Alan L. Yuille
Data, Assemble: Leveraging Multiple Datasets with Heterogeneous and Partial Labels
Sep 25, 2021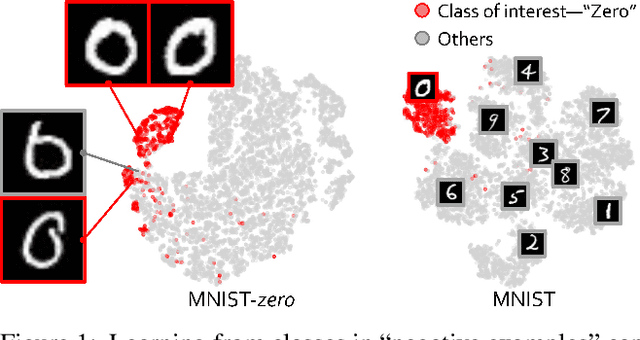
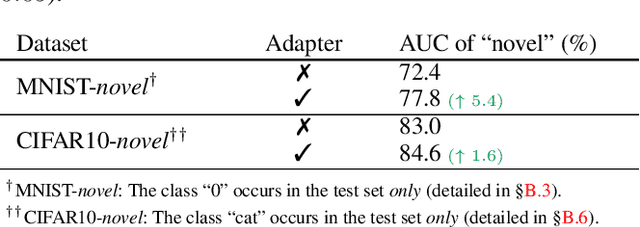
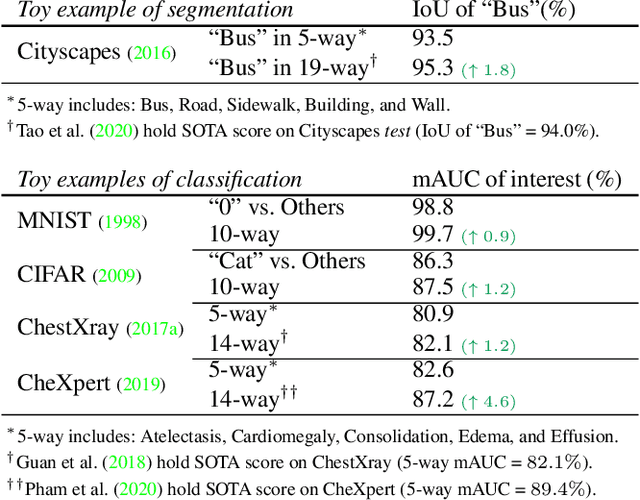
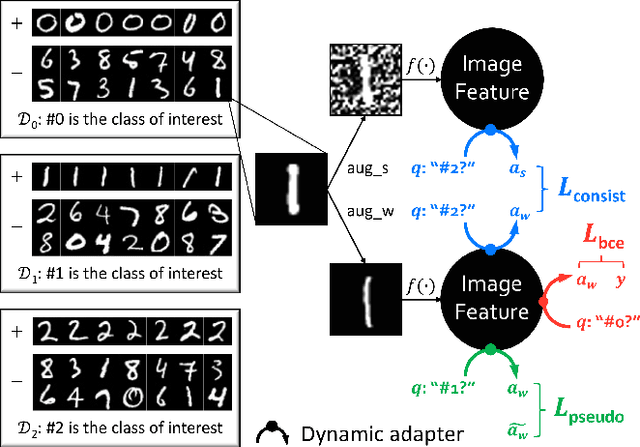

Abstract:The success of deep learning relies heavily on large datasets with extensive labels, but we often only have access to several small, heterogeneous datasets associated with partial labels, particularly in the field of medical imaging. When learning from multiple datasets, existing challenges include incomparable, heterogeneous, or even conflicting labeling protocols across datasets. In this paper, we propose a new initiative--"data, assemble"--which aims to unleash the full potential of partially labeled data and enormous unlabeled data from an assembly of datasets. To accommodate the supervised learning paradigm to partial labels, we introduce a dynamic adapter that encodes multiple visual tasks and aggregates image features in a question-and-answer manner. Furthermore, we employ pseudo-labeling and consistency constraints to harness images with missing labels and to mitigate the domain gap across datasets. From proof-of-concept studies on three natural imaging datasets and rigorous evaluations on two large-scale thorax X-ray benchmarks, we discover that learning from "negative examples" facilitates both classification and segmentation of classes of interest. This sheds new light on the computer-aided diagnosis of rare diseases and emerging pandemics, wherein "positive examples" are hard to collect, yet "negative examples" are relatively easier to assemble. As a result, besides exceeding the prior art in the NIH ChestXray benchmark, our model is particularly strong in identifying diseases of minority classes, yielding over 3-point improvement on average. Remarkably, when using existing partial labels, our model performance is on-par (p>0.05) with that using a fully curated dataset with exhaustive labels, eliminating the need for additional 40% annotation costs.
DeepLab2: A TensorFlow Library for Deep Labeling
Jun 17, 2021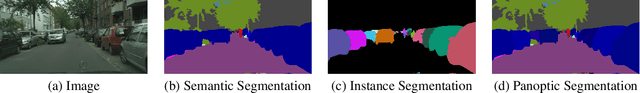
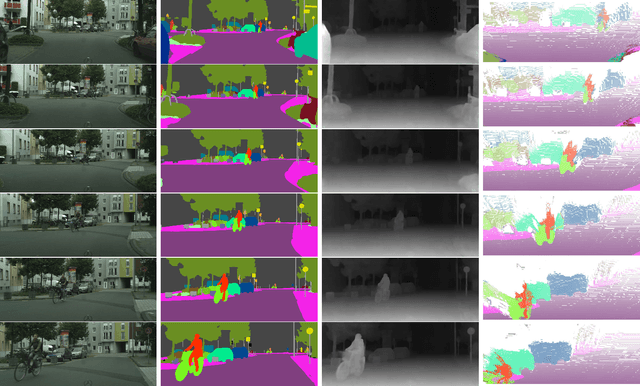


Abstract:DeepLab2 is a TensorFlow library for deep labeling, aiming to provide a state-of-the-art and easy-to-use TensorFlow codebase for general dense pixel prediction problems in computer vision. DeepLab2 includes all our recently developed DeepLab model variants with pretrained checkpoints as well as model training and evaluation code, allowing the community to reproduce and further improve upon the state-of-art systems. To showcase the effectiveness of DeepLab2, our Panoptic-DeepLab employing Axial-SWideRNet as network backbone achieves 68.0% PQ or 83.5% mIoU on Cityscaspes validation set, with only single-scale inference and ImageNet-1K pretrained checkpoints. We hope that publicly sharing our library could facilitate future research on dense pixel labeling tasks and envision new applications of this technology. Code is made publicly available at \url{https://github.com/google-research/deeplab2}.
Learning Inductive Attention Guidance for Partially Supervised Pancreatic Ductal Adenocarcinoma Prediction
May 31, 2021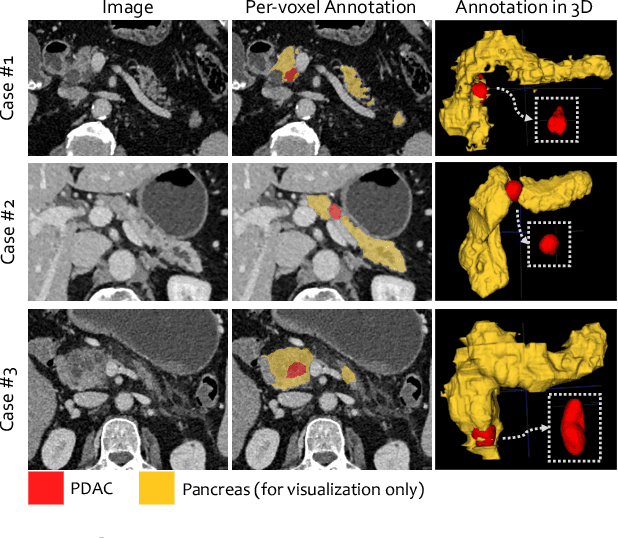
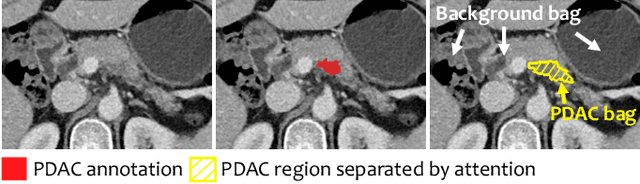
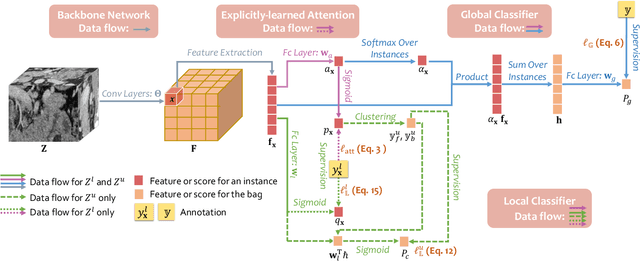
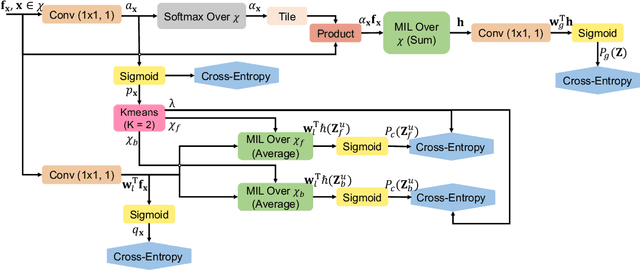
Abstract:Pancreatic ductal adenocarcinoma (PDAC) is the third most common cause of cancer death in the United States. Predicting tumors like PDACs (including both classification and segmentation) from medical images by deep learning is becoming a growing trend, but usually a large number of annotated data are required for training, which is very labor-intensive and time-consuming. In this paper, we consider a partially supervised setting, where cheap image-level annotations are provided for all the training data, and the costly per-voxel annotations are only available for a subset of them. We propose an Inductive Attention Guidance Network (IAG-Net) to jointly learn a global image-level classifier for normal/PDAC classification and a local voxel-level classifier for semi-supervised PDAC segmentation. We instantiate both the global and the local classifiers by multiple instance learning (MIL), where the attention guidance, indicating roughly where the PDAC regions are, is the key to bridging them: For global MIL based normal/PDAC classification, attention serves as a weight for each instance (voxel) during MIL pooling, which eliminates the distraction from the background; For local MIL based semi-supervised PDAC segmentation, the attention guidance is inductive, which not only provides bag-level pseudo-labels to training data without per-voxel annotations for MIL training, but also acts as a proxy of an instance-level classifier. Experimental results show that our IAG-Net boosts PDAC segmentation accuracy by more than 5% compared with the state-of-the-arts.
Sequential Learning on Liver Tumor Boundary Semantics and Prognostic Biomarker Mining
Mar 09, 2021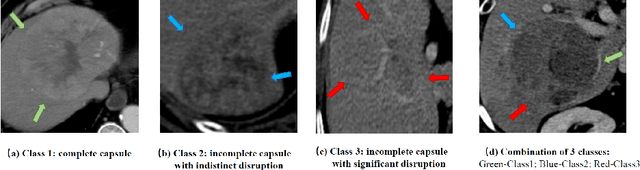
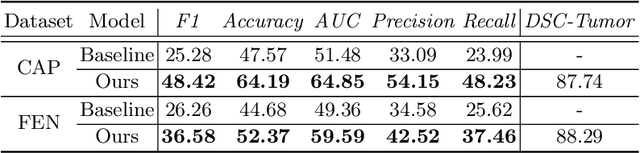
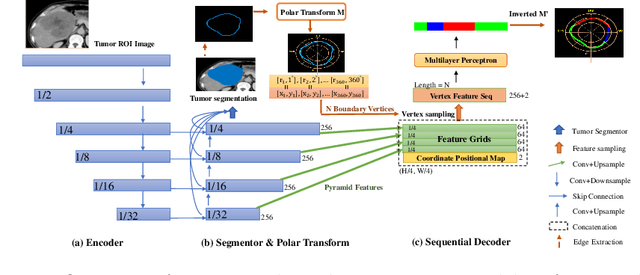
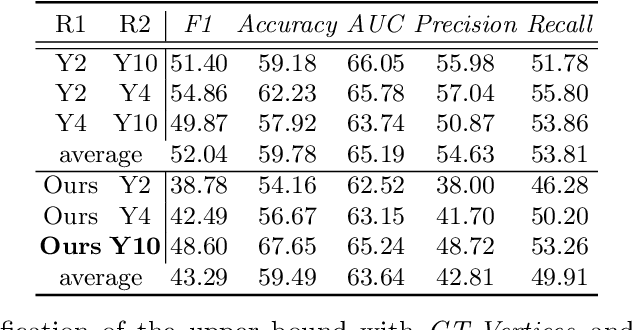
Abstract:The boundary of tumors (hepatocellular carcinoma, or HCC) contains rich semantics: capsular invasion, visibility, smoothness, folding and protuberance, etc. Capsular invasion on tumor boundary has proven to be clinically correlated with the prognostic indicator, microvascular invasion (MVI). Investigating tumor boundary semantics has tremendous clinical values. In this paper, we propose the first and novel computational framework that disentangles the task into two components: spatial vertex localization and sequential semantic classification. (1) A HCC tumor segmentor is built for tumor mask boundary extraction, followed by polar transform representing the boundary with radius and angle. Vertex generator is used to produce fixed-length boundary vertices where vertex features are sampled on the corresponding spatial locations. (2) The sampled deep vertex features with positional embedding are mapped into a sequential space and decoded by a multilayer perceptron (MLP) for semantic classification. Extensive experiments on tumor capsule semantics demonstrate the effectiveness of our framework. Mining the correlation between the boundary semantics and MVI status proves the feasibility to integrate this boundary semantics as a valid HCC prognostic biomarker.
Multi-phase Deformable Registration for Time-dependent Abdominal Organ Variations
Mar 08, 2021
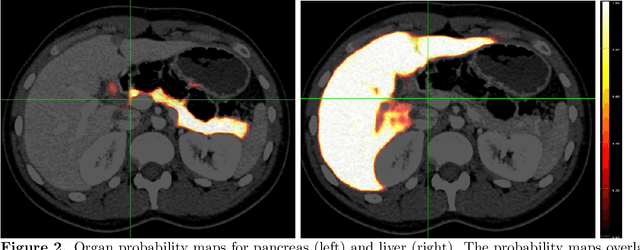
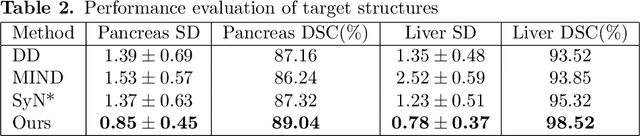
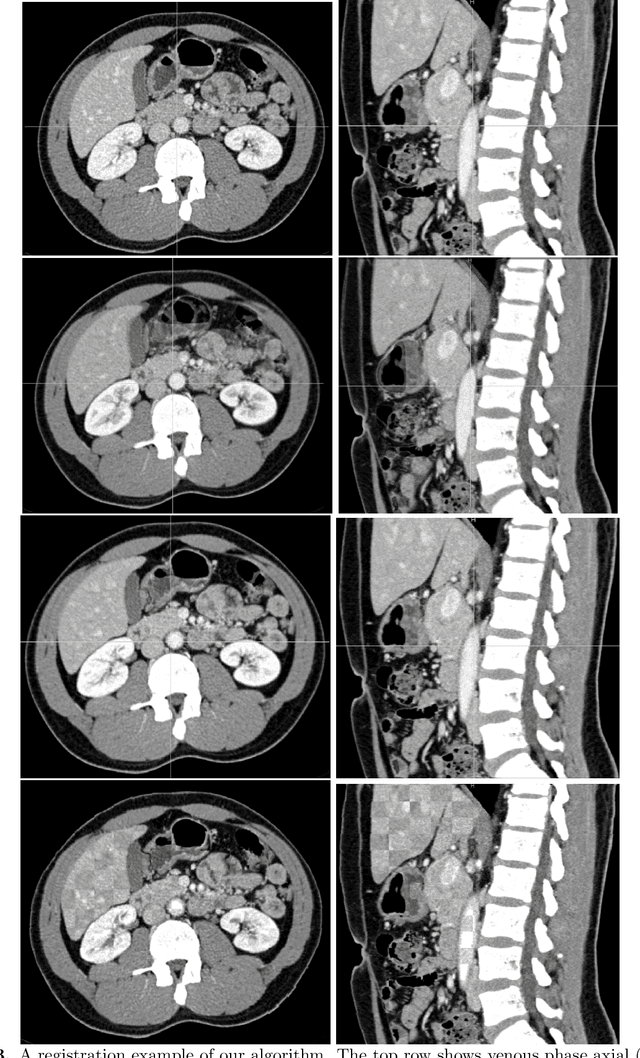
Abstract:Human body is a complex dynamic system composed of various sub-dynamic parts. Especially, thoracic and abdominal organs have complex internal shape variations with different frequencies by various reasons such as respiration with fast motion and peristalsis with slower motion. CT protocols for abdominal lesions are multi-phase scans for various tumor detection to use different vascular contrast, however, they are not aligned well enough to visually check the same area. In this paper, we propose a time-efficient and accurate deformable registration algorithm for multi-phase CT scans considering abdominal organ motions, which can be applied for differentiable or non-differentiable motions of abdominal organs. Experimental results shows the registration accuracy as 0.85 +/- 0.45mm (mean +/- STD) for pancreas within 1 minute for the whole abdominal region.
TransUNet: Transformers Make Strong Encoders for Medical Image Segmentation
Feb 08, 2021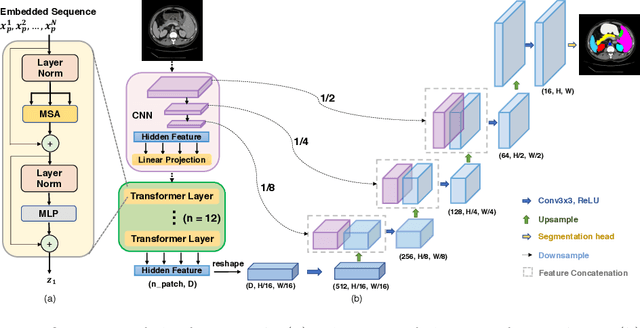
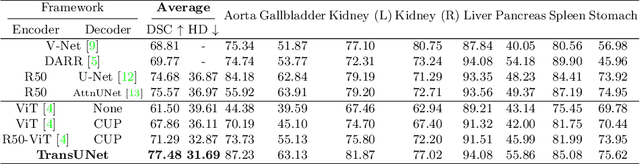
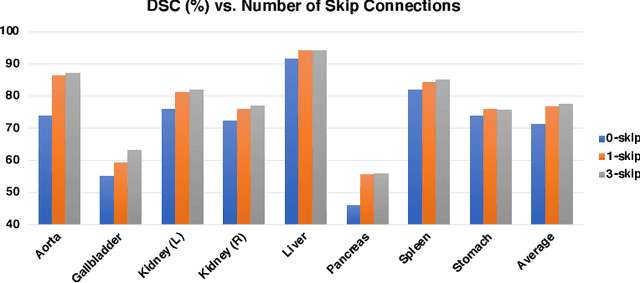
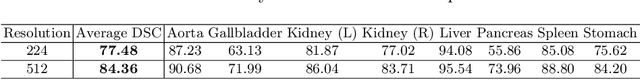
Abstract:Medical image segmentation is an essential prerequisite for developing healthcare systems, especially for disease diagnosis and treatment planning. On various medical image segmentation tasks, the u-shaped architecture, also known as U-Net, has become the de-facto standard and achieved tremendous success. However, due to the intrinsic locality of convolution operations, U-Net generally demonstrates limitations in explicitly modeling long-range dependency. Transformers, designed for sequence-to-sequence prediction, have emerged as alternative architectures with innate global self-attention mechanisms, but can result in limited localization abilities due to insufficient low-level details. In this paper, we propose TransUNet, which merits both Transformers and U-Net, as a strong alternative for medical image segmentation. On one hand, the Transformer encodes tokenized image patches from a convolution neural network (CNN) feature map as the input sequence for extracting global contexts. On the other hand, the decoder upsamples the encoded features which are then combined with the high-resolution CNN feature maps to enable precise localization. We argue that Transformers can serve as strong encoders for medical image segmentation tasks, with the combination of U-Net to enhance finer details by recovering localized spatial information. TransUNet achieves superior performances to various competing methods on different medical applications including multi-organ segmentation and cardiac segmentation. Code and models are available at https://github.com/Beckschen/TransUNet.
Nothing But Geometric Constraints: A Model-Free Method for Articulated Object Pose Estimation
Nov 30, 2020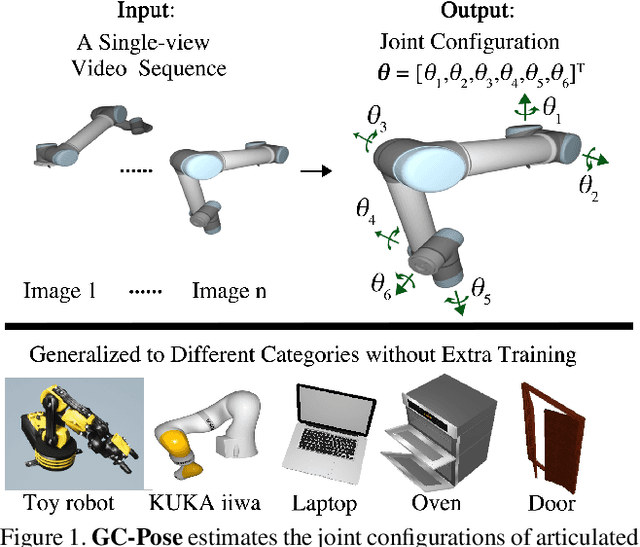

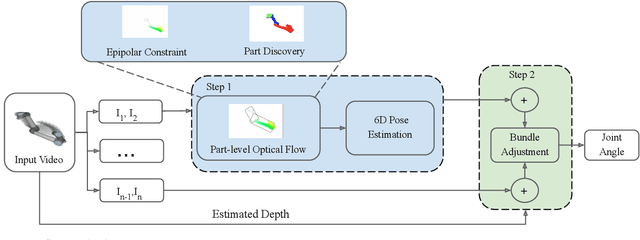
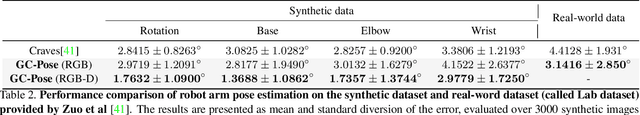
Abstract:We propose an unsupervised vision-based system to estimate the joint configurations of the robot arm from a sequence of RGB or RGB-D images without knowing the model a priori, and then adapt it to the task of category-independent articulated object pose estimation. We combine a classical geometric formulation with deep learning and extend the use of epipolar constraint to multi-rigid-body systems to solve this task. Given a video sequence, the optical flow is estimated to get the pixel-wise dense correspondences. After that, the 6D pose is computed by a modified PnP algorithm. The key idea is to leverage the geometric constraints and the constraint between multiple frames. Furthermore, we build a synthetic dataset with different kinds of robots and multi-joint articulated objects for the research of vision-based robot control and robotic vision. We demonstrate the effectiveness of our method on three benchmark datasets and show that our method achieves higher accuracy than the state-of-the-art supervised methods in estimating joint angles of robot arms and articulated objects.
Volumetric Medical Image Segmentation: A 3D Deep Coarse-to-fine Framework and Its Adversarial Examples
Oct 29, 2020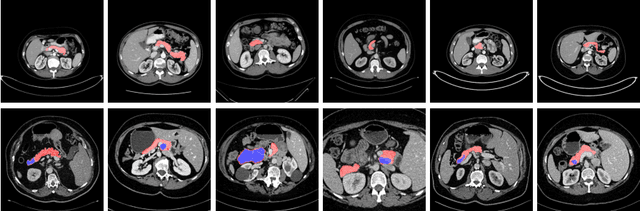
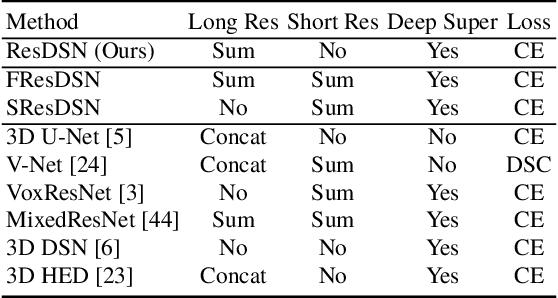
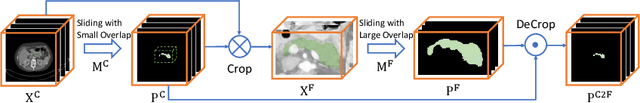
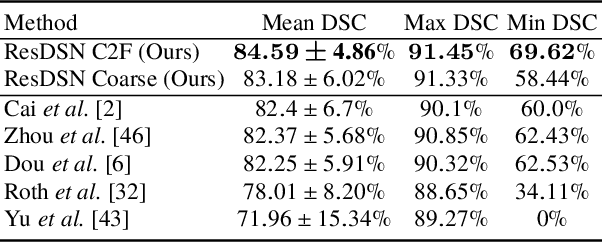
Abstract:Although deep neural networks have been a dominant method for many 2D vision tasks, it is still challenging to apply them to 3D tasks, such as medical image segmentation, due to the limited amount of annotated 3D data and limited computational resources. In this chapter, by rethinking the strategy to apply 3D Convolutional Neural Networks to segment medical images, we propose a novel 3D-based coarse-to-fine framework to efficiently tackle these challenges. The proposed 3D-based framework outperforms their 2D counterparts by a large margin since it can leverage the rich spatial information along all three axes. We further analyze the threat of adversarial attacks on the proposed framework and show how to defense against the attack. We conduct experiments on three datasets, the NIH pancreas dataset, the JHMI pancreas dataset and the JHMI pathological cyst dataset, where the first two and the last one contain healthy and pathological pancreases respectively, and achieve the current state-of-the-art in terms of Dice-Sorensen Coefficient (DSC) on all of them. Especially, on the NIH pancreas segmentation dataset, we outperform the previous best by an average of over $2\%$, and the worst case is improved by $7\%$ to reach almost $70\%$, which indicates the reliability of our framework in clinical applications.
Segmentation for Classification of Screening Pancreatic Neuroendocrine Tumors
Apr 04, 2020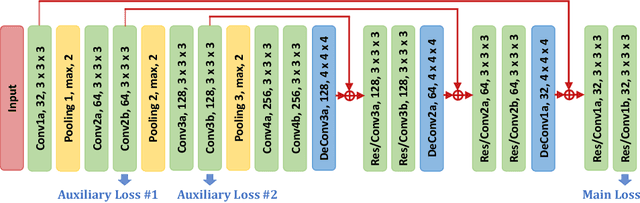
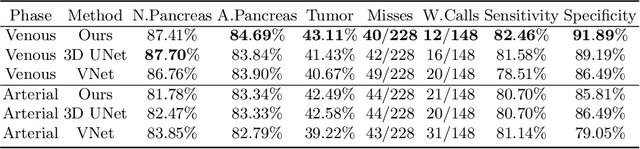
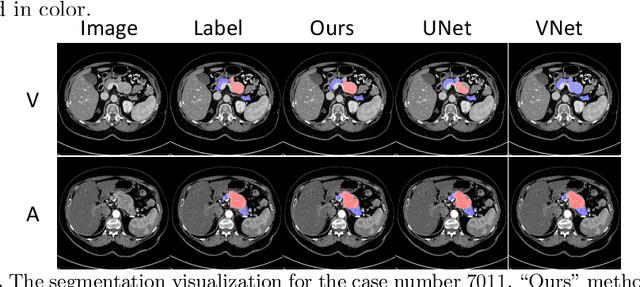
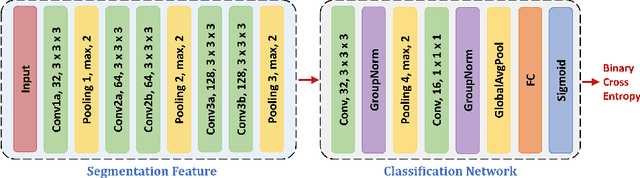
Abstract:This work presents comprehensive results to detect in the early stage the pancreatic neuroendocrine tumors (PNETs), a group of endocrine tumors arising in the pancreas, which are the second common type of pancreatic cancer, by checking the abdominal CT scans. To the best of our knowledge, this task has not been studied before as a computational task. To provide radiologists with tumor locations, we adopt a segmentation framework to classify CT volumes by checking if at least a sufficient number of voxels is segmented as tumors. To quantitatively analyze our method, we collect and voxelwisely label a new abdominal CT dataset containing $376$ cases with both arterial and venous phases available for each case, in which $228$ cases were diagnosed with PNETs while the remaining $148$ cases are normal, which is currently the largest dataset for PNETs to the best of our knowledge. In order to incorporate rich knowledge of radiologists to our framework, we annotate dilated pancreatic duct as well, which is regarded as the sign of high risk for pancreatic cancer. Quantitatively, our approach outperforms state-of-the-art segmentation networks and achieves a sensitivity of $89.47\%$ at a specificity of $81.08\%$, which indicates a potential direction to achieve a clinical impact related to cancer diagnosis by earlier tumor detection.
Detecting Pancreatic Adenocarcinoma in Multi-phase CT Scans via Alignment Ensemble
Apr 03, 2020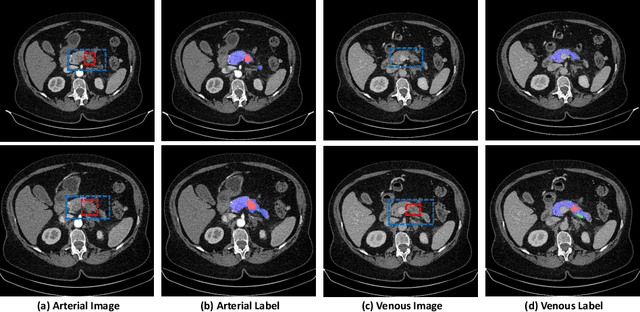
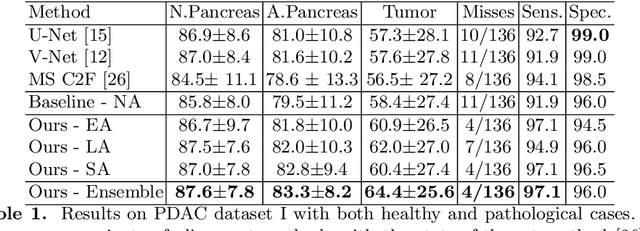
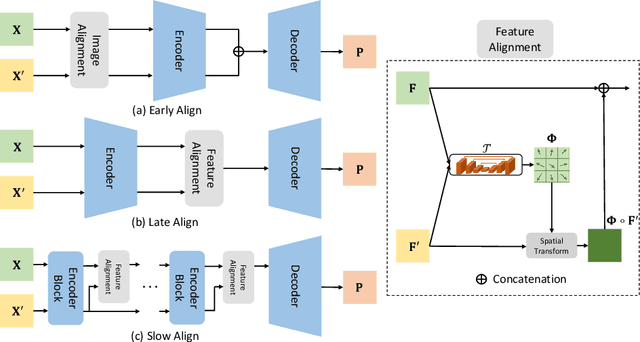
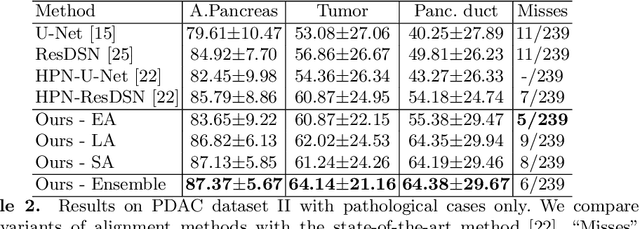
Abstract:Pancreatic ductal adenocarcinoma (PDAC) is one of the most lethal cancers among population. Screening for PDACs in dynamic contrast-enhanced CT is beneficial for early diagnose. In this paper, we investigate the problem of automated detecting PDACs in multi-phase (arterial and venous) CT scans. Multiple phases provide more information than single phase, but they are unaligned and inhomogeneous in texture, making it difficult to combine cross-phase information seamlessly. We study multiple phase alignment strategies, i.e., early alignment (image registration), late alignment (high-level feature registration) and slow alignment (multi-level feature registration), and suggest an ensemble of all these alignments as a promising way to boost the performance of PDAC detection. We provide an extensive empirical evaluation on two PDAC datasets and show that the proposed alignment ensemble significantly outperforms previous state-of-the-art approaches, illustrating strong potential for clinical use.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge