Guang Yang
Shanghai Key Laboratory of Magnetic Resonance, School of Physics and Electronic Science, East China Normal University, Shanghai, China, Institute of Magnetic Resonance and Molecular Imaging in Medicine, East China Normal University, Shanghai, China
LEGATO: Large-scale End-to-end Generalizable Approach to Typeset OMR
Jun 23, 2025Abstract:We propose Legato, a new end-to-end transformer model for optical music recognition (OMR). Legato is the first large-scale pretrained OMR model capable of recognizing full-page or multi-page typeset music scores and the first to generate documents in ABC notation, a concise, human-readable format for symbolic music. Bringing together a pretrained vision encoder with an ABC decoder trained on a dataset of more than 214K images, our model exhibits the strong ability to generalize across various typeset scores. We conduct experiments on a range of datasets and demonstrate that our model achieves state-of-the-art performance. Given the lack of a standardized evaluation for end-to-end OMR, we comprehensively compare our model against the previous state of the art using a diverse set of metrics.
An LLM-as-Judge Metric for Bridging the Gap with Human Evaluation in SE Tasks
May 27, 2025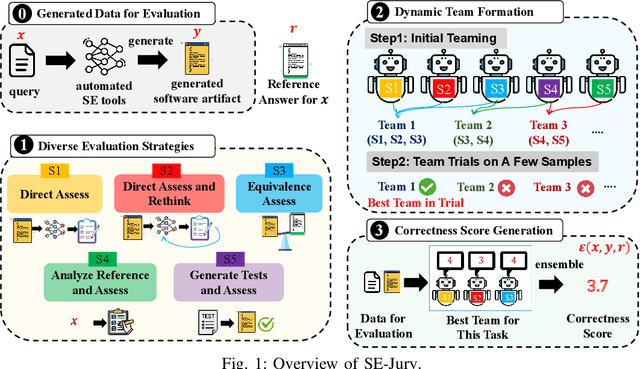
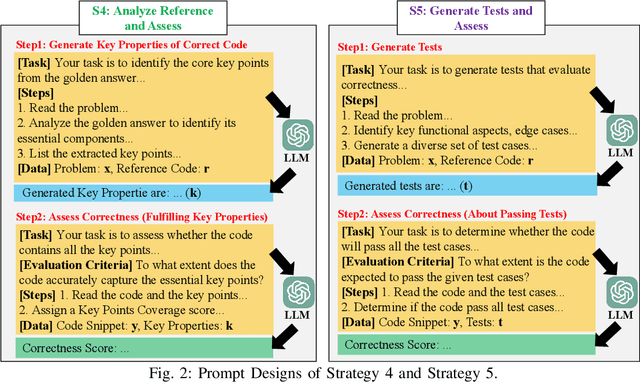
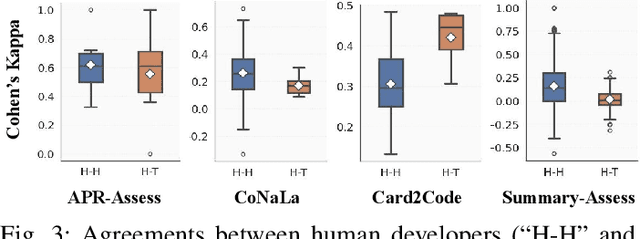
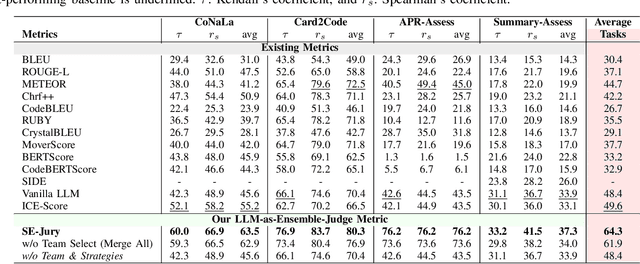
Abstract:Large Language Models (LLMs) and other automated techniques have been increasingly used to support software developers by generating software artifacts such as code snippets, patches, and comments. However, accurately assessing the correctness of these generated artifacts remains a significant challenge. On one hand, human evaluation provides high accuracy but is labor-intensive and lacks scalability. On the other hand, other existing automatic evaluation metrics are scalable and require minimal human effort, but they often fail to accurately reflect the actual correctness of generated software artifacts. In this paper, we present SWE-Judge, the first evaluation metric for LLM-as-Ensemble-Judge specifically designed to accurately assess the correctness of generated software artifacts. SWE-Judge first defines five distinct evaluation strategies, each implemented as an independent judge. A dynamic team selection mechanism then identifies the most appropriate subset of judges to produce a final correctness score through ensembling. We evaluate SWE-Judge across a diverse set of software engineering (SE) benchmarks, including CoNaLa, Card2Code, HumanEval-X, APPS, APR-Assess, and Summary-Assess. These benchmarks span three SE tasks: code generation, automated program repair, and code summarization. Experimental results demonstrate that SWE-Judge consistently achieves a higher correlation with human judgments, with improvements ranging from 5.9% to 183.8% over existing automatic metrics. Furthermore, SWE-Judge reaches agreement levels with human annotators that are comparable to inter-annotator agreement in code generation and program repair tasks. These findings underscore SWE-Judge's potential as a scalable and reliable alternative to human evaluation.
CODE-DITING: A Reasoning-Based Metric for Functional Alignment in Code Evaluation
May 26, 2025Abstract:Trustworthy evaluation methods for code snippets play a crucial role in neural code generation. Traditional methods, which either rely on reference solutions or require executable test cases, have inherent limitation in flexibility and scalability. The recent LLM-as-Judge methodology offers a promising alternative by directly evaluating functional consistency between the problem description and the generated code. To systematically understand the landscape of these LLM-as-Judge methods, we conduct a comprehensive empirical study across three diverse datasets. Our investigation reveals the pros and cons of two categories of LLM-as-Judge methods: the methods based on general foundation models can achieve good performance but require complex prompts and lack explainability, while the methods based on reasoning foundation models provide better explainability with simpler prompts but demand substantial computational resources due to their large parameter sizes. To address these limitations, we propose CODE-DITING, a novel code evaluation method that balances accuracy, efficiency and explainability. We develop a data distillation framework that effectively transfers reasoning capabilities from DeepSeek-R1671B to our CODE-DITING 1.5B and 7B models, significantly enhancing evaluation explainability and reducing the computational cost. With the majority vote strategy in the inference process, CODE-DITING 1.5B outperforms all models with the same magnitude of parameters and achieves performance which would normally exhibit in a model with 5 times of parameter scale. CODE-DITING 7B surpasses GPT-4o and DeepSeek-V3 671B, even though it only uses 1% of the parameter volume of these large models. Further experiments show that CODEDITING is robust to preference leakage and can serve as a promising alternative for code evaluation.
MMMG: a Comprehensive and Reliable Evaluation Suite for Multitask Multimodal Generation
May 23, 2025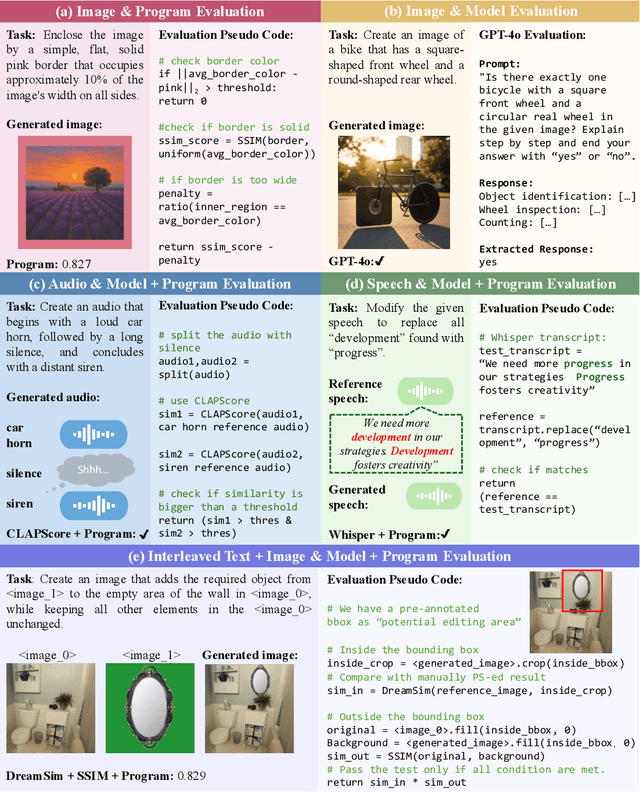
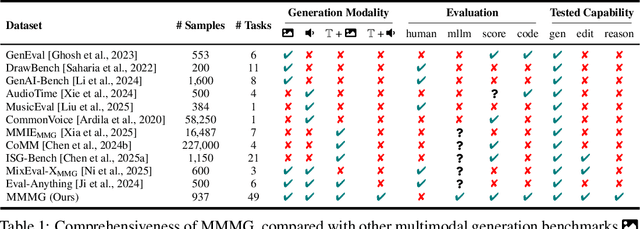
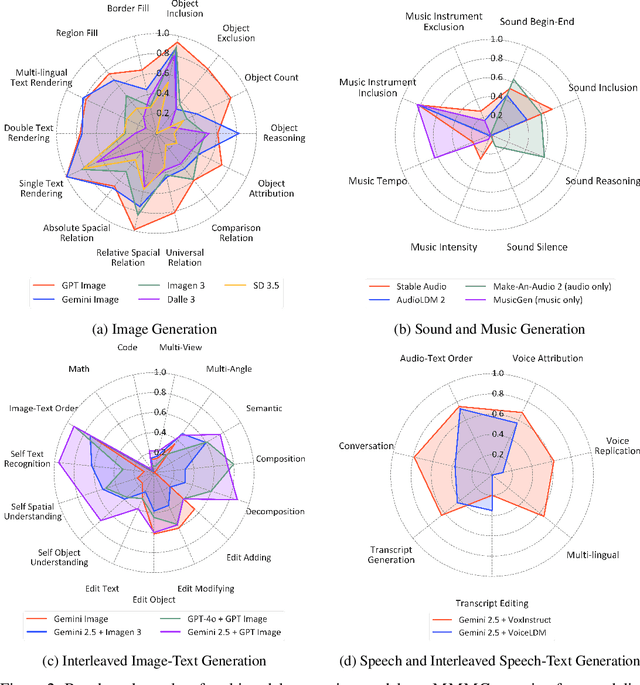
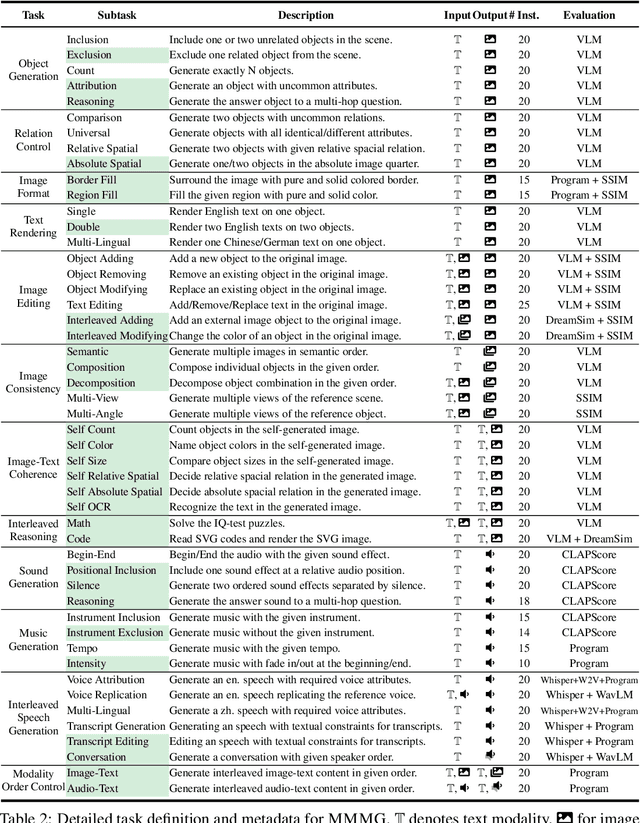
Abstract:Automatically evaluating multimodal generation presents a significant challenge, as automated metrics often struggle to align reliably with human evaluation, especially for complex tasks that involve multiple modalities. To address this, we present MMMG, a comprehensive and human-aligned benchmark for multimodal generation across 4 modality combinations (image, audio, interleaved text and image, interleaved text and audio), with a focus on tasks that present significant challenges for generation models, while still enabling reliable automatic evaluation through a combination of models and programs. MMMG encompasses 49 tasks (including 29 newly developed ones), each with a carefully designed evaluation pipeline, and 937 instructions to systematically assess reasoning, controllability, and other key capabilities of multimodal generation models. Extensive validation demonstrates that MMMG is highly aligned with human evaluation, achieving an average agreement of 94.3%. Benchmarking results on 24 multimodal generation models reveal that even though the state-of-the-art model, GPT Image, achieves 78.3% accuracy for image generation, it falls short on multimodal reasoning and interleaved generation. Furthermore, results suggest considerable headroom for improvement in audio generation, highlighting an important direction for future research.
The Staircase of Ethics: Probing LLM Value Priorities through Multi-Step Induction to Complex Moral Dilemmas
May 23, 2025Abstract:Ethical decision-making is a critical aspect of human judgment, and the growing use of LLMs in decision-support systems necessitates a rigorous evaluation of their moral reasoning capabilities. However, existing assessments primarily rely on single-step evaluations, failing to capture how models adapt to evolving ethical challenges. Addressing this gap, we introduce the Multi-step Moral Dilemmas (MMDs), the first dataset specifically constructed to evaluate the evolving moral judgments of LLMs across 3,302 five-stage dilemmas. This framework enables a fine-grained, dynamic analysis of how LLMs adjust their moral reasoning across escalating dilemmas. Our evaluation of nine widely used LLMs reveals that their value preferences shift significantly as dilemmas progress, indicating that models recalibrate moral judgments based on scenario complexity. Furthermore, pairwise value comparisons demonstrate that while LLMs often prioritize the value of care, this value can sometimes be superseded by fairness in certain contexts, highlighting the dynamic and context-dependent nature of LLM ethical reasoning. Our findings call for a shift toward dynamic, context-aware evaluation paradigms, paving the way for more human-aligned and value-sensitive development of LLMs.
Dynamic Dual Buffer with Divide-and-Conquer Strategy for Online Continual Learning
May 23, 2025Abstract:Online Continual Learning (OCL) presents a complex learning environment in which new data arrives in a batch-to-batch online format, and the risk of catastrophic forgetting can significantly impair model efficacy. In this study, we address OCL by introducing an innovative memory framework that incorporates a short-term memory system to retain dynamic information and a long-term memory system to archive enduring knowledge. Specifically, the long-term memory system comprises a collection of sub-memory buffers, each linked to a cluster prototype and designed to retain data samples from distinct categories. We propose a novel $K$-means-based sample selection method to identify cluster prototypes for each encountered category. To safeguard essential and critical samples, we introduce a novel memory optimisation strategy that selectively retains samples in the appropriate sub-memory buffer by evaluating each cluster prototype against incoming samples through an optimal transportation mechanism. This approach specifically promotes each sub-memory buffer to retain data samples that exhibit significant discrepancies from the corresponding cluster prototype, thereby ensuring the preservation of semantically rich information. In addition, we propose a novel Divide-and-Conquer (DAC) approach that formulates the memory updating as an optimisation problem and divides it into several subproblems. As a result, the proposed DAC approach can solve these subproblems separately and thus can significantly reduce computations of the proposed memory updating process. We conduct a series of experiments across standard and imbalanced learning settings, and the empirical findings indicate that the proposed memory framework achieves state-of-the-art performance in both learning contexts.
Enhancing Diffusion-Weighted Images (DWI) for Diffusion MRI: Is it Enough without Non-Diffusion-Weighted B=0 Reference?
May 19, 2025Abstract:Diffusion MRI (dMRI) is essential for studying brain microstructure, but high-resolution imaging remains challenging due to the inherent trade-offs between acquisition time and signal-to-noise ratio (SNR). Conventional methods often optimize only the diffusion-weighted images (DWIs) without considering their relationship with the non-diffusion-weighted (b=0) reference images. However, calculating diffusion metrics, such as the apparent diffusion coefficient (ADC) and diffusion tensor with its derived metrics like fractional anisotropy (FA) and mean diffusivity (MD), relies on the ratio between each DWI and the b=0 image, which is crucial for clinical observation and diagnostics. In this study, we demonstrate that solely enhancing DWIs using a conventional pixel-wise mean squared error (MSE) loss is insufficient, as the error in ratio between generated DWIs and b=0 diverges. We propose a novel ratio loss, defined as the MSE loss between the predicted and ground-truth log of DWI/b=0 ratios. Our results show that incorporating the ratio loss significantly improves the convergence of this ratio error, achieving lower ratio MSE and slightly enhancing the peak signal-to-noise ratio (PSNR) of generated DWIs. This leads to improved dMRI super-resolution and better preservation of b=0 ratio-based features for the derivation of diffusion metrics.
Multimodal Cancer Survival Analysis via Hypergraph Learning with Cross-Modality Rebalance
May 17, 2025Abstract:Multimodal pathology-genomic analysis has become increasingly prominent in cancer survival prediction. However, existing studies mainly utilize multi-instance learning to aggregate patch-level features, neglecting the information loss of contextual and hierarchical details within pathology images. Furthermore, the disparity in data granularity and dimensionality between pathology and genomics leads to a significant modality imbalance. The high spatial resolution inherent in pathology data renders it a dominant role while overshadowing genomics in multimodal integration. In this paper, we propose a multimodal survival prediction framework that incorporates hypergraph learning to effectively capture both contextual and hierarchical details from pathology images. Moreover, it employs a modality rebalance mechanism and an interactive alignment fusion strategy to dynamically reweight the contributions of the two modalities, thereby mitigating the pathology-genomics imbalance. Quantitative and qualitative experiments are conducted on five TCGA datasets, demonstrating that our model outperforms advanced methods by over 3.4\% in C-Index performance.
Decoupling Multi-Contrast Super-Resolution: Pairing Unpaired Synthesis with Implicit Representations
May 09, 2025Abstract:Magnetic Resonance Imaging (MRI) is critical for clinical diagnostics but is often limited by long acquisition times and low signal-to-noise ratios, especially in modalities like diffusion and functional MRI. The multi-contrast nature of MRI presents a valuable opportunity for cross-modal enhancement, where high-resolution (HR) modalities can serve as references to boost the quality of their low-resolution (LR) counterparts-motivating the development of Multi-Contrast Super-Resolution (MCSR) techniques. Prior work has shown that leveraging complementary contrasts can improve SR performance; however, effective feature extraction and fusion across modalities with varying resolutions remains a major challenge. Moreover, existing MCSR methods often assume fixed resolution settings and all require large, perfectly paired training datasets-conditions rarely met in real-world clinical environments. To address these challenges, we propose a novel Modular Multi-Contrast Super-Resolution (MCSR) framework that eliminates the need for paired training data and supports arbitrary upscaling. Our method decouples the MCSR task into two stages: (1) Unpaired Cross-Modal Synthesis (U-CMS), which translates a high-resolution reference modality into a synthesized version of the target contrast, and (2) Unsupervised Super-Resolution (U-SR), which reconstructs the final output using implicit neural representations (INRs) conditioned on spatial coordinates. This design enables scale-agnostic and anatomically faithful reconstruction by bridging un-paired cross-modal synthesis with unsupervised resolution enhancement. Experiments show that our method achieves superior performance at 4x and 8x upscaling, with improved fidelity and anatomical consistency over existing baselines. Our framework demonstrates strong potential for scalable, subject-specific, and data-efficient MCSR in real-world clinical settings.
PoseX: AI Defeats Physics Approaches on Protein-Ligand Cross Docking
May 03, 2025Abstract:Recently, significant progress has been made in protein-ligand docking, especially in modern deep learning methods, and some benchmarks were proposed, e.g., PoseBench, Plinder. However, these benchmarks suffer from less practical evaluation setups (e.g., blind docking, self docking), or heavy framework that involves training, raising challenges to assess docking methods efficiently. To fill this gap, we proposed PoseX, an open-source benchmark focusing on self-docking and cross-docking, to evaluate the algorithmic advances practically and comprehensively. Specifically, first, we curate a new evaluation dataset with 718 entries for self docking and 1,312 for cross docking; second, we incorporate 22 docking methods across three methodological categories, including (1) traditional physics-based methods (e.g., Schr\"odinger Glide), (2) AI docking methods (e.g., DiffDock), (3) AI co-folding methods (e.g., AlphaFold3); third, we design a relaxation method as post-processing to minimize conformation energy and refine binding pose; fourth, we released a leaderboard to rank submitted models in real time. We draw some key insights via extensive experiments: (1) AI-based approaches have already surpassed traditional physics-based approaches in overall docking accuracy (RMSD). The longstanding generalization issues that have plagued AI molecular docking have been significantly alleviated in the latest models. (2) The stereochemical deficiencies of AI-based approaches can be greatly alleviated with post-processing relaxation. Combining AI docking methods with the enhanced relaxation method achieves the best performance to date. (3) AI co-folding methods commonly face ligand chirality issues, which cannot be resolved by relaxation. The code, curated dataset and leaderboard are released at https://github.com/CataAI/PoseX.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge