"Image": models, code, and papers
Transformation-Grounded Image Generation Network for Novel 3D View Synthesis
Mar 08, 2017
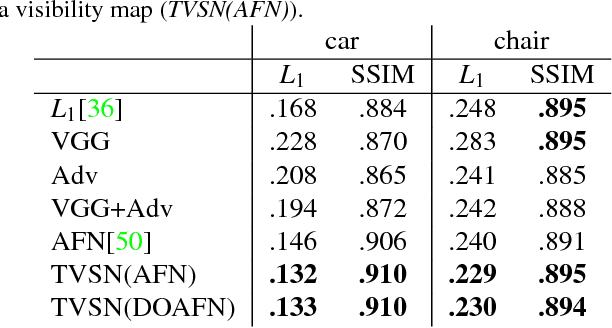
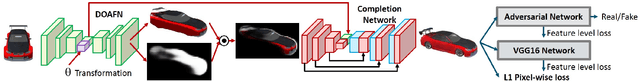

We present a transformation-grounded image generation network for novel 3D view synthesis from a single image. Instead of taking a 'blank slate' approach, we first explicitly infer the parts of the geometry visible both in the input and novel views and then re-cast the remaining synthesis problem as image completion. Specifically, we both predict a flow to move the pixels from the input to the novel view along with a novel visibility map that helps deal with occulsion/disocculsion. Next, conditioned on those intermediate results, we hallucinate (infer) parts of the object invisible in the input image. In addition to the new network structure, training with a combination of adversarial and perceptual loss results in a reduction in common artifacts of novel view synthesis such as distortions and holes, while successfully generating high frequency details and preserving visual aspects of the input image. We evaluate our approach on a wide range of synthetic and real examples. Both qualitative and quantitative results show our method achieves significantly better results compared to existing methods.
PointGuard: Provably Robust 3D Point Cloud Classification
Mar 04, 2021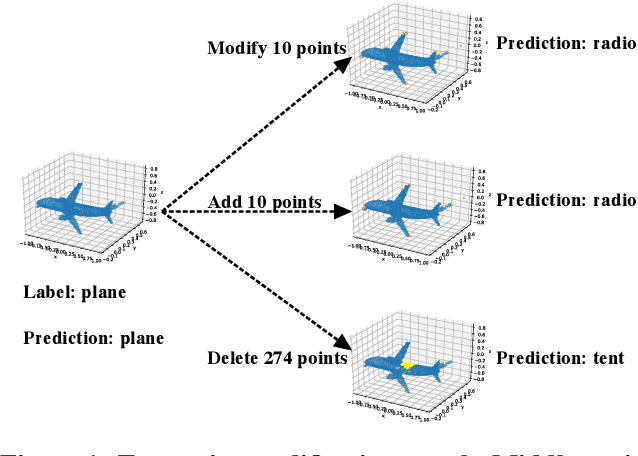
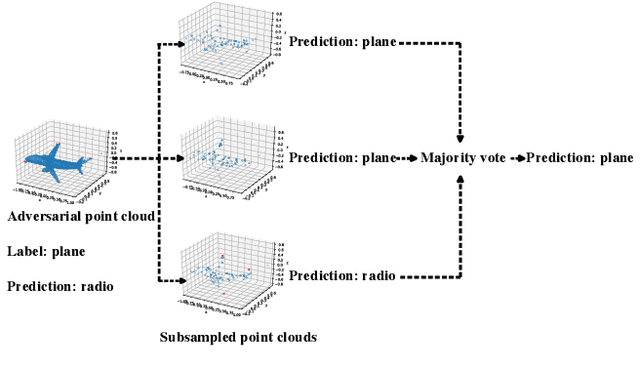
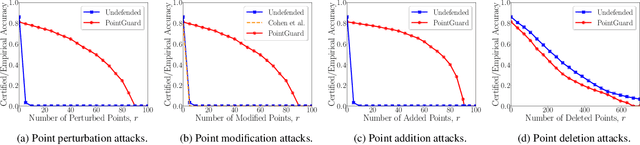
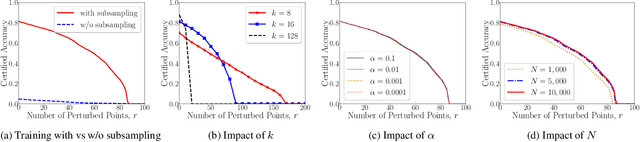
3D point cloud classification has many safety-critical applications such as autonomous driving and robotic grasping. However, several studies showed that it is vulnerable to adversarial attacks. In particular, an attacker can make a classifier predict an incorrect label for a 3D point cloud via carefully modifying, adding, and/or deleting a small number of its points. Randomized smoothing is state-of-the-art technique to build certifiably robust 2D image classifiers. However, when applied to 3D point cloud classification, randomized smoothing can only certify robustness against adversarially {modified} points. In this work, we propose PointGuard, the first defense that has provable robustness guarantees against adversarially modified, added, and/or deleted points. Specifically, given a 3D point cloud and an arbitrary point cloud classifier, our PointGuard first creates multiple subsampled point clouds, each of which contains a random subset of the points in the original point cloud; then our PointGuard predicts the label of the original point cloud as the majority vote among the labels of the subsampled point clouds predicted by the point cloud classifier. Our first major theoretical contribution is that we show PointGuard provably predicts the same label for a 3D point cloud when the number of adversarially modified, added, and/or deleted points is bounded. Our second major theoretical contribution is that we prove the tightness of our derived bound when no assumptions on the point cloud classifier are made. Moreover, we design an efficient algorithm to compute our certified robustness guarantees. We also empirically evaluate PointGuard on ModelNet40 and ScanNet benchmark datasets.
Cross-domain CNN for Hyperspectral Image Classification
May 02, 2018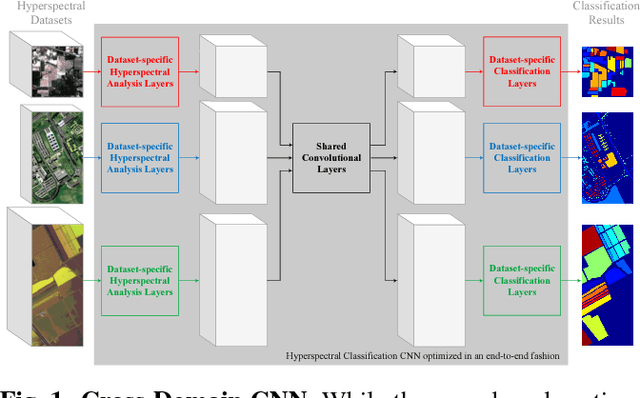
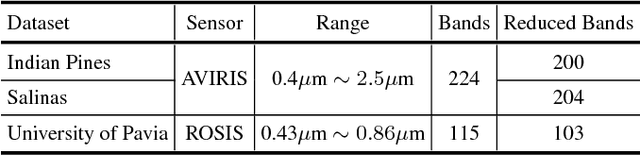
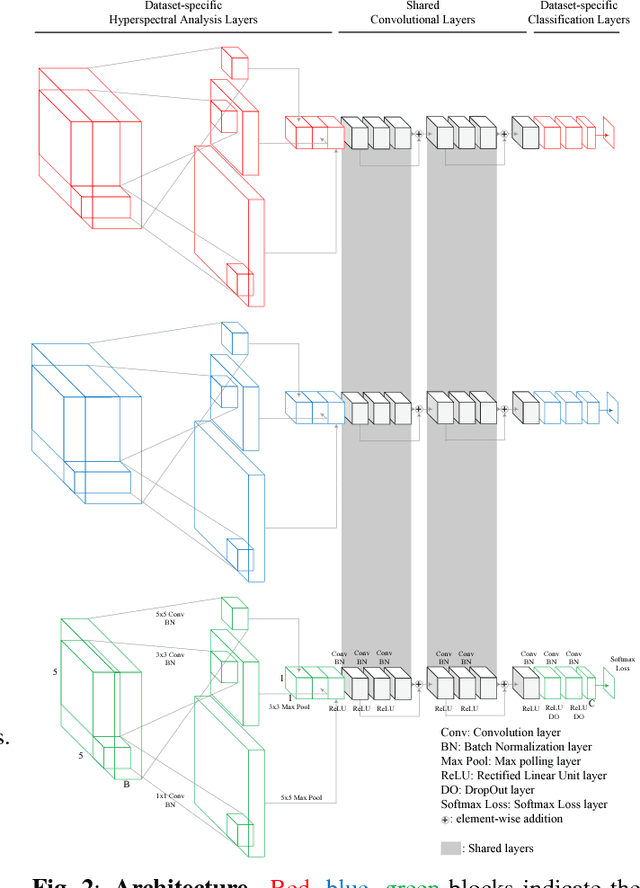
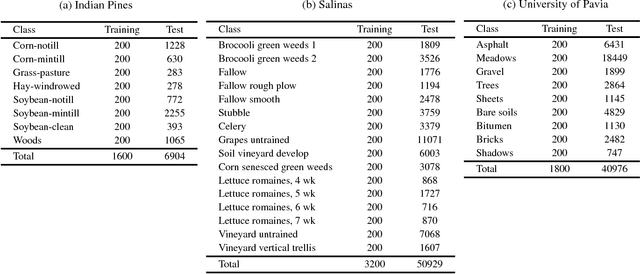
In this paper, we address the dataset scarcity issue with the hyperspectral image classification. As only a few thousands of pixels are available for training, it is difficult to effectively learn high-capacity Convolutional Neural Networks (CNNs). To cope with this problem, we propose a novel cross-domain CNN containing the shared parameters which can co-learn across multiple hyperspectral datasets. The network also contains the non-shared portions designed to handle the dataset specific spectral characteristics and the associated classification tasks. Our approach is the first attempt to learn a CNN for multiple hyperspectral datasets, in an end-to-end fashion. Moreover, we have experimentally shown that the proposed network trained on three of the widely used datasets outperform all the baseline networks which are trained on single dataset.
On the Space-Time Statistics of Motion Pictures
Jan 29, 2021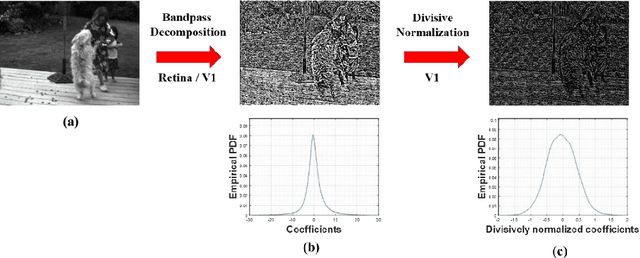
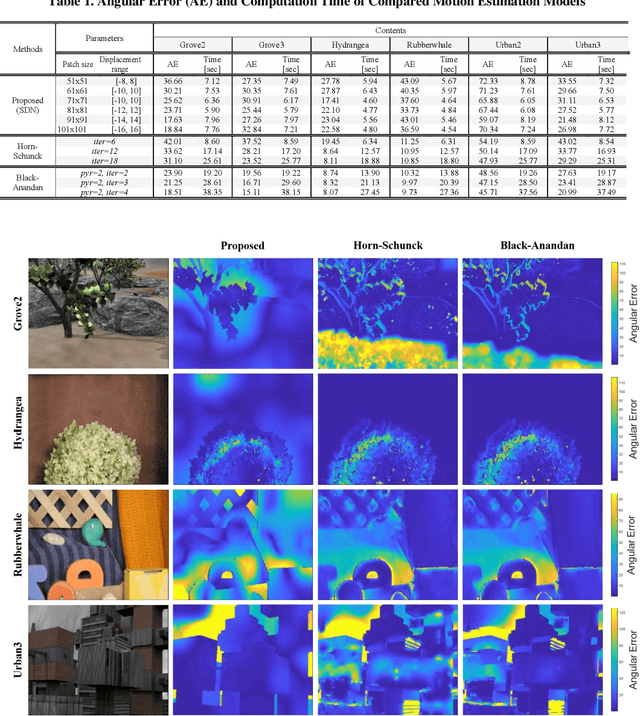
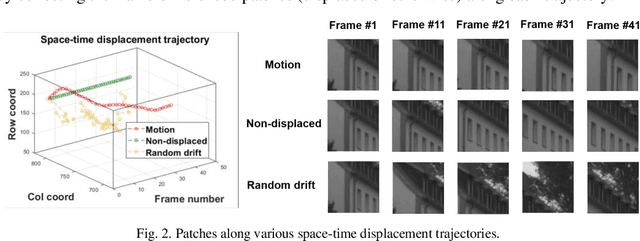
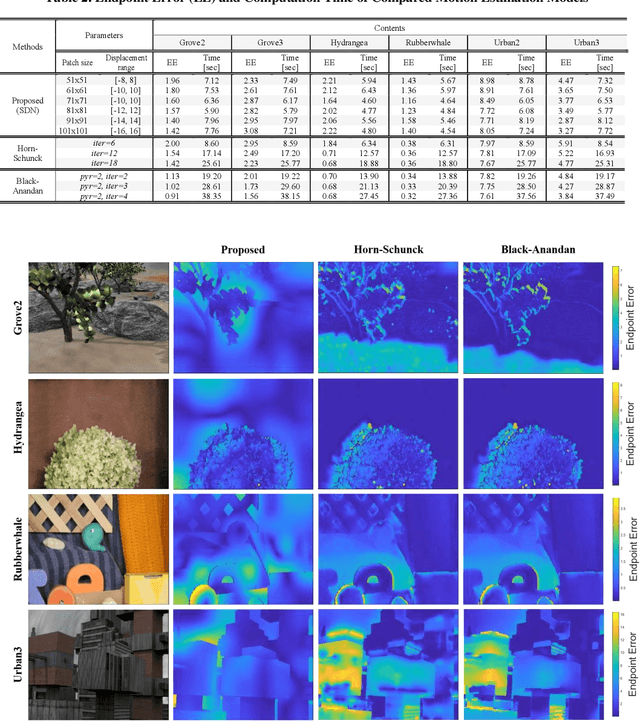
It is well-known that natural images possess statistical regularities that can be captured by bandpass decomposition and divisive normalization processes that approximate early neural processing in the human visual system. We expand on these studies and present new findings on the properties of space-time natural statistics that are inherent in motion pictures. Our model relies on the concept of temporal bandpass (e.g. lag) filtering in LGN and area V1, which is similar to smoothed frame differencing of video frames. Specifically, we model the statistics of the differences between adjacent or neighboring video frames that have been slightly spatially displaced relative to one another. We find that when these space-time differences are further subjected to locally pooled divisive normalization, statistical regularities (or lack thereof) arise that depend on the local motion trajectory. We find that bandpass and divisively normalized frame-differences that are displaced along the motion direction exhibit stronger statistical regularities than for other displacements. Conversely, the direction-dependent regularities of displaced frame differences can be used to estimate the image motion (optical flow) by finding the space-time displacement paths that best preserve statistical regularity.
Automated Deep Learning Analysis of Angiography Video Sequences for Coronary Artery Disease
Jan 29, 2021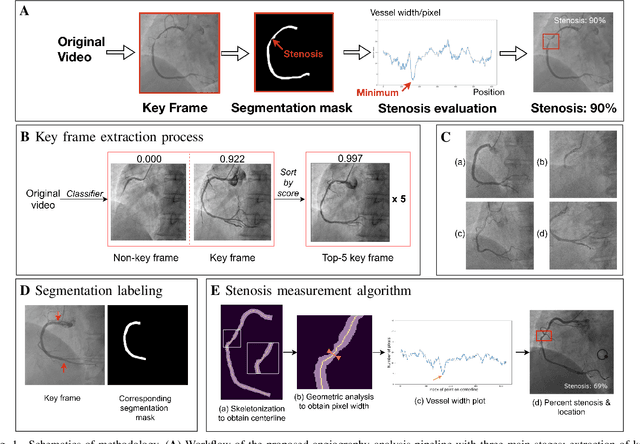
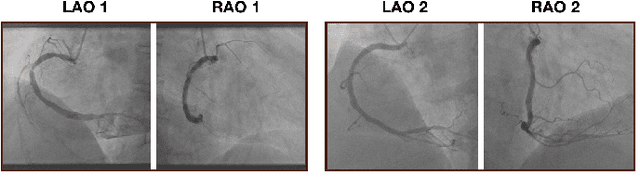
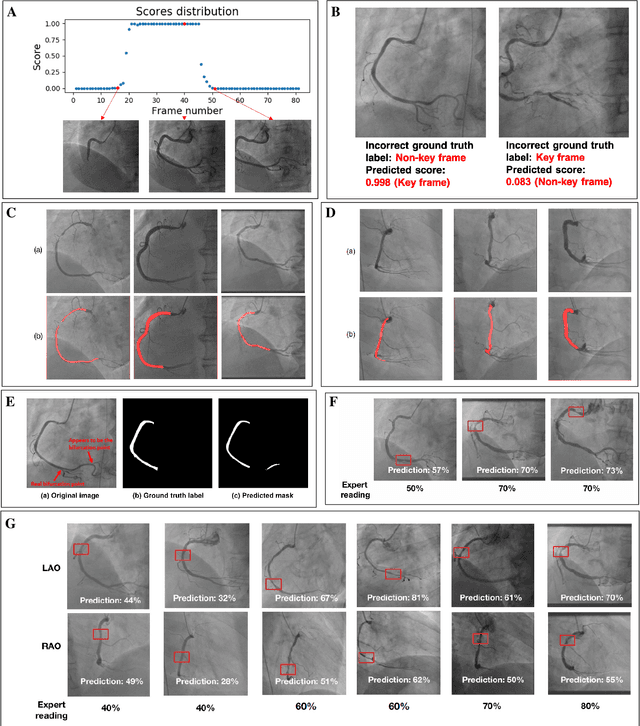
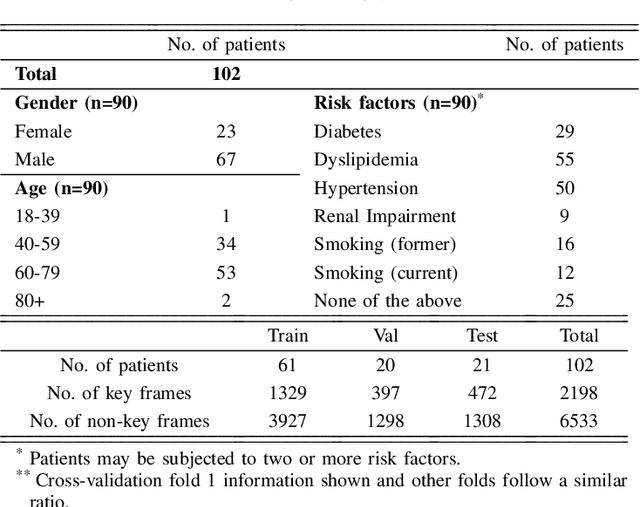
The evaluation of obstructions (stenosis) in coronary arteries is currently done by a physician's visual assessment of coronary angiography video sequences. It is laborious, and can be susceptible to interobserver variation. Prior studies have attempted to automate this process, but few have demonstrated an integrated suite of algorithms for the end-to-end analysis of angiograms. We report an automated analysis pipeline based on deep learning to rapidly and objectively assess coronary angiograms, highlight coronary vessels of interest, and quantify potential stenosis. We propose a 3-stage automated analysis method consisting of key frame extraction, vessel segmentation, and stenosis measurement. We combined powerful deep learning approaches such as ResNet and U-Net with traditional image processing and geometrical analysis. We trained and tested our algorithms on the Left Anterior Oblique (LAO) view of the right coronary artery (RCA) using anonymized angiograms obtained from a tertiary cardiac institution, then tested the generalizability of our technique to the Right Anterior Oblique (RAO) view. We demonstrated an overall improvement on previous work, with key frame extraction top-5 precision of 98.4%, vessel segmentation F1-Score of 0.891 and stenosis measurement 20.7% Type I Error rate.
Learning Neural Light Transport
Jun 05, 2020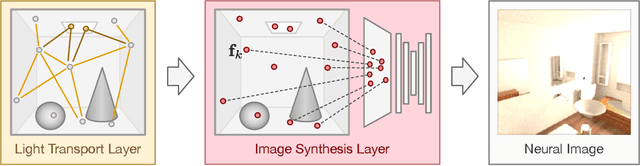
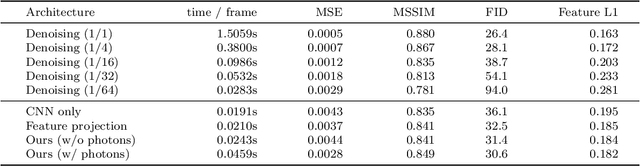
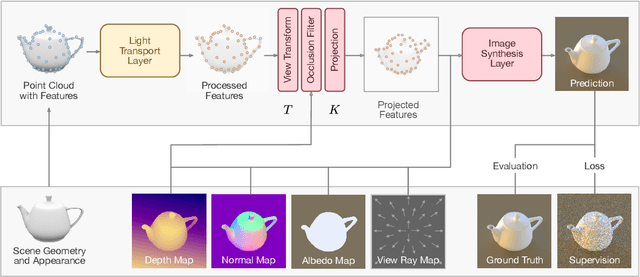
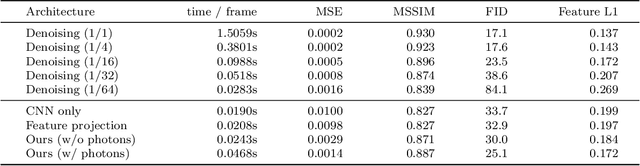
In recent years, deep generative models have gained significance due to their ability to synthesize natural-looking images with applications ranging from virtual reality to data augmentation for training computer vision models. While existing models are able to faithfully learn the image distribution of the training set, they often lack controllability as they operate in 2D pixel space and do not model the physical image formation process. In this work, we investigate the importance of 3D reasoning for photorealistic rendering. We present an approach for learning light transport in static and dynamic 3D scenes using a neural network with the goal of predicting photorealistic images. In contrast to existing approaches that operate in the 2D image domain, our approach reasons in both 3D and 2D space, thus enabling global illumination effects and manipulation of 3D scene geometry. Experimentally, we find that our model is able to produce photorealistic renderings of static and dynamic scenes. Moreover, it compares favorably to baselines which combine path tracing and image denoising at the same computational budget.
Classification of Seeds using Domain Randomization on Self-Supervised Learning Frameworks
Mar 29, 2021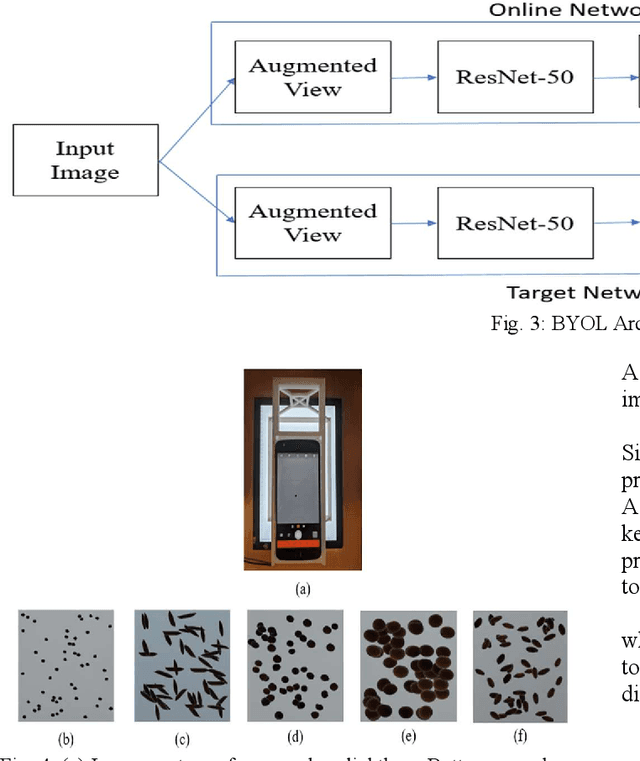
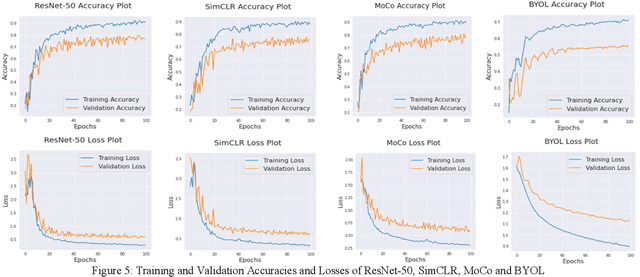
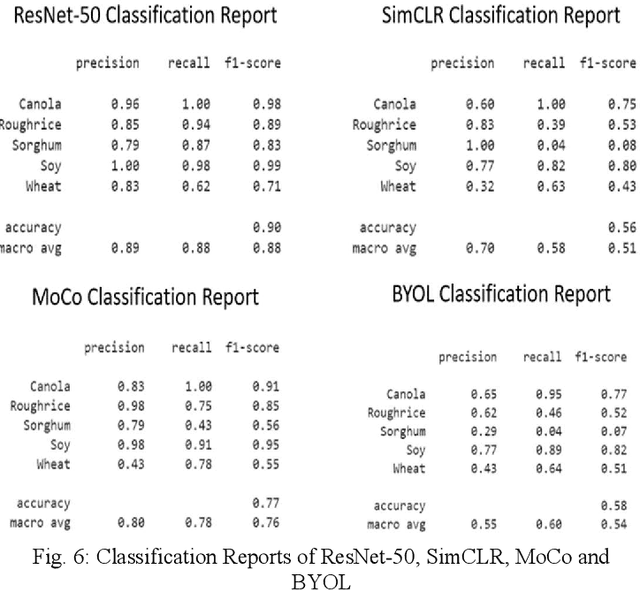
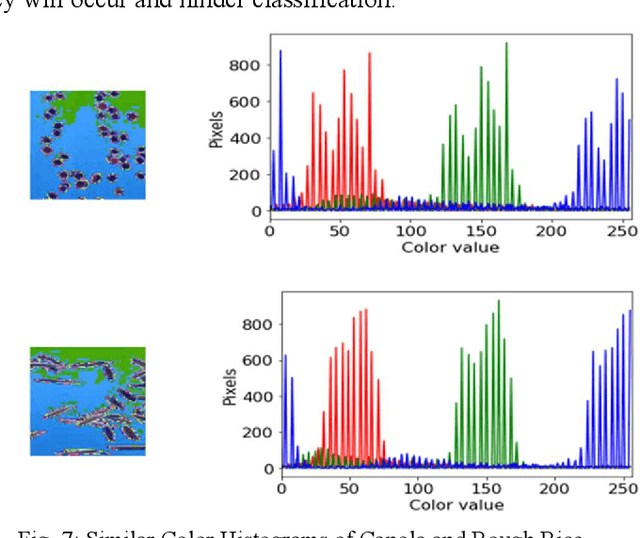
The first step toward Seed Phenotyping i.e. the comprehensive assessment of complex seed traits such as growth, development, tolerance, resistance, ecology, yield, and the measurement of pa-rameters that form more complex traits is the identification of seed type. Generally, a plant re-searcher inspects the visual attributes of a seed such as size, shape, area, color and texture to identify the seed type, a process that is tedious and labor-intensive. Advances in the areas of computer vision and deep learning have led to the development of convolutional neural networks (CNN) that aid in classification using images. While they classify efficiently, a key bottleneck is the need for an extensive amount of labelled data to train the CNN before it can be put to the task of classification. The work leverages the concepts of Contrastive Learning and Domain Randomi-zation in order to achieve the same. Briefly, domain randomization is the technique of applying models trained on images containing simulated objects to real-world objects. The use of synthetic images generated from a representational sample crop of real-world images alleviates the need for a large volume of test subjects. As part of the work, synthetic image datasets of five different types of seed images namely, canola, rough rice, sorghum, soy and wheat are applied to three different self-supervised learning frameworks namely, SimCLR, Momentum Contrast (MoCo) and Build Your Own Latent (BYOL) where ResNet-50 is used as the backbone in each of the networks. When the self-supervised models are fine-tuned with only 5% of the labels from the synthetic dataset, results show that MoCo, the model that yields the best performance of the self-supervised learning frameworks in question, achieves an accuracy of 77% on the test dataset which is only ~13% less than the accuracy of 90% achieved by ResNet-50 trained on 100% of the labels.
Boundary-Aware Geometric Encoding for Semantic Segmentation of Point Clouds
Jan 07, 2021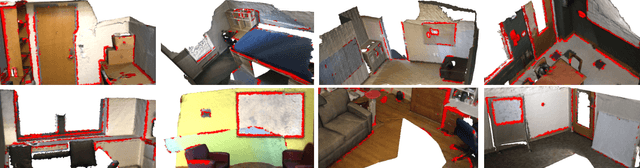
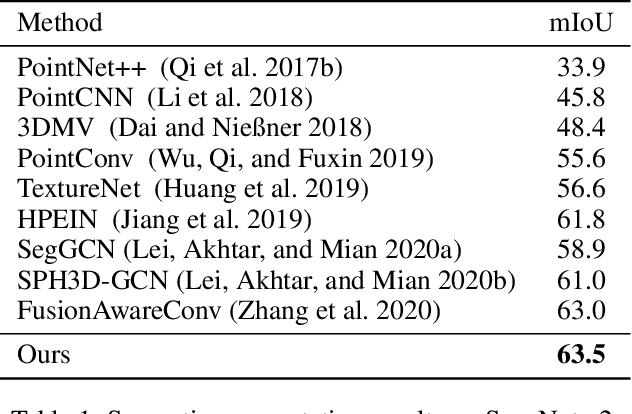
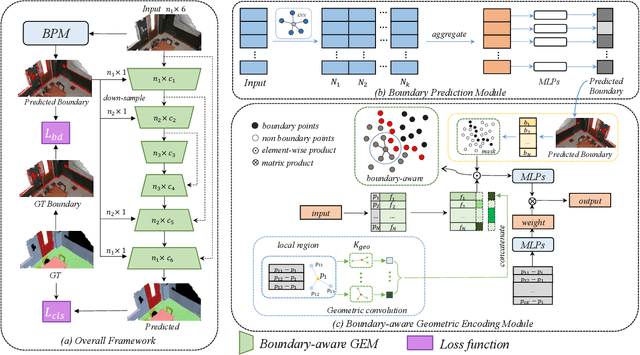
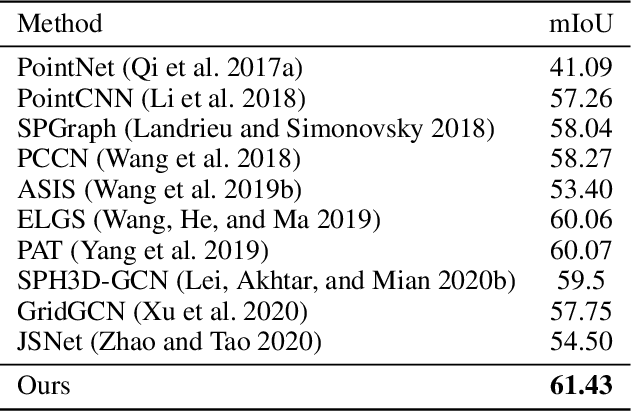
Boundary information plays a significant role in 2D image segmentation, while usually being ignored in 3D point cloud segmentation where ambiguous features might be generated in feature extraction, leading to misclassification in the transition area between two objects. In this paper, firstly, we propose a Boundary Prediction Module (BPM) to predict boundary points. Based on the predicted boundary, a boundary-aware Geometric Encoding Module (GEM) is designed to encode geometric information and aggregate features with discrimination in a neighborhood, so that the local features belonging to different categories will not be polluted by each other. To provide extra geometric information for boundary-aware GEM, we also propose a light-weight Geometric Convolution Operation (GCO), making the extracted features more distinguishing. Built upon the boundary-aware GEM, we build our network and test it on benchmarks like ScanNet v2, S3DIS. Results show our methods can significantly improve the baseline and achieve state-of-the-art performance. Code is available at https://github.com/JchenXu/BoundaryAwareGEM.
Patch-based Texture Synthesis for Image Inpainting
May 05, 2016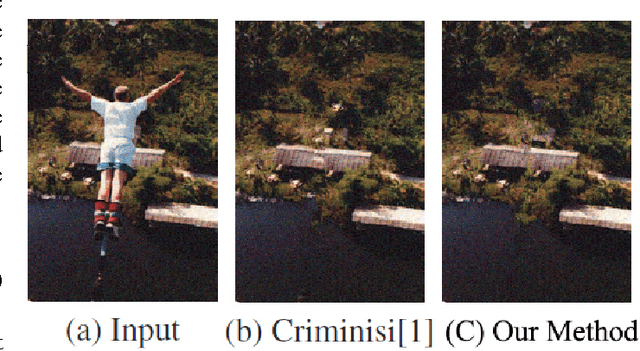
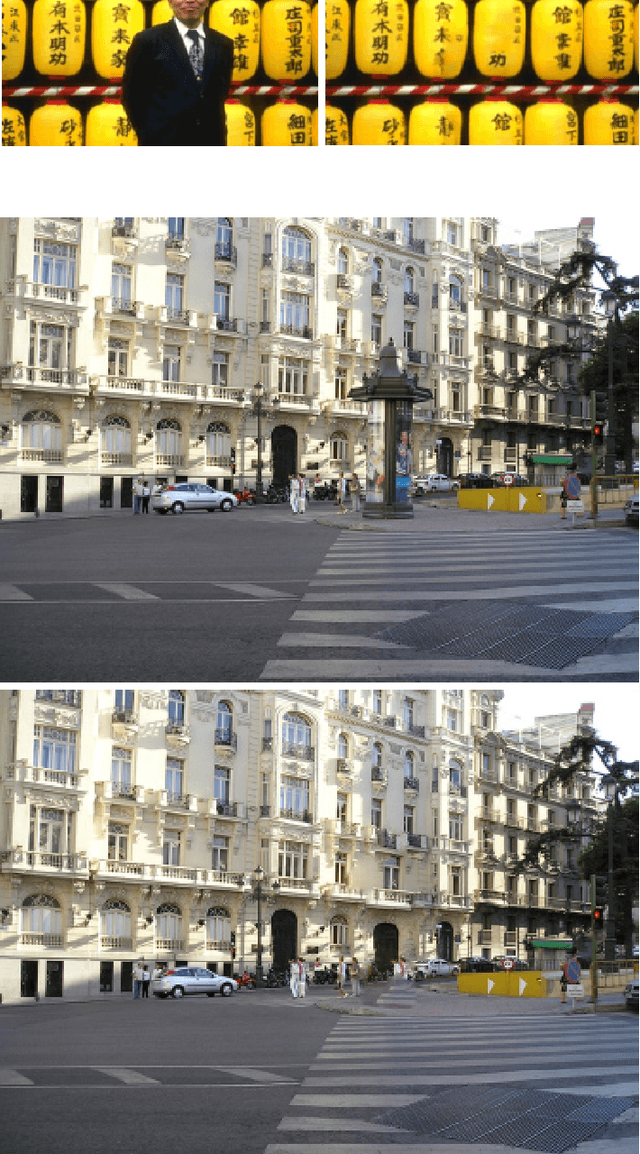
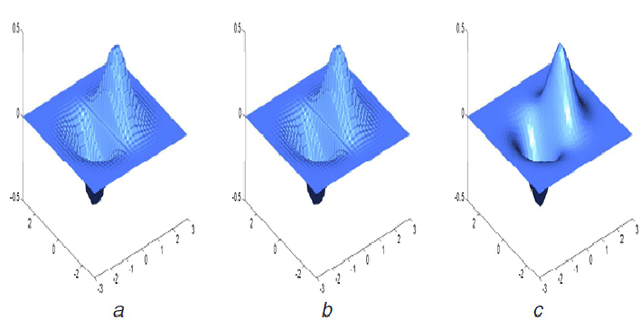

Image inpaiting is an important task in image processing and vision. In this paper, we develop a general method for patch-based image inpainting by synthesizing new textures from existing one. A novel framework is introduced to find several optimal candidate patches and generate a new texture patch in the process. We form it as an optimization problem that identifies the potential patches for synthesis from an coarse-to-fine manner. We use the texture descriptor as a clue in searching for matching patches from the known region. To ensure the structure faithful to the original image, a geometric constraint metric is formally defined that is applied directly to the patch synthesis procedure. We extensively conducted our experiments on a wide range of testing images on various scenarios and contents by arbitrarily specifying the target the regions for inference followed by using existing evaluation metrics to verify its texture coherency and structural consistency. Our results demonstrate the high accuracy and desirable output that can be potentially used for numerous applications: object removal, background subtraction, and image retrieval.
Quantization of Fully Convolutional Networks for Accurate Biomedical Image Segmentation
Mar 13, 2018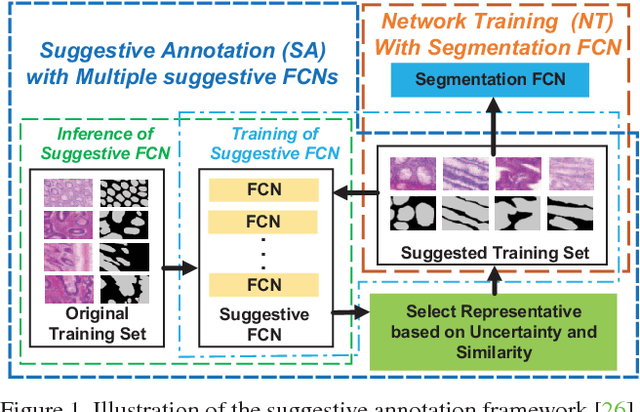
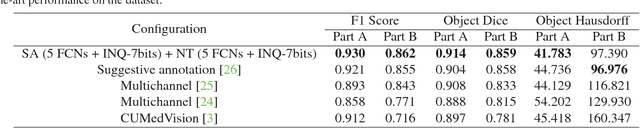
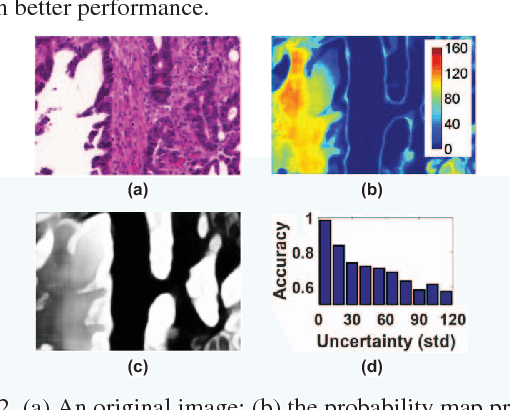
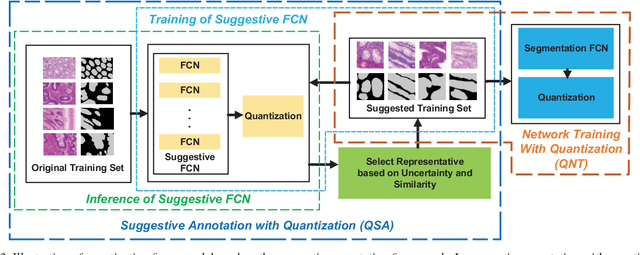
With pervasive applications of medical imaging in health-care, biomedical image segmentation plays a central role in quantitative analysis, clinical diagno- sis, and medical intervention. Since manual anno- tation su ers limited reproducibility, arduous e orts, and excessive time, automatic segmentation is desired to process increasingly larger scale histopathological data. Recently, deep neural networks (DNNs), par- ticularly fully convolutional networks (FCNs), have been widely applied to biomedical image segmenta- tion, attaining much improved performance. At the same time, quantization of DNNs has become an ac- tive research topic, which aims to represent weights with less memory (precision) to considerably reduce memory and computation requirements of DNNs while maintaining acceptable accuracy. In this paper, we apply quantization techniques to FCNs for accurate biomedical image segmentation. Unlike existing litera- ture on quantization which primarily targets memory and computation complexity reduction, we apply quan- tization as a method to reduce over tting in FCNs for better accuracy. Speci cally, we focus on a state-of- the-art segmentation framework, suggestive annotation [22], which judiciously extracts representative annota- tion samples from the original training dataset, obtain- ing an e ective small-sized balanced training dataset. We develop two new quantization processes for this framework: (1) suggestive annotation with quantiza- tion for highly representative training samples, and (2) network training with quantization for high accuracy. Extensive experiments on the MICCAI Gland dataset show that both quantization processes can improve the segmentation performance, and our proposed method exceeds the current state-of-the-art performance by up to 1%. In addition, our method has a reduction of up to 6.4x on memory usage.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge