Hongyu Guo
Rigidity-Aware Geometric Pretraining for Protein Design and Conformational Ensembles
Mar 02, 2026Abstract:Generative models have recently advanced $\textit{de novo}$ protein design by learning the statistical regularities of natural structures. However, current approaches face three key limitations: (1) Existing methods cannot jointly learn protein geometry and design tasks, where pretraining can be a solution; (2) Current pretraining methods mostly rely on local, non-rigid atomic representations for property prediction downstream tasks, limiting global geometric understanding for protein generation tasks; and (3) Existing approaches have yet to effectively model the rich dynamic and conformational information of protein structures. To overcome these issues, we introduce $\textbf{RigidSSL}$ ($\textit{Rigidity-Aware Self-Supervised Learning}$), a geometric pretraining framework that front-loads geometry learning prior to generative finetuning. Phase I (RigidSSL-Perturb) learns geometric priors from 432K structures from the AlphaFold Protein Structure Database with simulated perturbations. Phase II (RigidSSL-MD) refines these representations on 1.3K molecular dynamics trajectories to capture physically realistic transitions. Underpinning both phases is a bi-directional, rigidity-aware flow matching objective that jointly optimizes translational and rotational dynamics to maximize mutual information between conformations. Empirically, RigidSSL variants improve designability by up to 43\% while enhancing novelty and diversity in unconditional generation. Furthermore, RigidSSL-Perturb improves the success rate by 5.8\% in zero-shot motif scaffolding and RigidSSL-MD captures more biophysically realistic conformational ensembles in G protein-coupled receptor modeling. The code is available at: https://github.com/ZhanghanNi/RigidSSL.git.
PerturbDiff: Functional Diffusion for Single-Cell Perturbation Modeling
Feb 23, 2026Abstract:Building Virtual Cells that can accurately simulate cellular responses to perturbations is a long-standing goal in systems biology. A fundamental challenge is that high-throughput single-cell sequencing is destructive: the same cell cannot be observed both before and after a perturbation. Thus, perturbation prediction requires mapping unpaired control and perturbed populations. Existing models address this by learning maps between distributions, but typically assume a single fixed response distribution when conditioned on observed cellular context (e.g., cell type) and the perturbation type. In reality, responses vary systematically due to unobservable latent factors such as microenvironmental fluctuations and complex batch effects, forming a manifold of possible distributions for the same observed conditions. To account for this variability, we introduce PerturbDiff, which shifts modeling from individual cells to entire distributions. By embedding distributions as points in a Hilbert space, we define a diffusion-based generative process operating directly over probability distributions. This allows PerturbDiff to capture population-level response shifts across hidden factors. Benchmarks on established datasets show that PerturbDiff achieves state-of-the-art performance in single-cell response prediction and generalizes substantially better to unseen perturbations. See our project page (https://katarinayuan.github.io/PerturbDiff-ProjectPage/), where code and data will be made publicly available (https://github.com/DeepGraphLearning/PerturbDiff).
InertialAR: Autoregressive 3D Molecule Generation with Inertial Frames
Oct 31, 2025Abstract:Transformer-based autoregressive models have emerged as a unifying paradigm across modalities such as text and images, but their extension to 3D molecule generation remains underexplored. The gap stems from two fundamental challenges: (1) tokenizing molecules into a canonical 1D sequence of tokens that is invariant to both SE(3) transformations and atom index permutations, and (2) designing an architecture capable of modeling hybrid atom-based tokens that couple discrete atom types with continuous 3D coordinates. To address these challenges, we introduce InertialAR. InertialAR devises a canonical tokenization that aligns molecules to their inertial frames and reorders atoms to ensure SE(3) and permutation invariance. Moreover, InertialAR equips the attention mechanism with geometric awareness via geometric rotary positional encoding (GeoRoPE). In addition, it utilizes a hierarchical autoregressive paradigm to predict the next atom-based token, predicting the atom type first and then its 3D coordinates via Diffusion loss. Experimentally, InertialAR achieves state-of-the-art performance on 7 of the 10 evaluation metrics for unconditional molecule generation across QM9, GEOM-Drugs, and B3LYP. Moreover, it significantly outperforms strong baselines in controllable generation for targeted chemical functionality, attaining state-of-the-art results across all 5 metrics.
LacMaterial: Large Language Models as Analogical Chemists for Materials Discovery
Oct 25, 2025Abstract:Analogical reasoning, the transfer of relational structures across contexts (e.g., planet is to sun as electron is to nucleus), is fundamental to scientific discovery. Yet human insight is often constrained by domain expertise and surface-level biases, limiting access to deeper, structure-driven analogies both within and across disciplines. Large language models (LLMs), trained on vast cross-domain data, present a promising yet underexplored tool for analogical reasoning in science. Here, we demonstrate that LLMs can generate novel battery materials by (1) retrieving cross-domain analogs and analogy-guided exemplars to steer exploration beyond conventional dopant substitutions, and (2) constructing in-domain analogical templates from few labeled examples to guide targeted exploitation. These explicit analogical reasoning strategies yield candidates outside established compositional spaces and outperform standard prompting baselines. Our findings position LLMs as interpretable, expert-like hypothesis generators that leverage analogy-driven generalization for scientific innovation.
UniFGVC: Universal Training-Free Few-Shot Fine-Grained Vision Classification via Attribute-Aware Multimodal Retrieval
Aug 06, 2025Abstract:Few-shot fine-grained visual classification (FGVC) aims to leverage limited data to enable models to discriminate subtly distinct categories. Recent works mostly finetuned the pre-trained visual language models to achieve performance gain, yet suffering from overfitting and weak generalization. To deal with this, we introduce UniFGVC, a universal training-free framework that reformulates few-shot FGVC as multimodal retrieval. First, we propose the Category-Discriminative Visual Captioner (CDV-Captioner) to exploit the open-world knowledge of multimodal large language models (MLLMs) to generate a structured text description that captures the fine-grained attribute features distinguishing closely related classes. CDV-Captioner uses chain-of-thought prompting and visually similar reference images to reduce hallucination and enhance discrimination of generated captions. Using it we can convert each image into an image-description pair, enabling more comprehensive feature representation, and construct the multimodal category templates using few-shot samples for the subsequent retrieval pipeline. Then, off-the-shelf vision and text encoders embed query and template pairs, and FGVC is accomplished by retrieving the nearest template in the joint space. UniFGVC ensures broad compatibility with diverse MLLMs and encoders, offering reliable generalization and adaptability across few-shot FGVC scenarios. Extensive experiments on 12 FGVC benchmarks demonstrate its consistent superiority over prior few-shot CLIP-based methods and even several fully-supervised MLLMs-based approaches.
DisProtEdit: Exploring Disentangled Representations for Multi-Attribute Protein Editing
Jun 17, 2025Abstract:We introduce DisProtEdit, a controllable protein editing framework that leverages dual-channel natural language supervision to learn disentangled representations of structural and functional properties. Unlike prior approaches that rely on joint holistic embeddings, DisProtEdit explicitly separates semantic factors, enabling modular and interpretable control. To support this, we construct SwissProtDis, a large-scale multimodal dataset where each protein sequence is paired with two textual descriptions, one for structure and one for function, automatically decomposed using a large language model. DisProtEdit aligns protein and text embeddings using alignment and uniformity objectives, while a disentanglement loss promotes independence between structural and functional semantics. At inference time, protein editing is performed by modifying one or both text inputs and decoding from the updated latent representation. Experiments on protein editing and representation learning benchmarks demonstrate that DisProtEdit performs competitively with existing methods while providing improved interpretability and controllability. On a newly constructed multi-attribute editing benchmark, the model achieves a both-hit success rate of up to 61.7%, highlighting its effectiveness in coordinating simultaneous structural and functional edits.
OBELiX: A Curated Dataset of Crystal Structures and Experimentally Measured Ionic Conductivities for Lithium Solid-State Electrolytes
Feb 20, 2025



Abstract:Solid-state electrolyte batteries are expected to replace liquid electrolyte lithium-ion batteries in the near future thanks to their higher theoretical energy density and improved safety. However, their adoption is currently hindered by their lower effective ionic conductivity, a quantity that governs charge and discharge rates. Identifying highly ion-conductive materials using conventional theoretical calculations and experimental validation is both time-consuming and resource-intensive. While machine learning holds the promise to expedite this process, relevant ionic conductivity and structural data is scarce. Here, we present OBELiX, a domain-expert-curated database of $\sim$600 synthesized solid electrolyte materials and their experimentally measured room temperature ionic conductivities gathered from literature. Each material is described by their measured composition, space group and lattice parameters. A full-crystal description in the form of a crystallographic information file (CIF) is provided for ~320 structures for which atomic positions were available. We discuss various statistics and features of the dataset and provide training and testing splits that avoid data leakage. Finally, we benchmark seven existing ML models on the task of predicting ionic conductivity and discuss their performance. The goal of this work is to facilitate the use of machine learning for solid-state electrolyte materials discovery.
Baichuan-Omni-1.5 Technical Report
Jan 26, 2025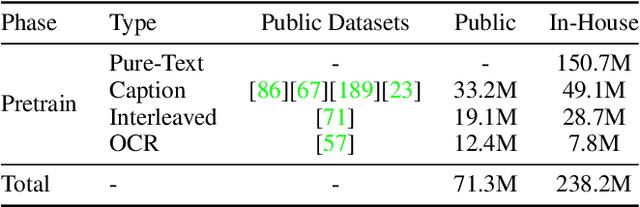
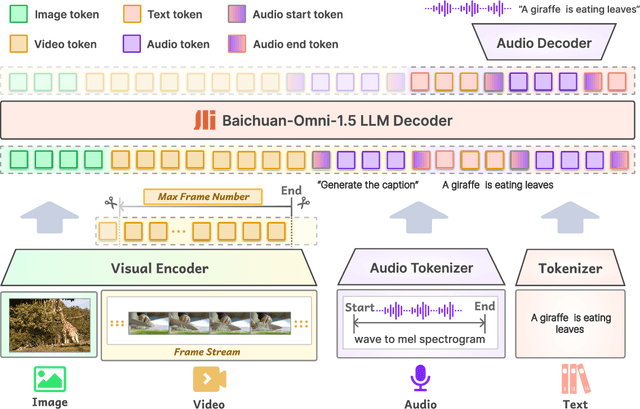
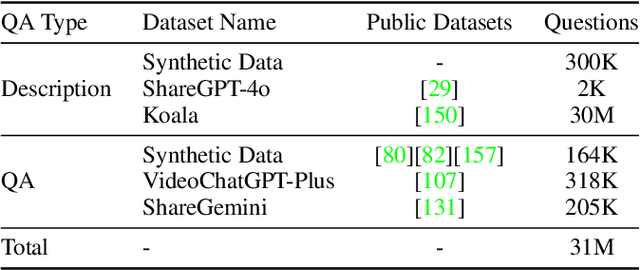
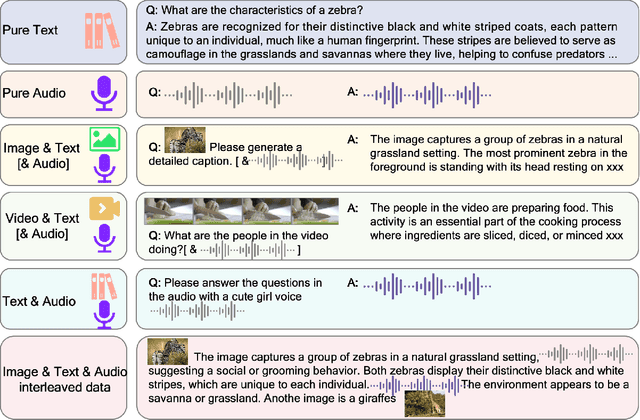
Abstract:We introduce Baichuan-Omni-1.5, an omni-modal model that not only has omni-modal understanding capabilities but also provides end-to-end audio generation capabilities. To achieve fluent and high-quality interaction across modalities without compromising the capabilities of any modality, we prioritized optimizing three key aspects. First, we establish a comprehensive data cleaning and synthesis pipeline for multimodal data, obtaining about 500B high-quality data (text, audio, and vision). Second, an audio-tokenizer (Baichuan-Audio-Tokenizer) has been designed to capture both semantic and acoustic information from audio, enabling seamless integration and enhanced compatibility with MLLM. Lastly, we designed a multi-stage training strategy that progressively integrates multimodal alignment and multitask fine-tuning, ensuring effective synergy across all modalities. Baichuan-Omni-1.5 leads contemporary models (including GPT4o-mini and MiniCPM-o 2.6) in terms of comprehensive omni-modal capabilities. Notably, it achieves results comparable to leading models such as Qwen2-VL-72B across various multimodal medical benchmarks.
A High Energy-Efficiency Multi-core Neuromorphic Architecture for Deep SNN Training
Dec 10, 2024



Abstract:There is a growing necessity for edge training to adapt to dynamically changing environment. Neuromorphic computing represents a significant pathway for high-efficiency intelligent computation in energy-constrained edges, but existing neuromorphic architectures lack the ability of directly training spiking neural networks (SNNs) based on backpropagation. We develop a multi-core neuromorphic architecture with Feedforward-Propagation, Back-Propagation, and Weight-Gradient engines in each core, supporting high efficient parallel computing at both the engine and core levels. It combines various data flows and sparse computation optimization by fully leveraging the sparsity in SNN training, obtaining a high energy efficiency of 1.05TFLOPS/W@ FP16 @ 28nm, 55 ~ 85% reduction of DRAM access compared to A100 GPU in SNN trainings, and a 20-core deep SNN training and a 5-worker federated learning on FPGAs. Our study develops the first multi-core neuromorphic architecture supporting the direct SNN training, facilitating the neuromorphic computing in edge-learnable applications.
Structure Language Models for Protein Conformation Generation
Oct 24, 2024



Abstract:Proteins adopt multiple structural conformations to perform their diverse biological functions, and understanding these conformations is crucial for advancing drug discovery. Traditional physics-based simulation methods often struggle with sampling equilibrium conformations and are computationally expensive. Recently, deep generative models have shown promise in generating protein conformations as a more efficient alternative. However, these methods predominantly rely on the diffusion process within a 3D geometric space, which typically centers around the vicinity of metastable states and is often inefficient in terms of runtime. In this paper, we introduce Structure Language Modeling (SLM) as a novel framework for efficient protein conformation generation. Specifically, the protein structures are first encoded into a compact latent space using a discrete variational auto-encoder, followed by conditional language modeling that effectively captures sequence-specific conformation distributions. This enables a more efficient and interpretable exploration of diverse ensemble modes compared to existing methods. Based on this general framework, we instantiate SLM with various popular LM architectures as well as proposing the ESMDiff, a novel BERT-like structure language model fine-tuned from ESM3 with masked diffusion. We verify our approach in various scenarios, including the equilibrium dynamics of BPTI, conformational change pairs, and intrinsically disordered proteins. SLM provides a highly efficient solution, offering a 20-100x speedup than existing methods in generating diverse conformations, shedding light on promising avenues for future research.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge