Hwee Kuan Lee
Prefix-Tuning+: Modernizing Prefix-Tuning through Attention Independent Prefix Data
Jun 16, 2025Abstract:Parameter-Efficient Fine-Tuning (PEFT) methods have become crucial for rapidly adapting large language models (LLMs) to downstream tasks. Prefix-Tuning, an early and effective PEFT technique, demonstrated the ability to achieve performance comparable to full fine-tuning with significantly reduced computational and memory overhead. However, despite its earlier success, its effectiveness in training modern state-of-the-art LLMs has been very limited. In this work, we demonstrate empirically that Prefix-Tuning underperforms on LLMs because of an inherent tradeoff between input and prefix significance within the attention head. This motivates us to introduce Prefix-Tuning+, a novel architecture that generalizes the principles of Prefix-Tuning while addressing its shortcomings by shifting the prefix module out of the attention head itself. We further provide an overview of our construction process to guide future users when constructing their own context-based methods. Our experiments show that, across a diverse set of benchmarks, Prefix-Tuning+ consistently outperforms existing Prefix-Tuning methods. Notably, it achieves performance on par with the widely adopted LoRA method on several general benchmarks, highlighting the potential modern extension of Prefix-Tuning approaches. Our findings suggest that by overcoming its inherent limitations, Prefix-Tuning can remain a competitive and relevant research direction in the landscape of parameter-efficient LLM adaptation.
D-LORD for Motion Stylization
Dec 05, 2024Abstract:This paper introduces a novel framework named D-LORD (Double Latent Optimization for Representation Disentanglement), which is designed for motion stylization (motion style transfer and motion retargeting). The primary objective of this framework is to separate the class and content information from a given motion sequence using a data-driven latent optimization approach. Here, class refers to person-specific style, such as a particular emotion or an individual's identity, while content relates to the style-agnostic aspect of an action, such as walking or jumping, as universally understood concepts. The key advantage of D-LORD is its ability to perform style transfer without needing paired motion data. Instead, it utilizes class and content labels during the latent optimization process. By disentangling the representation, the framework enables the transformation of one motion sequences style to another's style using Adaptive Instance Normalization. The proposed D-LORD framework is designed with a focus on generalization, allowing it to handle different class and content labels for various applications. Additionally, it can generate diverse motion sequences when specific class and content labels are provided. The framework's efficacy is demonstrated through experimentation on three datasets: the CMU XIA dataset for motion style transfer, the MHAD dataset, and the RRIS Ability dataset for motion retargeting. Notably, this paper presents the first generalized framework for motion style transfer and motion retargeting, showcasing its potential contributions in this area.
Exact Conversion of In-Context Learning to Model Weights in Linearized-Attention Transformers
Jun 06, 2024


Abstract:In-Context Learning (ICL) has been a powerful emergent property of large language models that has attracted increasing attention in recent years. In contrast to regular gradient-based learning, ICL is highly interpretable and does not require parameter updates. In this paper, we show that, for linearized transformer networks, ICL can be made explicit and permanent through the inclusion of bias terms. We mathematically demonstrate the equivalence between a model with ICL demonstration prompts and the same model with the additional bias terms. Our algorithm (ICLCA) allows for exact conversion in an inexpensive manner. Existing methods are not exact and require expensive parameter updates. We demonstrate the efficacy of our approach through experiments that show the exact incorporation of ICL tokens into a linear transformer. We further suggest how our method can be adapted to achieve cheap approximate conversion of ICL tokens, even in regular transformer networks that are not linearized. Our experiments on GPT-2 show that, even though the conversion is only approximate, the model still gains valuable context from the included bias terms.
Investigating and unmasking feature-level vulnerabilities of CNNs to adversarial perturbations
May 31, 2024Abstract:This study explores the impact of adversarial perturbations on Convolutional Neural Networks (CNNs) with the aim of enhancing the understanding of their underlying mechanisms. Despite numerous defense methods proposed in the literature, there is still an incomplete understanding of this phenomenon. Instead of treating the entire model as vulnerable, we propose that specific feature maps learned during training contribute to the overall vulnerability. To investigate how the hidden representations learned by a CNN affect its vulnerability, we introduce the Adversarial Intervention framework. Experiments were conducted on models trained on three well-known computer vision datasets, subjecting them to attacks of different nature. Our focus centers on the effects that adversarial perturbations to a model's initial layer have on the overall behavior of the model. Empirical results revealed compelling insights: a) perturbing selected channel combinations in shallow layers causes significant disruptions; b) the channel combinations most responsible for the disruptions are common among different types of attacks; c) despite shared vulnerable combinations of channels, different attacks affect hidden representations with varying magnitudes; d) there exists a positive correlation between a kernel's magnitude and its vulnerability. In conclusion, this work introduces a novel framework to study the vulnerability of a CNN model to adversarial perturbations, revealing insights that contribute to a deeper understanding of the phenomenon. The identified properties pave the way for the development of efficient ad-hoc defense mechanisms in future applications.
Finding Meaningful Distributions of ML Black-boxes under Forensic Investigation
May 10, 2023



Abstract:Given a poorly documented neural network model, we take the perspective of a forensic investigator who wants to find out the model's data domain (e.g. whether on face images or traffic signs). Although existing methods such as membership inference and model inversion can be used to uncover some information about an unknown model, they still require knowledge of the data domain to start with. In this paper, we propose solving this problem by leveraging on comprehensive corpus such as ImageNet to select a meaningful distribution that is close to the original training distribution and leads to high performance in follow-up investigations. The corpus comprises two components, a large dataset of samples and meta information such as hierarchical structure and textual information on the samples. Our goal is to select a set of samples from the corpus for the given model. The core of our method is an objective function that considers two criteria on the selected samples: the model functional properties (derived from the dataset), and semantics (derived from the metadata). We also give an algorithm to efficiently search the large space of all possible subsets w.r.t. the objective function. Experimentation results show that the proposed method is effective. For example, cloning a given model (originally trained with CIFAR-10) by using Caltech 101 can achieve 45.5% accuracy. By using datasets selected by our method, the accuracy is improved to 72.0%.
An End-to-End Breast Tumour Classification Model Using Context-Based Patch Modelling- A BiLSTM Approach for Image Classification
Jun 05, 2021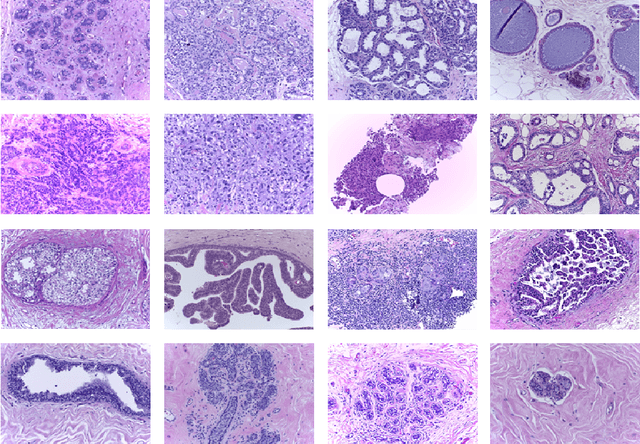
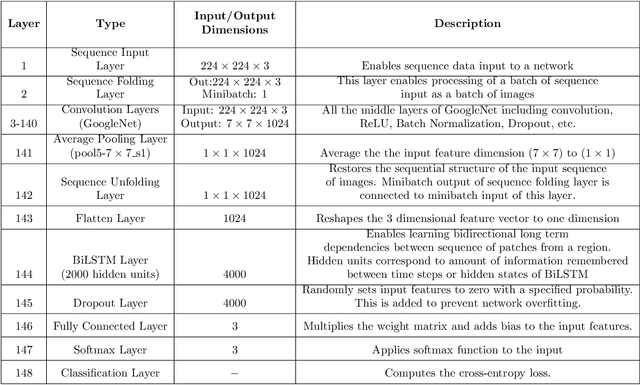
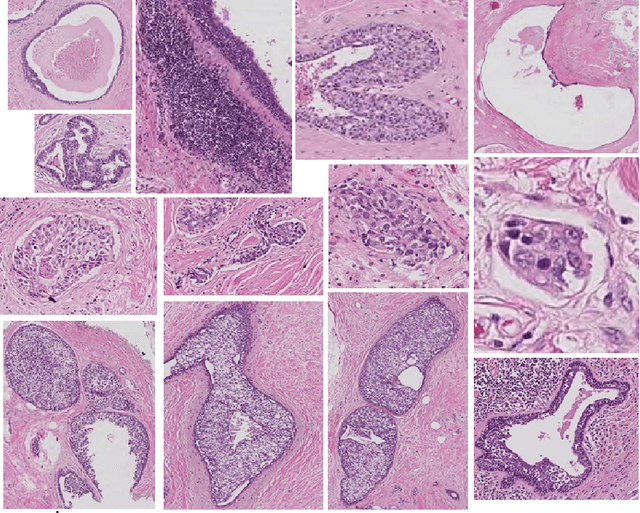
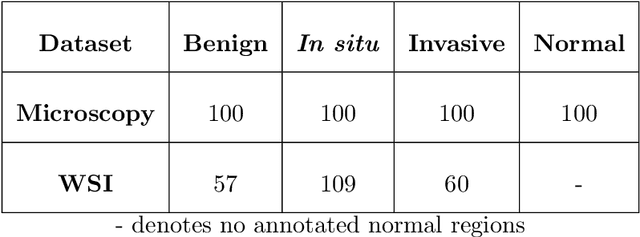
Abstract:Researchers working on computational analysis of Whole Slide Images (WSIs) in histopathology have primarily resorted to patch-based modelling due to large resolution of each WSI. The large resolution makes WSIs infeasible to be fed directly into the machine learning models due to computational constraints. However, due to patch-based analysis, most of the current methods fail to exploit the underlying spatial relationship among the patches. In our work, we have tried to integrate this relationship along with feature-based correlation among the extracted patches from the particular tumorous region. For the given task of classification, we have used BiLSTMs to model both forward and backward contextual relationship. RNN based models eliminate the limitation of sequence size by allowing the modelling of variable size images within a deep learning model. We have also incorporated the effect of spatial continuity by exploring different scanning techniques used to sample patches. To establish the efficiency of our approach, we trained and tested our model on two datasets, microscopy images and WSI tumour regions. After comparing with contemporary literature we achieved the better performance with accuracy of 90% for microscopy image dataset. For WSI tumour region dataset, we compared the classification results with deep learning networks such as ResNet, DenseNet, and InceptionV3 using maximum voting technique. We achieved the highest performance accuracy of 84%. We found out that BiLSTMs with CNN features have performed much better in modelling patches into an end-to-end Image classification network. Additionally, the variable dimensions of WSI tumour regions were used for classification without the need for resizing. This suggests that our method is independent of tumour image size and can process large dimensional images without losing the resolution details.
* 36 pages, 5 figures, 9 tables. Published in Computerized Medical Imaging and Graphics
Automated Deep Learning Analysis of Angiography Video Sequences for Coronary Artery Disease
Jan 29, 2021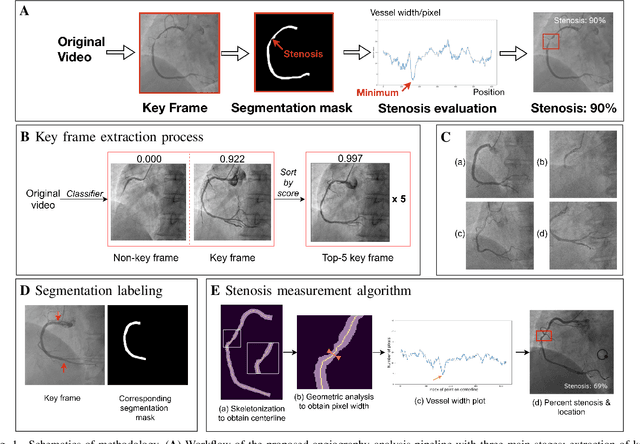
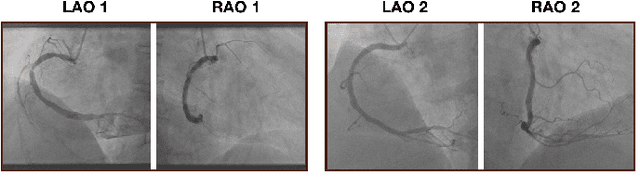
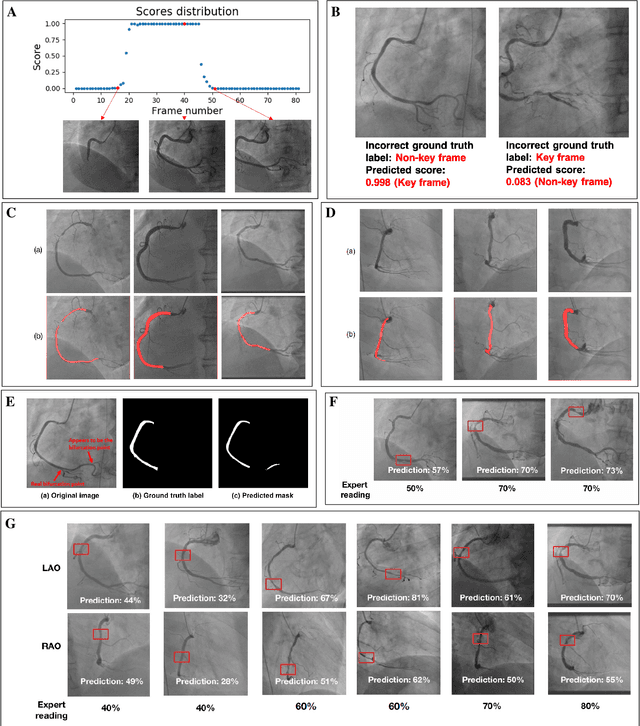
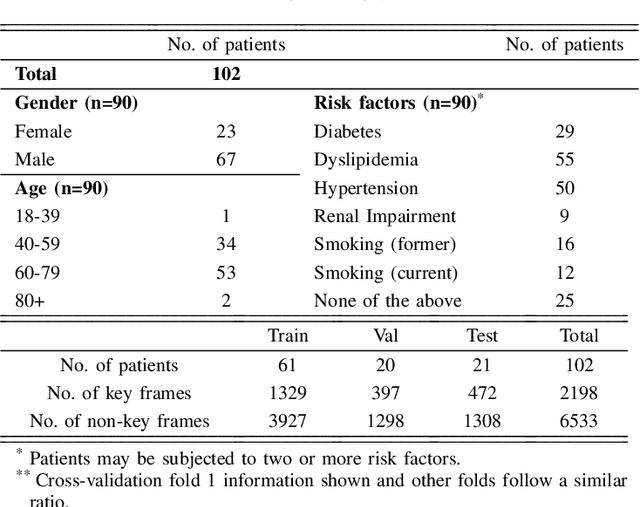
Abstract:The evaluation of obstructions (stenosis) in coronary arteries is currently done by a physician's visual assessment of coronary angiography video sequences. It is laborious, and can be susceptible to interobserver variation. Prior studies have attempted to automate this process, but few have demonstrated an integrated suite of algorithms for the end-to-end analysis of angiograms. We report an automated analysis pipeline based on deep learning to rapidly and objectively assess coronary angiograms, highlight coronary vessels of interest, and quantify potential stenosis. We propose a 3-stage automated analysis method consisting of key frame extraction, vessel segmentation, and stenosis measurement. We combined powerful deep learning approaches such as ResNet and U-Net with traditional image processing and geometrical analysis. We trained and tested our algorithms on the Left Anterior Oblique (LAO) view of the right coronary artery (RCA) using anonymized angiograms obtained from a tertiary cardiac institution, then tested the generalizability of our technique to the Right Anterior Oblique (RAO) view. We demonstrated an overall improvement on previous work, with key frame extraction top-5 precision of 98.4%, vessel segmentation F1-Score of 0.891 and stenosis measurement 20.7% Type I Error rate.
Machine-Learning Study using Improved Correlation Configuration and Application to Quantum Monte Carlo Simulation
Jul 29, 2020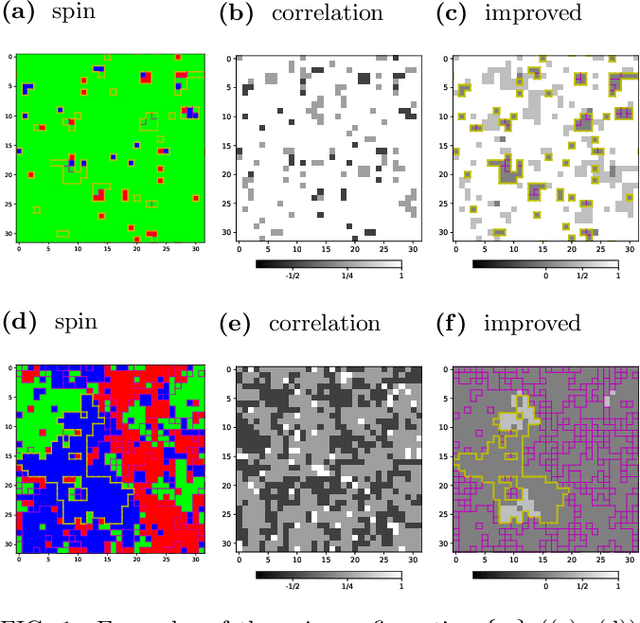
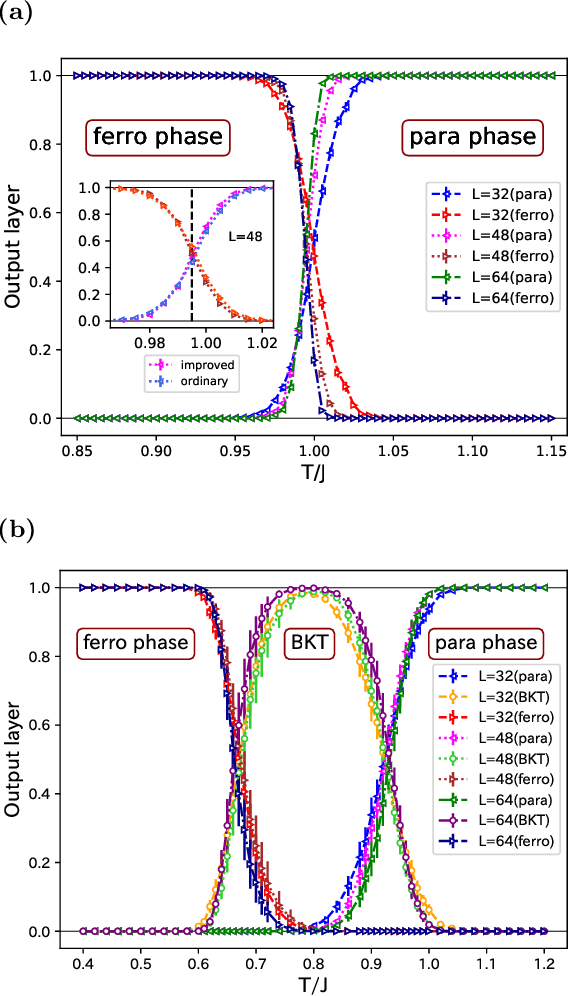
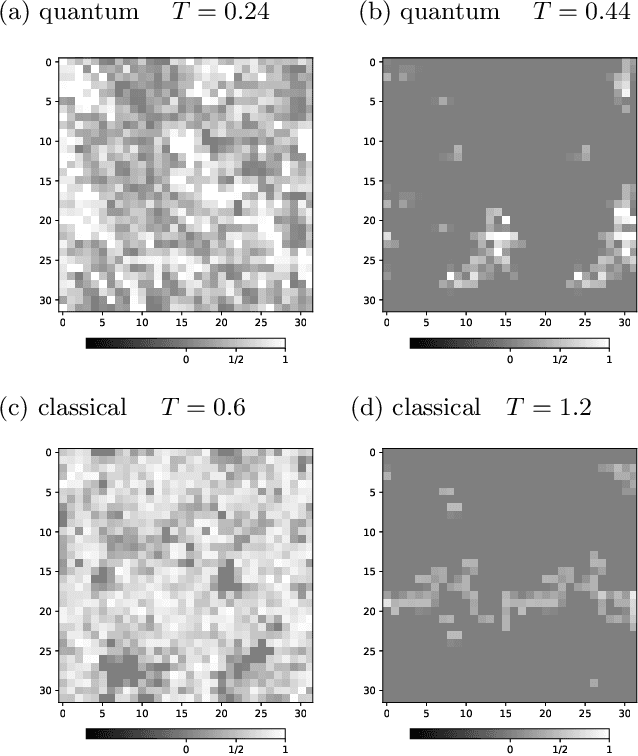
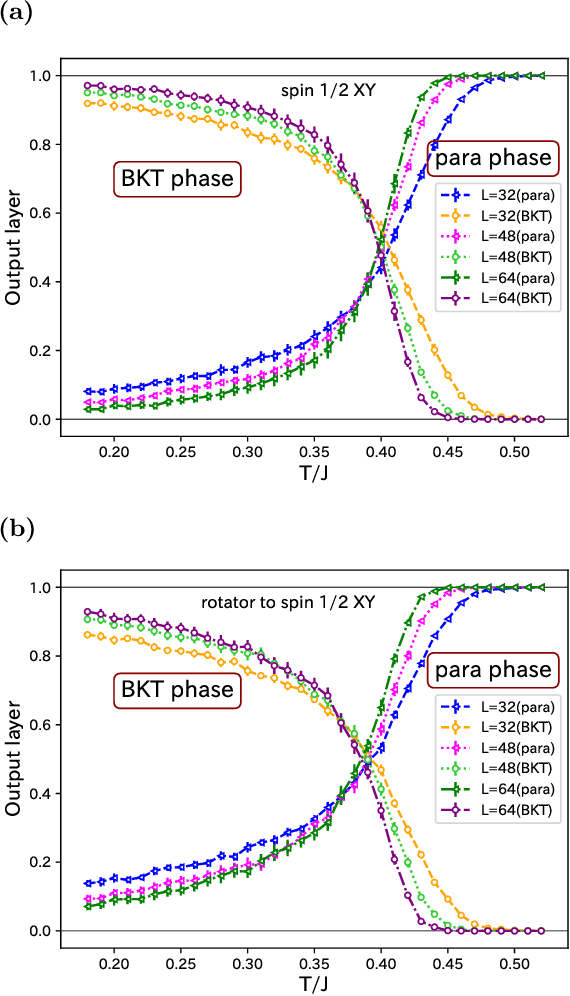
Abstract:We use the Fortuin-Kasteleyn representation based improved estimator of the correlation configuration as an alternative to the ordinary correlation configuration in the machine-learning study of the phase classification of spin models. The phases of classical spin models are classified using the improved estimators, and the method is also applied to the quantum Monte Carlo simulation using the loop algorithm. We analyze the Berezinskii-Kosterlitz-Thouless (BKT) transition of the spin 1/2 quantum XY model on the square lattice. We classify the BKT phase and the paramagnetic phase of the quantum XY model using the machine-learning approach. We show that the classification of the quantum XY model can be performed by using the training data of the classical XY model.
Studying The Effect of MIL Pooling Filters on MIL Tasks
Jun 02, 2020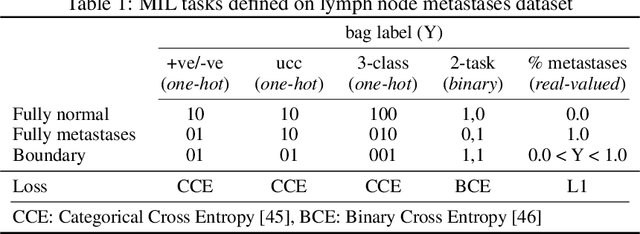
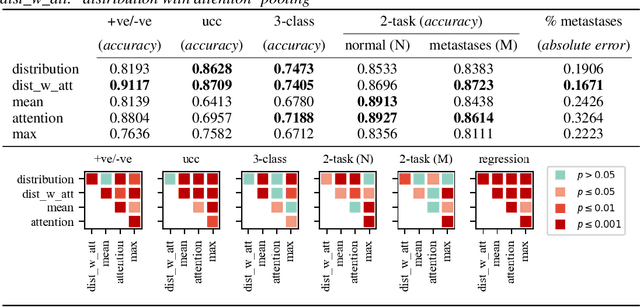
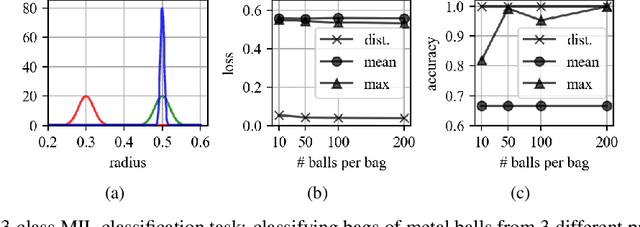
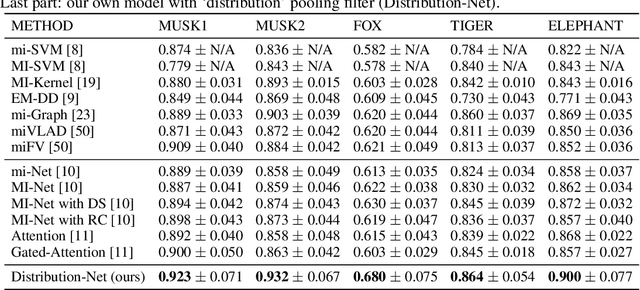
Abstract:There are different multiple instance learning (MIL) pooling filters used in MIL models. In this paper, we study the effect of different MIL pooling filters on the performance of MIL models in real world MIL tasks. We designed a neural network based MIL framework with 5 different MIL pooling filters: `max', `mean', `attention', `distribution' and `distribution with attention'. We also formulated 5 different MIL tasks on a real world lymph node metastases dataset. We found that the performance of our framework in a task is different for different filters. We also observed that the performances of the five pooling filters are also different from task to task. Hence, the selection of a correct MIL pooling filter for each MIL task is crucial for better performance. Furthermore, we noticed that models with `distribution' and `distribution with attention' pooling filters consistently perform well in almost all of the tasks. We attribute this phenomena to the amount of information captured by `distribution' based pooling filters. While point estimate based pooling filters, like `max' and `mean', produce point estimates of distributions, `distribution' based pooling filters capture the full information in distributions. Lastly, we compared the performance of our neural network model with `distribution' pooling filter with the performance of the best MIL methods in the literature on classical MIL datasets and our model outperformed the others.
Detection and Recovery of Adversarial Attacks with Injected Attractors
Mar 05, 2020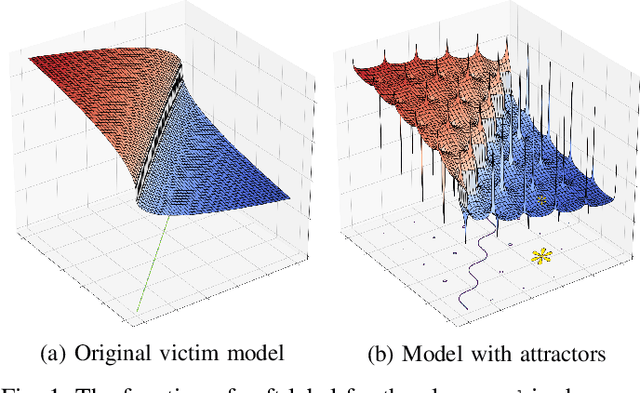
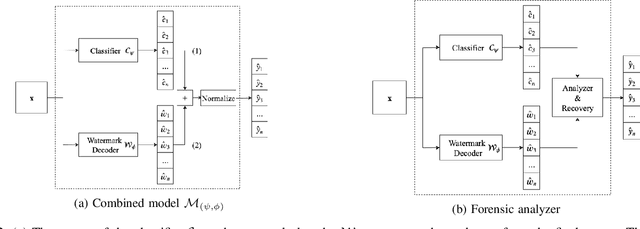
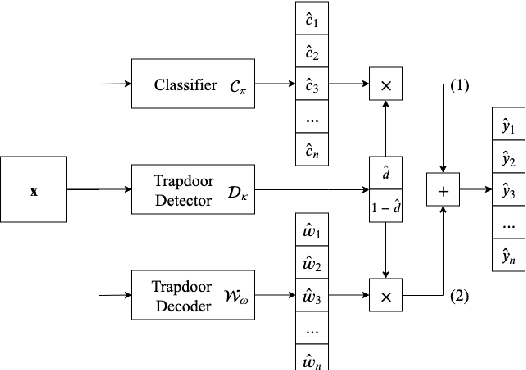
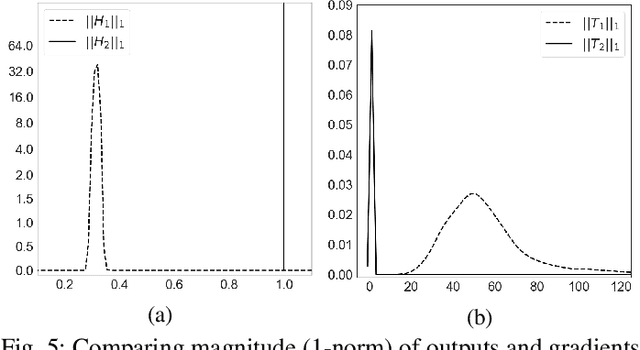
Abstract:Many machine learning adversarial attacks find adversarial samples of a victim model ${\mathcal M}$ by following the gradient of some functions, either explicitly or implicitly. To detect and recover from such attacks, we take the proactive approach that modifies those functions with the goal of misleading the attacks to some local minimals, or to some designated regions that can be easily picked up by a forensic analyzer. To achieve the goal, we propose adding a large number of artifacts, which we called $attractors$, onto the otherwise smooth function. An attractor is a point in the input space, which has a neighborhood of samples with gradients pointing toward it. We observe that decoders of watermarking schemes exhibit properties of attractors, and give a generic method that injects attractors from a watermark decoder into the victim model ${\mathcal M}$. This principled approach allows us to leverage on known watermarking schemes for scalability and robustness. Experimental studies show that our method has competitive performance. For instance, for un-targeted attacks on CIFAR-10 dataset, we can reduce the overall attack success rate of DeepFool to 1.9%, whereas known defence LID, FS and MagNet can reduce the rate to 90.8%, 98.5% and 78.5% respectively.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge