Lucy Lu Wang
ReFinE: Streamlining UI Mockup Iteration with Research Findings
Apr 06, 2026Abstract:Although HCI research papers offer valuable design insights, designers often struggle to apply them in design workflows due to difficulties in finding relevant literature, understanding technical jargon, the lack of contextualization, and limited actionability. To address these challenges, we present ReFinE, a Figma plugin that supports real-time design iteration by surfacing contextualized insights from research papers. ReFinE identifies and synthesizes design implications from HCI literature relevant to the mockup's design context, and tailors this research evidence to a specific design mockup by providing actionable visual guidance on how to update the mockup. To assess the system's effectiveness, we conducted a technical evaluation and a user study. Results show that ReFinE effectively synthesizes and contextualizes design implications, reducing cognitive load and improving designers' ability to integrate research evidence into UI mockups. This work contributes to bridging the gap between research and design practice by presenting a tool for embedding scholarly insights into the UI design process.
Clarify or Answer: Reinforcement Learning for Agentic VQA with Context Under-specification
Jan 23, 2026Abstract:Real-world visual question answering (VQA) is often context-dependent: an image-question pair may be under-specified, such that the correct answer depends on external information that is not observable in the image. In such cases, directly answering can lead to confident but incorrect predictions. We propose CoA(Clarify-or-Answer), an ask-or-answer agent that separately models the decision to ask or answer, and what to ask if needed. CoA first determines whether clarification is necessary; if so, it asks a single focused question and then incorporates the response to produce the final answer. We introduce CONTEXTCLARIFY with a set of ambiguous VQA questions and the contrast set that is non-ambiguous. We further introduce GRPO-CR (Clarification Reasoning), a reinforcement learning approach that optimizes clarification question generation with multiple reward signals encouraging well-formed, focused, non-trivial questions that resolve ambiguity. Across three VLLMs and three datasets, CoA achieves consistent improvements at both the module and system levels, improving end-to-end VQA accuracy by an average of +15.3 points (83%) over prompting-based baselines
Leveraging Hierarchical Organization for Medical Multi-document Summarization
Oct 27, 2025Abstract:Medical multi-document summarization (MDS) is a complex task that requires effectively managing cross-document relationships. This paper investigates whether incorporating hierarchical structures in the inputs of MDS can improve a model's ability to organize and contextualize information across documents compared to traditional flat summarization methods. We investigate two ways of incorporating hierarchical organization across three large language models (LLMs), and conduct comprehensive evaluations of the resulting summaries using automated metrics, model-based metrics, and domain expert evaluation of preference, understandability, clarity, complexity, relevance, coverage, factuality, and coherence. Our results show that human experts prefer model-generated summaries over human-written summaries. Hierarchical approaches generally preserve factuality, coverage, and coherence of information, while also increasing human preference for summaries. Additionally, we examine whether simulated judgments from GPT-4 align with human judgments, finding higher agreement along more objective evaluation facets. Our findings demonstrate that hierarchical structures can improve the clarity of medical summaries generated by models while maintaining content coverage, providing a practical way to improve human preference for generated summaries.
MMMG: a Comprehensive and Reliable Evaluation Suite for Multitask Multimodal Generation
May 23, 2025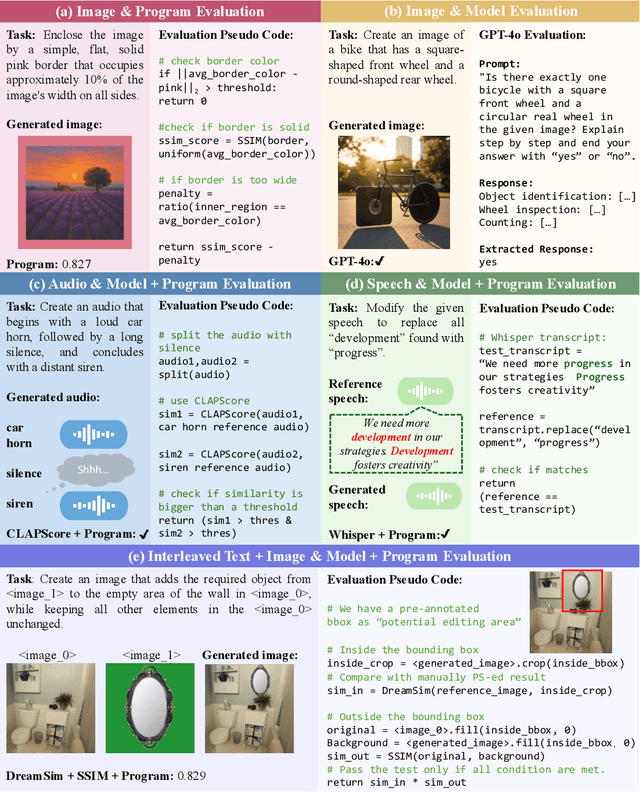
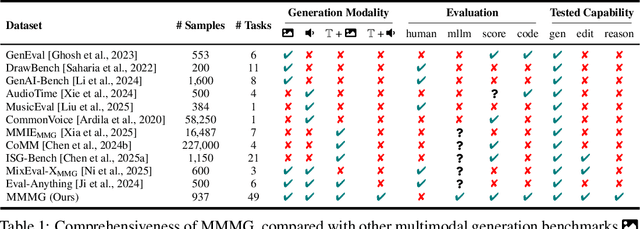
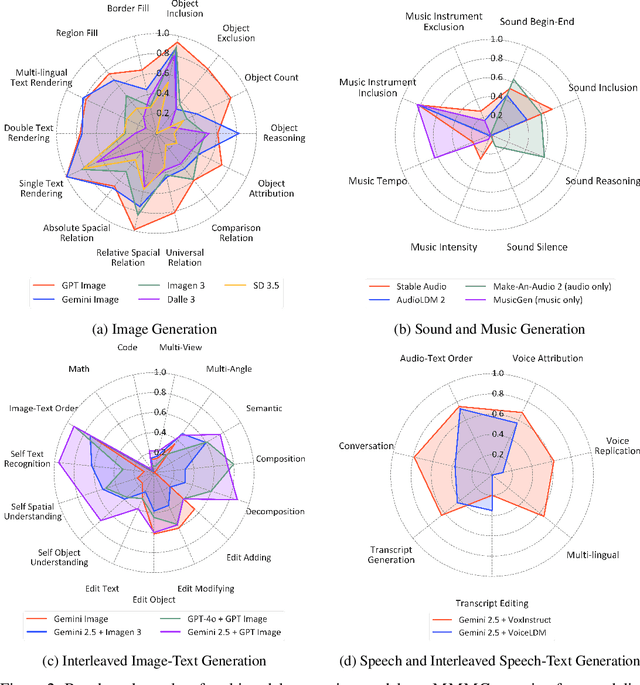
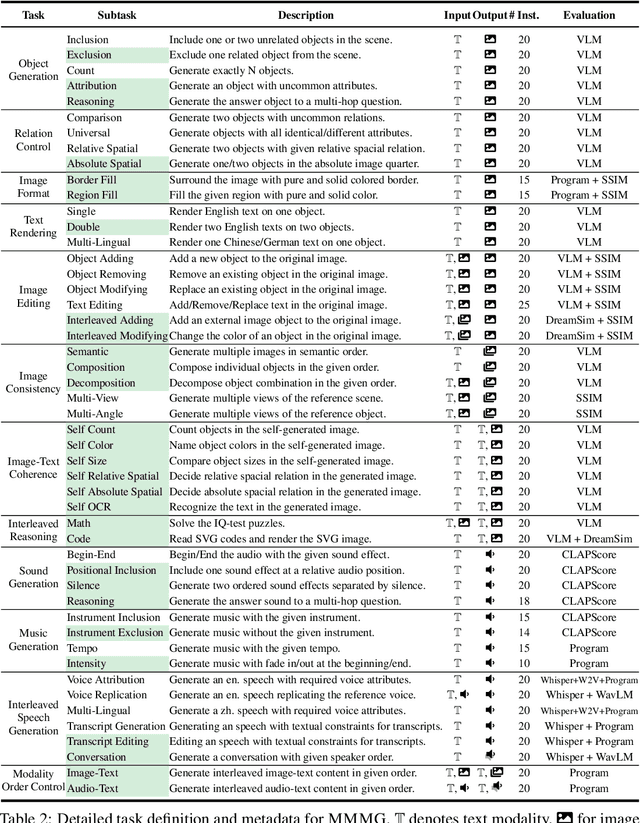
Abstract:Automatically evaluating multimodal generation presents a significant challenge, as automated metrics often struggle to align reliably with human evaluation, especially for complex tasks that involve multiple modalities. To address this, we present MMMG, a comprehensive and human-aligned benchmark for multimodal generation across 4 modality combinations (image, audio, interleaved text and image, interleaved text and audio), with a focus on tasks that present significant challenges for generation models, while still enabling reliable automatic evaluation through a combination of models and programs. MMMG encompasses 49 tasks (including 29 newly developed ones), each with a carefully designed evaluation pipeline, and 937 instructions to systematically assess reasoning, controllability, and other key capabilities of multimodal generation models. Extensive validation demonstrates that MMMG is highly aligned with human evaluation, achieving an average agreement of 94.3%. Benchmarking results on 24 multimodal generation models reveal that even though the state-of-the-art model, GPT Image, achieves 78.3% accuracy for image generation, it falls short on multimodal reasoning and interleaved generation. Furthermore, results suggest considerable headroom for improvement in audio generation, highlighting an important direction for future research.
FACTS&EVIDENCE: An Interactive Tool for Transparent Fine-Grained Factual Verification of Machine-Generated Text
Mar 19, 2025



Abstract:With the widespread consumption of AI-generated content, there has been an increased focus on developing automated tools to verify the factual accuracy of such content. However, prior research and tools developed for fact verification treat it as a binary classification or a linear regression problem. Although this is a useful mechanism as part of automatic guardrails in systems, we argue that such tools lack transparency in the prediction reasoning and diversity in source evidence to provide a trustworthy user experience. We develop Facts&Evidence - an interactive and transparent tool for user-driven verification of complex text. The tool facilitates the intricate decision-making involved in fact-verification, presenting its users a breakdown of complex input texts to visualize the credibility of individual claims along with an explanation of model decisions and attribution to multiple, diverse evidence sources. Facts&Evidence aims to empower consumers of machine-generated text and give them agency to understand, verify, selectively trust and use such text.
Explainable AI for Clinical Outcome Prediction: A Survey of Clinician Perceptions and Preferences
Feb 27, 2025Abstract:Explainable AI (XAI) techniques are necessary to help clinicians make sense of AI predictions and integrate predictions into their decision-making workflow. In this work, we conduct a survey study to understand clinician preference among different XAI techniques when they are used to interpret model predictions over text-based EHR data. We implement four XAI techniques (LIME, Attention-based span highlights, exemplar patient retrieval, and free-text rationales generated by LLMs) on an outcome prediction model that uses ICU admission notes to predict a patient's likelihood of experiencing in-hospital mortality. Using these XAI implementations, we design and conduct a survey study of 32 practicing clinicians, collecting their feedback and preferences on the four techniques. We synthesize our findings into a set of recommendations describing when each of the XAI techniques may be more appropriate, their potential limitations, as well as recommendations for improvement.
Varying Shades of Wrong: Aligning LLMs with Wrong Answers Only
Oct 14, 2024



Abstract:In the absence of abundant reliable annotations for challenging tasks and contexts, how can we expand the frontier of LLM capabilities with potentially wrong answers? We focus on two research questions: (1) Can LLMs generate reliable preferences among wrong options? And if so, (2) Would alignment with such wrong-over-wrong preferences be helpful? We employ methods based on self-consistency, token probabilities, and LLM-as-a-judge to elicit wrong-over-wrong preferences, and fine-tune language models with preference optimization approaches using these synthesized preferences. Extensive experiments with seven LLMs and eight datasets demonstrate that (1) LLMs do have preliminary capability in distinguishing various shades of wrong, achieving up to 20.9% higher performance than random guess; (2) Alignment with wrong-over-wrong preferences helps LLMs to produce less wrong and sometimes even outright correct answers, while overall improving model calibration.
Know Your Limits: A Survey of Abstention in Large Language Models
Aug 08, 2024



Abstract:Abstention, the refusal of large language models (LLMs) to provide an answer, is increasingly recognized for its potential to mitigate hallucinations and enhance safety in LLM systems. In this survey, we introduce a framework to examine abstention from three perspectives: the query, the model, and human values. We organize the literature on abstention methods, benchmarks, and evaluation metrics using this framework, and discuss merits and limitations of prior work. We further identify and motivate areas for future work, centered around whether abstention can be achieved as a meta-capability that transcends specific tasks or domains, while still providing opportunities to optimize abstention abilities based on context.
The Art of Refusal: A Survey of Abstention in Large Language Models
Jul 25, 2024



Abstract:Abstention, the refusal of large language models (LLMs) to provide an answer, is increasingly recognized for its potential to mitigate hallucinations and enhance safety in building LLM systems. In this survey, we introduce a framework to examine abstention behavior from three perspectives: the query, the model, and human values. We review the literature on abstention methods (categorized based on the development stages of LLMs), benchmarks, and evaluation metrics, and discuss the merits and limitations of prior work. We further identify and motivate areas for future research, such as encouraging the study of abstention as a meta-capability across tasks and customizing abstention abilities based on context. In doing so, we aim to broaden the scope and impact of abstention methodologies in AI systems.
CHIME: LLM-Assisted Hierarchical Organization of Scientific Studies for Literature Review Support
Jul 23, 2024



Abstract:Literature review requires researchers to synthesize a large amount of information and is increasingly challenging as the scientific literature expands. In this work, we investigate the potential of LLMs for producing hierarchical organizations of scientific studies to assist researchers with literature review. We define hierarchical organizations as tree structures where nodes refer to topical categories and every node is linked to the studies assigned to that category. Our naive LLM-based pipeline for hierarchy generation from a set of studies produces promising yet imperfect hierarchies, motivating us to collect CHIME, an expert-curated dataset for this task focused on biomedicine. Given the challenging and time-consuming nature of building hierarchies from scratch, we use a human-in-the-loop process in which experts correct errors (both links between categories and study assignment) in LLM-generated hierarchies. CHIME contains 2,174 LLM-generated hierarchies covering 472 topics, and expert-corrected hierarchies for a subset of 100 topics. Expert corrections allow us to quantify LLM performance, and we find that while they are quite good at generating and organizing categories, their assignment of studies to categories could be improved. We attempt to train a corrector model with human feedback which improves study assignment by 12.6 F1 points. We release our dataset and models to encourage research on developing better assistive tools for literature review.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge