state-of-the-art unsupervised segmentation methods by more than 10 percentage points. Besides, our approach outperforms directly using SAM for zero-shot inference, and its performance is close to fully supervised learning.
DiffPAD: Denoising Diffusion-based Adversarial Patch Decontamination
Oct 31, 2024Abstract:In the ever-evolving adversarial machine learning landscape, developing effective defenses against patch attacks has become a critical challenge, necessitating reliable solutions to safeguard real-world AI systems. Although diffusion models have shown remarkable capacity in image synthesis and have been recently utilized to counter $\ell_p$-norm bounded attacks, their potential in mitigating localized patch attacks remains largely underexplored. In this work, we propose DiffPAD, a novel framework that harnesses the power of diffusion models for adversarial patch decontamination. DiffPAD first performs super-resolution restoration on downsampled input images, then adopts binarization, dynamic thresholding scheme and sliding window for effective localization of adversarial patches. Such a design is inspired by the theoretically derived correlation between patch size and diffusion restoration error that is generalized across diverse patch attack scenarios. Finally, DiffPAD applies inpainting techniques to the original input images with the estimated patch region being masked. By integrating closed-form solutions for super-resolution restoration and image inpainting into the conditional reverse sampling process of a pre-trained diffusion model, DiffPAD obviates the need for text guidance or fine-tuning. Through comprehensive experiments, we demonstrate that DiffPAD not only achieves state-of-the-art adversarial robustness against patch attacks but also excels in recovering naturalistic images without patch remnants.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge