Lijun Liu
Co-Diffusion: An Affinity-Aware Two-Stage Latent Diffusion Framework for Generalizable Drug-Target Affinity Prediction
Mar 11, 2026Abstract:Predicting drug-target affinity is fundamental to virtual screening and lead optimization. However, existing deep models often suffer from representation collapse in stringent cold-start regimes, where the scarcity of labels and domain shifts prevent the learning of transferable pharmacophores and binding motifs. In this paper, we propose Co-Diffusion, a novel affinity-aware framework that redefines DTA prediction as a constrained latent denoising process to enhance generalization. Co-Diffusion employs a two-stage paradigm: Stage I establishes an affinity-steered latent manifold by aligning drug and target embeddings under an explicit supervised objective, ensuring that the latent space reflects the intrinsic binding landscape. Stage II introduces modality-specific latent diffusion as a stochastic perturb-and-denoise regularizer, forcing the model to recover consistent affinity semantics from noisy structural representations. This approach effectively mitigates the reconstruction-regression conflict common in generative DTA models. Theoretically, we show that Co-Diffusion maximizes a variational lower bound on the joint likelihood of drug structures, protein sequences, and binding strength. Extensive experiments across multiple benchmarks demonstrate that Co-Diffusion significantly outperforms state-of-the-art baselines, particularly yielding superior zero-shot generalization on unseen molecular scaffolds and novel protein families-paving a robust path for in silico drug prioritization in unexplored chemical spaces.
CORE-Acu: Structured Reasoning Traces and Knowledge Graph Safety Verification for Acupuncture Clinical Decision Support
Mar 09, 2026Abstract:Large language models (LLMs) show significant potential for clinical decision support (CDS), yet their black-box nature -- characterized by untraceable reasoning and probabilistic hallucinations -- poses severe challenges in acupuncture, a field demanding rigorous interpretability and safety. To address this, we propose CORE-Acu, a neuro-symbolic framework for acupuncture clinical decision support that integrates Structured Chain-of-Thought (S-CoT) with knowledge graph (KG) safety verification. First, we construct the first acupuncture Structured Reasoning Trace dataset and a schema-constrained fine-tuning framework. By enforcing an explicit causal chain from pattern identification to treatment principles, treatment plans, and acupoint selection, we transform implicit Traditional Chinese Medicine (TCM) reasoning into interpretable generation constraints, mitigating the opacity of LLM-based CDS. Furthermore, we construct a TCM safety knowledge graph and establish a ``Generate--Verify--Revise'' closed-loop inference system based on a Symbolic Veto Mechanism, employing deterministic rules to intercept hallucinations and enforce hard safety boundaries. Finally, we introduce the Lexicon-Matched Entity-Reweighted Loss (LMERL), which corrects terminology drift caused by the frequency--importance mismatch in general optimization by adaptively amplifying gradient contributions of high-risk entities during fine-tuning. Experiments on 1,000 held-out cases demonstrate CORE-Acu's superior entity fidelity and reasoning quality. Crucially, CORE-Acu achieved 0/1,000 observed safety violations (95\% CI: 0--0.37\%), whereas GPT-4o exhibited an 8.5\% violation rate under identical rules. These results establish CORE-Acu as a robust neuro-symbolic framework for acupuncture clinical decision support, guaranteeing both reasoning auditability and strict safety compliance.
SkinFlow: Efficient Information Transmission for Open Dermatological Diagnosis via Dynamic Visual Encoding and Staged RL
Jan 14, 2026Abstract:General-purpose Large Vision-Language Models (LVLMs), despite their massive scale, often falter in dermatology due to "diffuse attention" - the inability to disentangle subtle pathological lesions from background noise. In this paper, we challenge the assumption that parameter scaling is the only path to medical precision. We introduce SkinFlow, a framework that treats diagnosis as an optimization of visual information transmission efficiency. Our approach utilizes a Virtual-Width Dynamic Vision Encoder (DVE) to "unfold" complex pathological manifolds without physical parameter expansion, coupled with a two-stage Reinforcement Learning strategy. This strategy sequentially aligns explicit medical descriptions (Stage I) and reconstructs implicit diagnostic textures (Stage II) within a constrained semantic space. Furthermore, we propose a clinically grounded evaluation protocol that prioritizes diagnostic safety and hierarchical relevance over rigid label matching. Empirical results are compelling: our 7B model establishes a new state-of-the-art on the Fitzpatrick17k benchmark, achieving a +12.06% gain in Top-1 accuracy and a +28.57% boost in Top-6 accuracy over the massive general-purpose models (e.g., Qwen3VL-235B and GPT-5.2). These findings demonstrate that optimizing geometric capacity and information flow yields superior diagnostic reasoning compared to raw parameter scaling.
Efficient Medical VIE via Reinforcement Learning
Jun 16, 2025Abstract:Visual Information Extraction (VIE) converts unstructured document images into structured formats like JSON, critical for medical applications such as report analysis and online consultations. Traditional methods rely on OCR and language models, while end-to-end multimodal models offer direct JSON generation. However, domain-specific schemas and high annotation costs limit their effectiveness in medical VIE. We base our approach on the Reinforcement Learning with Verifiable Rewards (RLVR) framework to address these challenges using only 100 annotated samples. Our approach ensures dataset diversity, a balanced precision-recall reward mechanism to reduce hallucinations and improve field coverage, and innovative sampling strategies to enhance reasoning capabilities. Fine-tuning Qwen2.5-VL-7B with our RLVR method, we achieve state-of-the-art performance on medical VIE tasks, significantly improving F1, precision, and recall. While our models excel on tasks similar to medical datasets, performance drops on dissimilar tasks, highlighting the need for domain-specific optimization. Case studies further demonstrate the value of reasoning during training and inference for VIE.
Baichuan-Omni-1.5 Technical Report
Jan 26, 2025
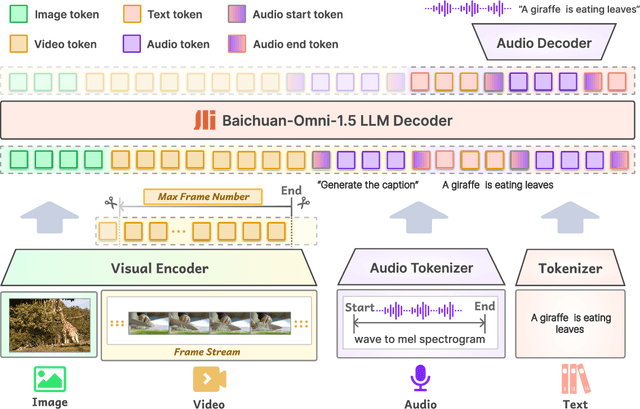
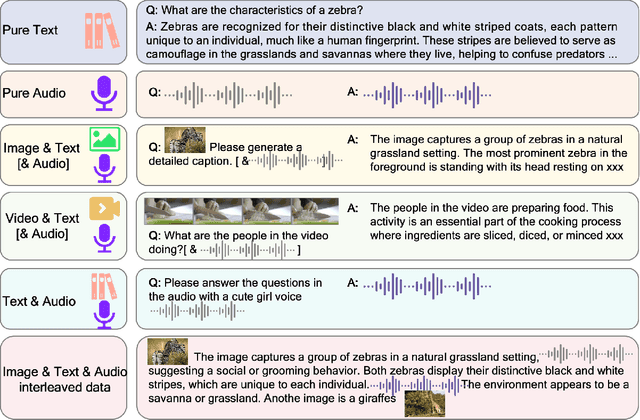
Abstract:We introduce Baichuan-Omni-1.5, an omni-modal model that not only has omni-modal understanding capabilities but also provides end-to-end audio generation capabilities. To achieve fluent and high-quality interaction across modalities without compromising the capabilities of any modality, we prioritized optimizing three key aspects. First, we establish a comprehensive data cleaning and synthesis pipeline for multimodal data, obtaining about 500B high-quality data (text, audio, and vision). Second, an audio-tokenizer (Baichuan-Audio-Tokenizer) has been designed to capture both semantic and acoustic information from audio, enabling seamless integration and enhanced compatibility with MLLM. Lastly, we designed a multi-stage training strategy that progressively integrates multimodal alignment and multitask fine-tuning, ensuring effective synergy across all modalities. Baichuan-Omni-1.5 leads contemporary models (including GPT4o-mini and MiniCPM-o 2.6) in terms of comprehensive omni-modal capabilities. Notably, it achieves results comparable to leading models such as Qwen2-VL-72B across various multimodal medical benchmarks.
AAVDiff: Experimental Validation of Enhanced Viability and Diversity in Recombinant Adeno-Associated Virus (AAV) Capsids through Diffusion Generation
Apr 17, 2024Abstract:Recombinant adeno-associated virus (rAAV) vectors have revolutionized gene therapy, but their broad tropism and suboptimal transduction efficiency limit their clinical applications. To overcome these limitations, researchers have focused on designing and screening capsid libraries to identify improved vectors. However, the large sequence space and limited resources present challenges in identifying viable capsid variants. In this study, we propose an end-to-end diffusion model to generate capsid sequences with enhanced viability. Using publicly available AAV2 data, we generated 38,000 diverse AAV2 viral protein (VP) sequences, and evaluated 8,000 for viral selection. The results attested the superiority of our model compared to traditional methods. Additionally, in the absence of AAV9 capsid data, apart from one wild-type sequence, we used the same model to directly generate a number of viable sequences with up to 9 mutations. we transferred the remaining 30,000 samples to the AAV9 domain. Furthermore, we conducted mutagenesis on AAV9 VP hypervariable regions VI and V, contributing to the continuous improvement of the AAV9 VP sequence. This research represents a significant advancement in the design and functional validation of rAAV vectors, offering innovative solutions to enhance specificity and transduction efficiency in gene therapy applications.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge