Dragomir Radev
EHRKit: A Python Natural Language Processing Toolkit for Electronic Health Record Texts
Apr 13, 2022
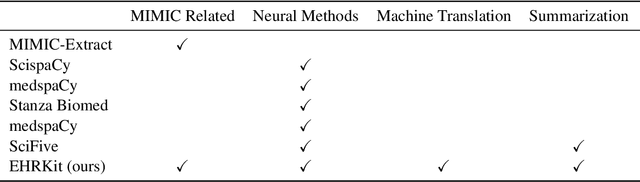

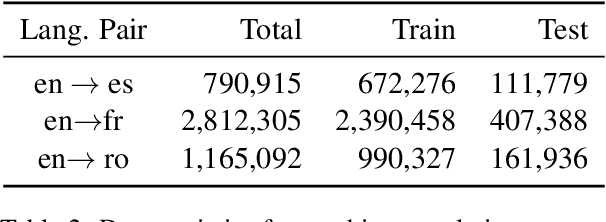
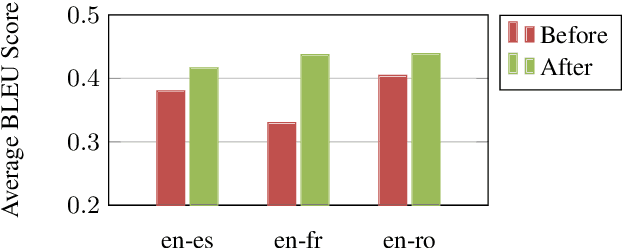
Abstract:The Electronic Health Record (EHR) is an essential part of the modern medical system and impacts healthcare delivery, operations, and research. Unstructured text is attracting much attention despite structured information in the EHRs and has become an exciting research field. The success of the recent neural Natural Language Processing (NLP) method has led to a new direction for processing unstructured clinical notes. In this work, we create a python library for clinical texts, EHRKit. This library contains two main parts: MIMIC-III-specific functions and tasks specific functions. The first part introduces a list of interfaces for accessing MIMIC-III NOTEEVENTS data, including basic search, information retrieval, and information extraction. The second part integrates many third-party libraries for up to 12 off-shelf NLP tasks such as named entity recognition, summarization, machine translation, etc.
BRIO: Bringing Order to Abstractive Summarization
Mar 31, 2022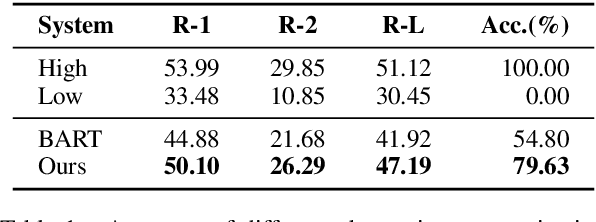
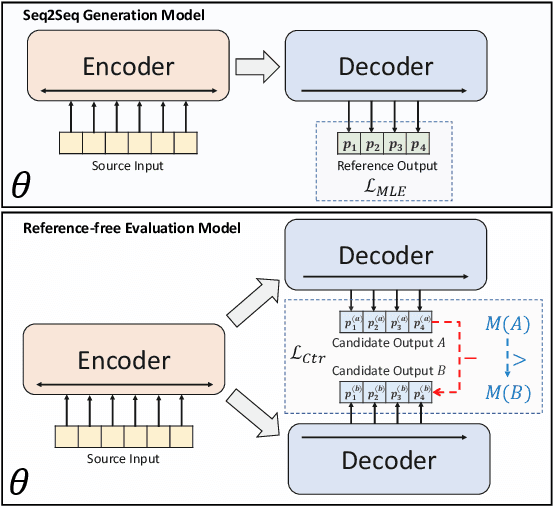
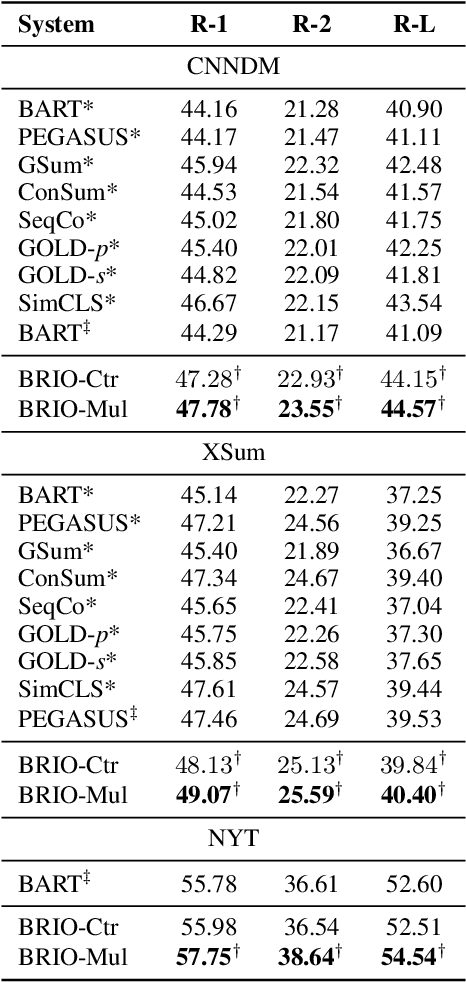
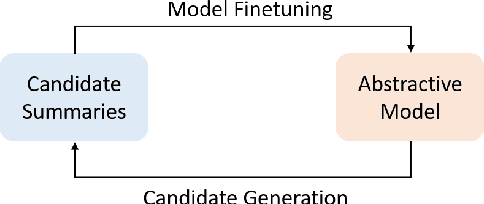
Abstract:Abstractive summarization models are commonly trained using maximum likelihood estimation, which assumes a deterministic (one-point) target distribution in which an ideal model will assign all the probability mass to the reference summary. This assumption may lead to performance degradation during inference, where the model needs to compare several system-generated (candidate) summaries that have deviated from the reference summary. To address this problem, we propose a novel training paradigm which assumes a non-deterministic distribution so that different candidate summaries are assigned probability mass according to their quality. Our method achieves a new state-of-the-art result on the CNN/DailyMail (47.78 ROUGE-1) and XSum (49.07 ROUGE-1) datasets. Further analysis also shows that our model can estimate probabilities of candidate summaries that are more correlated with their level of quality.
UnifiedSKG: Unifying and Multi-Tasking Structured Knowledge Grounding with Text-to-Text Language Models
Jan 20, 2022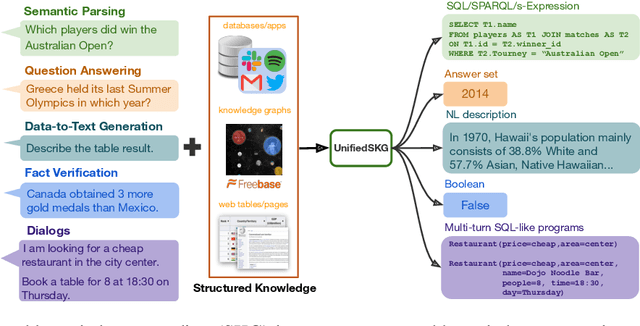
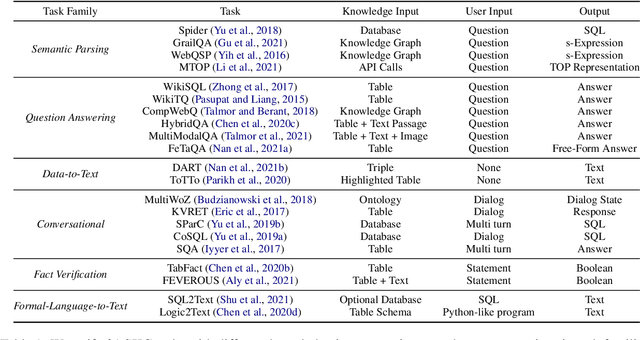
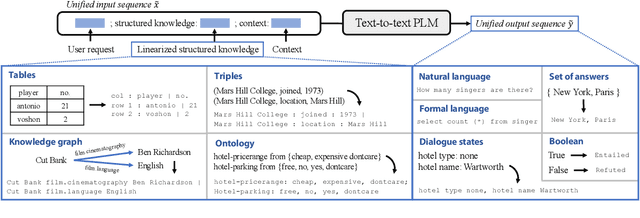
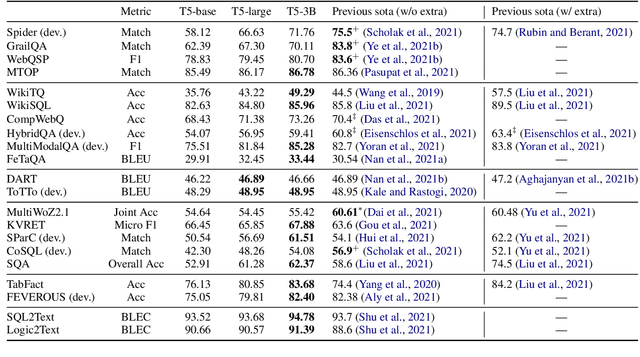
Abstract:Structured knowledge grounding (SKG) leverages structured knowledge to complete user requests, such as semantic parsing over databases and question answering over knowledge bases. Since the inputs and outputs of SKG tasks are heterogeneous, they have been studied separately by different communities, which limits systematic and compatible research on SKG. In this paper, we overcome this limitation by proposing the SKG framework, which unifies 21 SKG tasks into a text-to-text format, aiming to promote systematic SKG research, instead of being exclusive to a single task, domain, or dataset. We use UnifiedSKG to benchmark T5 with different sizes and show that T5, with simple modifications when necessary, achieves state-of-the-art performance on almost all of the 21 tasks. We further demonstrate that multi-task prefix-tuning improves the performance on most tasks, largely improving the overall performance. UnifiedSKG also facilitates the investigation of zero-shot and few-shot learning, and we show that T0, GPT-3, and Codex struggle in zero-shot and few-shot learning for SKG. We also use UnifiedSKG to conduct a series of controlled experiments on structured knowledge encoding variants across SKG tasks. UnifiedSKG is easily extensible to more tasks, and it is open-sourced at https://github.com/hkunlp/unifiedskg Latest collections at https://unifiedskg.com.
A Transfer Learning Pipeline for Educational Resource Discovery with Application in Leading Paragraph Generation
Jan 07, 2022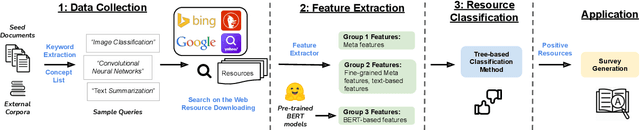

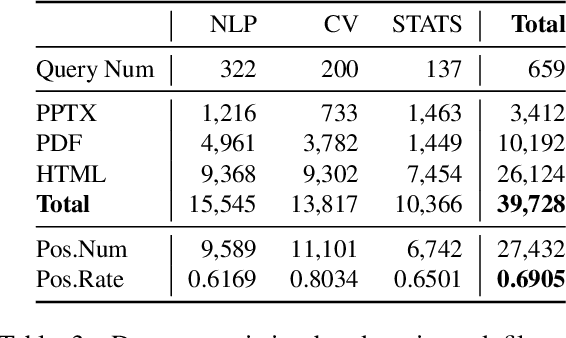
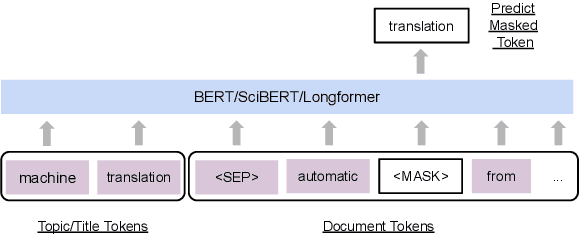
Abstract:Effective human learning depends on a wide selection of educational materials that align with the learner's current understanding of the topic. While the Internet has revolutionized human learning or education, a substantial resource accessibility barrier still exists. Namely, the excess of online information can make it challenging to navigate and discover high-quality learning materials. In this paper, we propose the educational resource discovery (ERD) pipeline that automates web resource discovery for novel domains. The pipeline consists of three main steps: data collection, feature extraction, and resource classification. We start with a known source domain and conduct resource discovery on two unseen target domains via transfer learning. We first collect frequent queries from a set of seed documents and search on the web to obtain candidate resources, such as lecture slides and introductory blog posts. Then we introduce a novel pretrained information retrieval deep neural network model, query-document masked language modeling (QD-MLM), to extract deep features of these candidate resources. We apply a tree-based classifier to decide whether the candidate is a positive learning resource. The pipeline achieves F1 scores of 0.94 and 0.82 when evaluated on two similar but novel target domains. Finally, we demonstrate how this pipeline can benefit an application: leading paragraph generation for surveys. This is the first study that considers various web resources for survey generation, to the best of our knowledge. We also release a corpus of 39,728 manually labeled web resources and 659 queries from NLP, Computer Vision (CV), and Statistics (STATS).
CONFIT: Toward Faithful Dialogue Summarization with Linguistically-Informed Contrastive Fine-tuning
Dec 16, 2021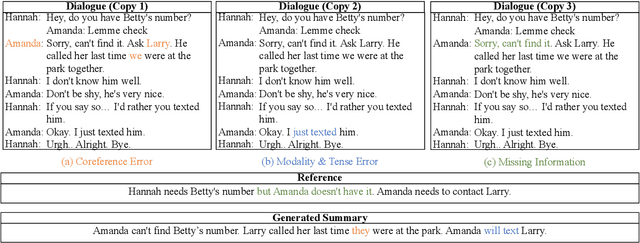
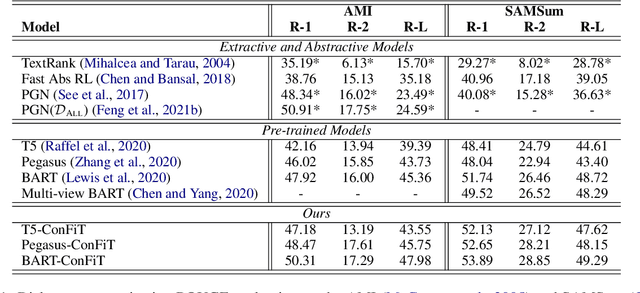
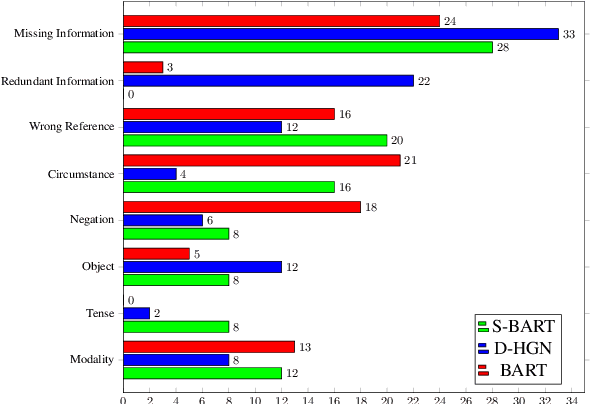
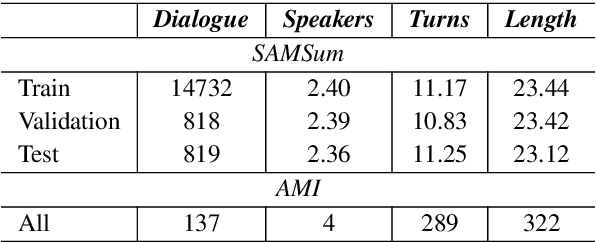
Abstract:Factual inconsistencies in generated summaries severely limit the practical applications of abstractive dialogue summarization. Although significant progress has been achieved by using pre-trained models, substantial amounts of hallucinated content are found during the human evaluation. Pre-trained models are most commonly fine-tuned with cross-entropy loss for text summarization, which may not be an optimal strategy. In this work, we provide a typology of factual errors with annotation data to highlight the types of errors and move away from a binary understanding of factuality. We further propose a training strategy that improves the factual consistency and overall quality of summaries via a novel contrastive fine-tuning, called ConFiT. Based on our linguistically-informed typology of errors, we design different modular objectives that each target a specific type. Specifically, we utilize hard negative samples with errors to reduce the generation of factual inconsistency. In order to capture the key information between speakers, we also design a dialogue-specific loss. Using human evaluation and automatic faithfulness metrics, we show that our model significantly reduces all kinds of factual errors on the dialogue summarization, SAMSum corpus. Moreover, our model could be generalized to the meeting summarization, AMI corpus, and it produces significantly higher scores than most of the baselines on both datasets regarding word-overlap metrics.
CLICKER: A Computational LInguistics Classification Scheme for Educational Resources
Dec 16, 2021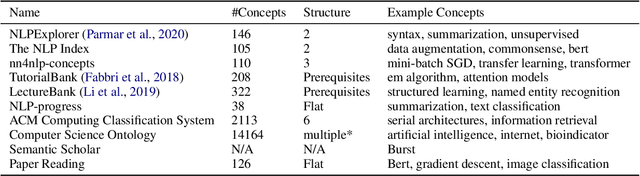
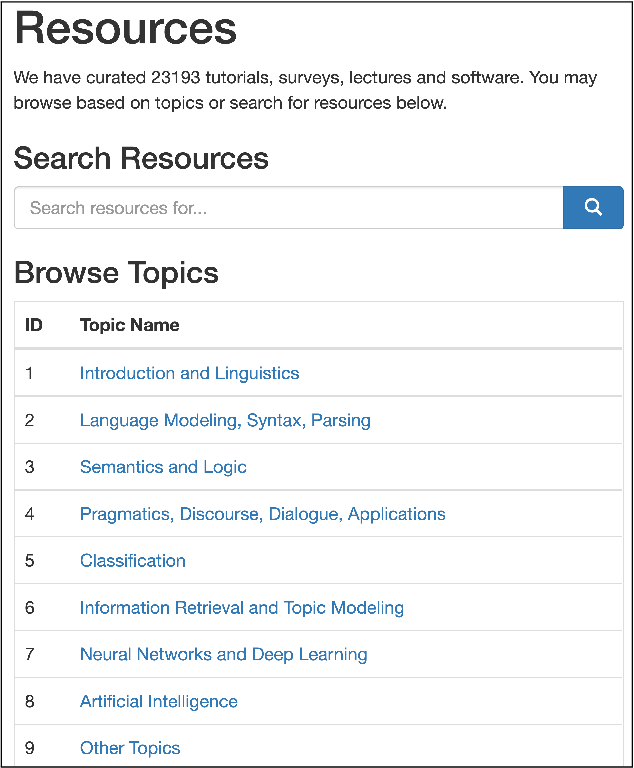
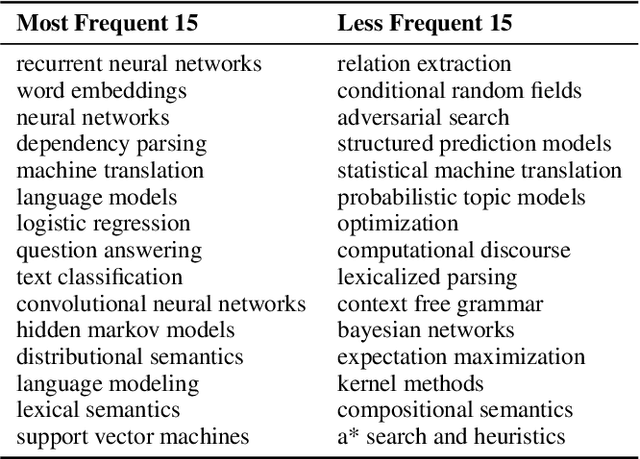
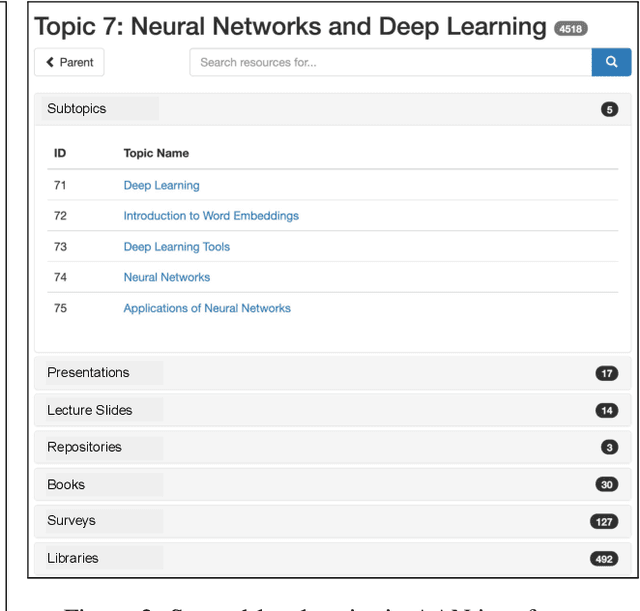
Abstract:A classification scheme of a scientific subject gives an overview of its body of knowledge. It can also be used to facilitate access to research articles and other materials related to the subject. For example, the ACM Computing Classification System (CCS) is used in the ACM Digital Library search interface and also for indexing computer science papers. We observed that a comprehensive classification system like CCS or Mathematics Subject Classification (MSC) does not exist for Computational Linguistics (CL) and Natural Language Processing (NLP). We propose a classification scheme -- CLICKER for CL/NLP based on the analysis of online lectures from 77 university courses on this subject. The currently proposed taxonomy includes 334 topics and focuses on educational aspects of CL/NLP; it is based primarily, but not exclusively, on lecture notes from NLP courses. We discuss how such a taxonomy can help in various real-world applications, including tutoring platforms, resource retrieval, resource recommendation, prerequisite chain learning, and survey generation.
Surfer100: Generating Surveys From Web Resources on Wikipedia-style
Dec 13, 2021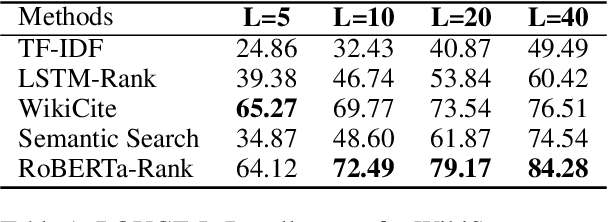
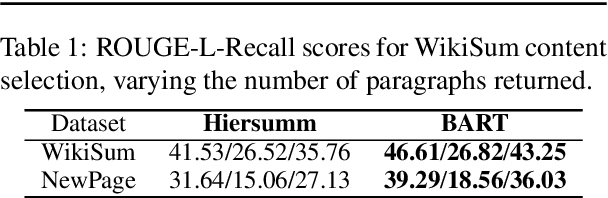

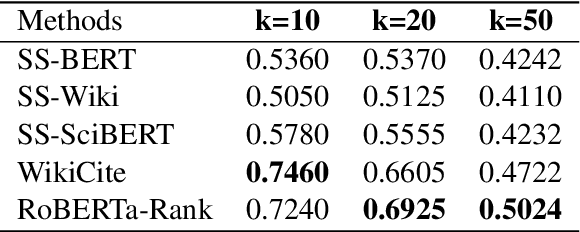
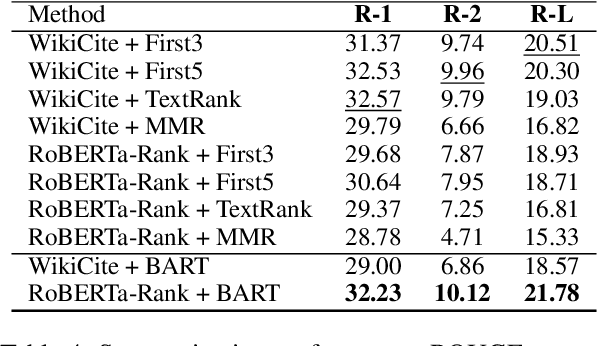
Abstract:Fast-developing fields such as Artificial Intelligence (AI) often outpace the efforts of encyclopedic sources such as Wikipedia, which either do not completely cover recently-introduced topics or lack such content entirely. As a result, methods for automatically producing content are valuable tools to address this information overload. We show that recent advances in pretrained language modeling can be combined for a two-stage extractive and abstractive approach for Wikipedia lead paragraph generation. We extend this approach to generate longer Wikipedia-style summaries with sections and examine how such methods struggle in this application through detailed studies with 100 reference human-collected surveys. This is the first study on utilizing web resources for long Wikipedia-style summaries to the best of our knowledge.
Summ^N: A Multi-Stage Summarization Framework for Long Input Dialogues and Documents
Oct 16, 2021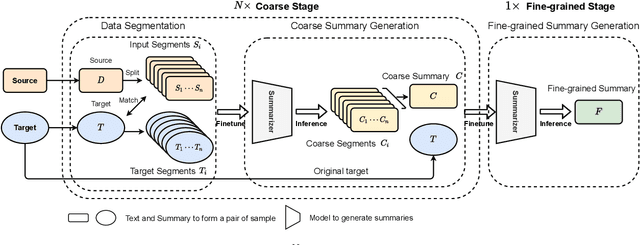
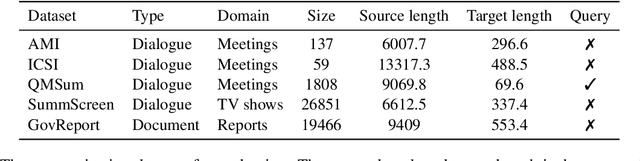
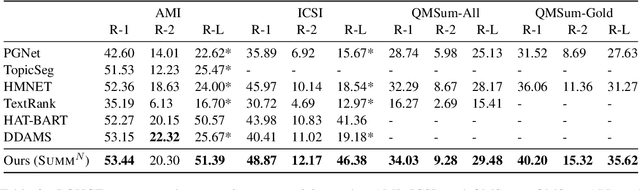
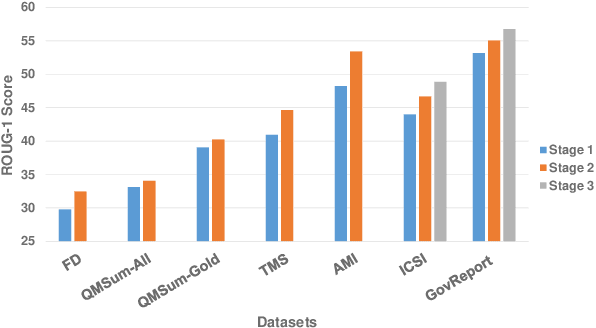
Abstract:Text summarization is an essential task to help readers capture salient information from documents, news, interviews, and meetings. However, most state-of-the-art pretrained language models are unable to efficiently process long text commonly seen in the summarization problem domain. In this paper, we propose Summ^N, a simple, flexible, and effective multi-stage framework for input texts that are longer than the maximum context lengths of typical pretrained LMs. Summ^N first generates the coarse summary in multiple stages and then produces the final fine-grained summary based on them. The framework can process input text of arbitrary length by adjusting the number of stages while keeping the LM context size fixed. Moreover, it can deal with both documents and dialogues and can be used on top of any underlying backbone abstractive summarization model. Our experiments demonstrate that Summ^N significantly outperforms previous state-of-the-art methods by improving ROUGE scores on three long meeting summarization datasets AMI, ICSI, and QMSum, two long TV series datasets from SummScreen, and a newly proposed long document summarization dataset GovReport. Our data and code are available at https://github.com/chatc/Summ-N.
DYLE: Dynamic Latent Extraction for Abstractive Long-Input Summarization
Oct 15, 2021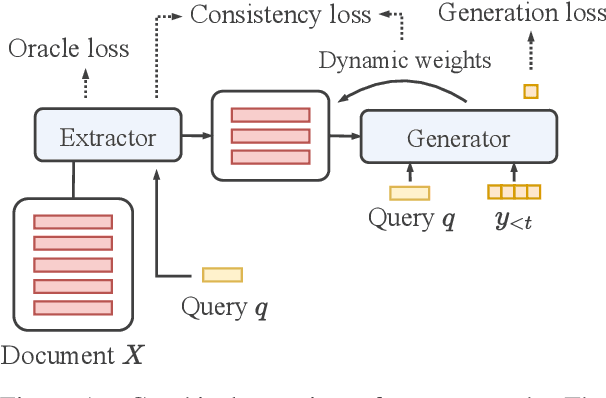
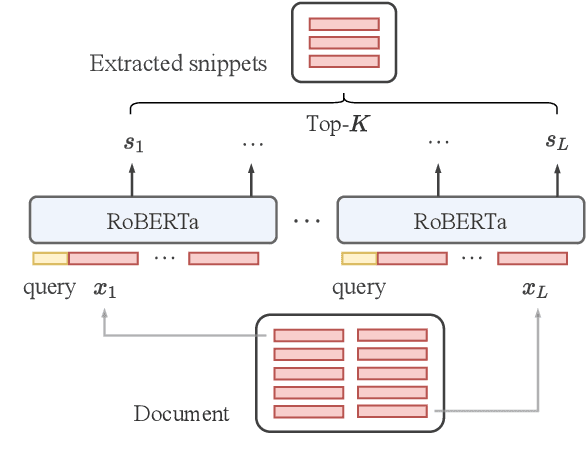
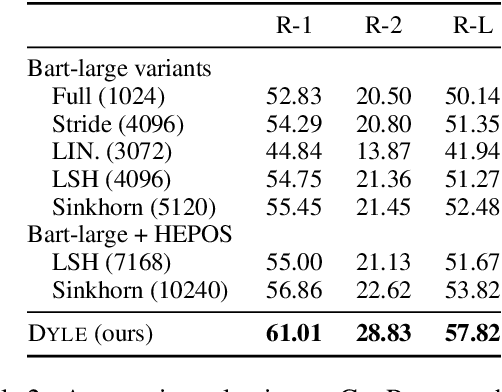
Abstract:Transformer-based models have achieved state-of-the-art performance on short text summarization. However, they still struggle with long-input summarization. In this paper, we present a new approach for long-input summarization: Dynamic Latent Extraction for Abstractive Summarization. We jointly train an extractor with an abstractor and treat the extracted text snippets as the latent variable. We propose extractive oracles to provide the extractor with a strong learning signal. We introduce consistency loss, which encourages the extractor to approximate the averaged dynamic weights predicted by the generator. We conduct extensive tests on two long-input summarization datasets, GovReport (document) and QMSum (dialogue). Our model significantly outperforms the current state-of-the-art, including a 6.21 ROUGE-2 improvement on GovReport and a 2.13 ROUGE-1 improvement on QMSum. Further analysis shows that the dynamic weights make our generation process highly interpretable. Our code will be publicly available upon publication.
3D-Transformer: Molecular Representation with Transformer in 3D Space
Oct 08, 2021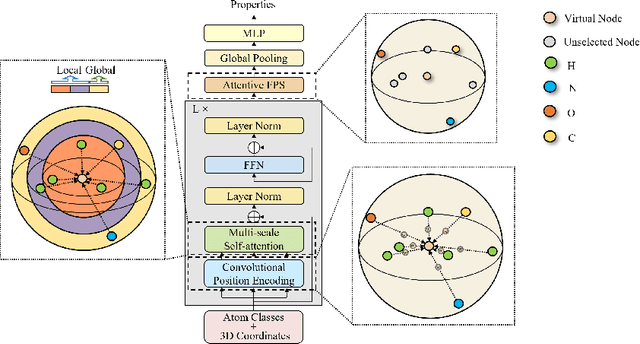
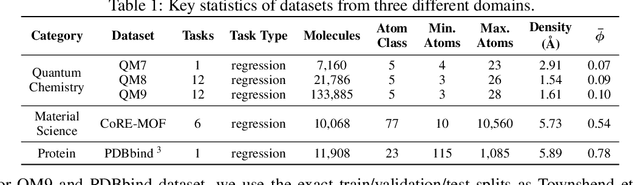
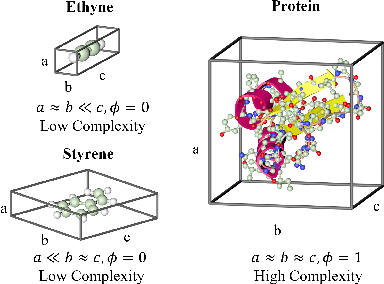
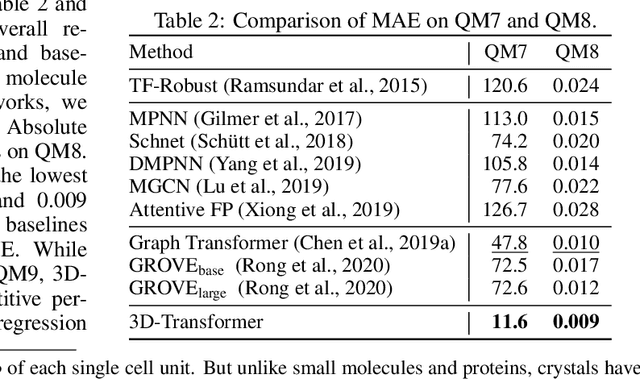
Abstract:Spatial structures in the 3D space are important to determine molecular properties. Recent papers use geometric deep learning to represent molecules and predict properties. These papers, however, are computationally expensive in capturing long-range dependencies of input atoms; and have not considered the non-uniformity of interatomic distances, thus failing to learn context-dependent representations at different scales. To deal with such issues, we introduce 3D-Transformer, a variant of the Transformer for molecular representations that incorporates 3D spatial information. 3D-Transformer operates on a fully-connected graph with direct connections between atoms. To cope with the non-uniformity of interatomic distances, we develop a multi-scale self-attention module that exploits local fine-grained patterns with increasing contextual scales. As molecules of different sizes rely on different kinds of spatial features, we design an adaptive position encoding module that adopts different position encoding methods for small and large molecules. Finally, to attain the molecular representation from atom embeddings, we propose an attentive farthest point sampling algorithm that selects a portion of atoms with the assistance of attention scores, overcoming handicaps of the virtual node and previous distance-dominant downsampling methods. We validate 3D-Transformer across three important scientific domains: quantum chemistry, material science, and proteomics. Our experiments show significant improvements over state-of-the-art models on the crystal property prediction task and the protein-ligand binding affinity prediction task, and show better or competitive performance in quantum chemistry molecular datasets. This work provides clear evidence that biochemical tasks can gain consistent benefits from 3D molecular representations and different tasks require different position encoding methods.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge