Weikai Huang
Imaginative Perception Tokens Enhance Spatial Reasoning in Multimodal Language Models
Jun 03, 2026Abstract:Vision language models (VLMs) excel at many tasks but still struggle with spatial reasoning when critical information is not directly observable. Many such problems require imaginative perception: inferring what would be seen from an unseen viewpoint, tracing paths through occluded spaces, or integrating partial observations into a coherent spatial representation. We introduce Imaginative Perception Tokens (IPT), intermediate perceptual representations that externalize what a VLM would perceive under alternative spatial configurations while remaining consistent with the observed input. To study this capability, we formulate three tasks, Perspective Taking (PET), Path Tracing (PT), and Multiview Counting (MVC), and construct datasets of approximately 20K examples with ground truth imaginations, answers, and evaluation benchmarks. Using the unified VLM BAGEL as the backbone, IPT supervision consistently improves spatial reasoning and often outperforms textual chain of thought training, even without generating images at inference time. On MVC, IPT improves accuracy by 3.4% and achieves competitive performance with strong closed-source models on PT. We further find that combining IPT and label-only supervision yields additional gains, whereas textual chain of thought can substantially degrade performance, suggesting a modality mismatch when spatial computation is forced through language. Overall, IPT provides a principled supervision signal for reasoning about unobserved spatial structure, improving generalization while producing interpretable intermediate representations.
MolmoAct2: Action Reasoning Models for Real-world Deployment
May 04, 2026Abstract:Vision-Language-Action (VLA) models aim to provide a single generalist controller for robots, but today's systems fall short on the criteria that matter for real-world deployment. Frontier models are closed, open-weight alternatives are tied to expensive hardware, reasoning-augmented policies pay prohibitive latency for their grounding, and fine-tuned success rates remain below the threshold for dependable use. We present MolmoAct2, a fully open action reasoning model built for practical deployment, advancing its predecessor along five axes. We introduce MolmoER, a VLM backbone specialized for spatial and embodied reasoning, trained on a 3.3M-sample corpus with a specialize-then-rehearse recipe. We release three new datasets spanning low-to-medium cost platforms, including MolmoAct2-BimanualYAM, 720 hours of teleoperated bimanual trajectories that constitute the largest open bimanual dataset to date, together with quality-filtered Franka (DROID) and SO100/101 subsets. We provide OpenFAST, an open-weight, open-data action tokenizer trained on millions of trajectories across five embodiments. We redesign the architecture to graft a flow-matching continuous-action expert onto a discrete-token VLM via per-layer KV-cache conditioning. Finally, we propose MolmoThink, an adaptive-depth reasoning variant that re-predicts depth tokens only for scene regions that change between timesteps, retaining geometric grounding at a fraction of prior latency. In the most extensive empirical study of any open VLA to date, spanning 7 simulation and real-world benchmarks, MolmoAct2 outperforms strong baselines including Pi-05, while MolmoER surpasses GPT-5 and Gemini Robotics ER-1.5 across 13 embodied-reasoning benchmarks. We release model weights, training code, and complete training data. Project page: https://allenai.org/blog/molmoact2
WildDet3D: Scaling Promptable 3D Detection in the Wild
Apr 09, 2026Abstract:Understanding objects in 3D from a single image is a cornerstone of spatial intelligence. A key step toward this goal is monocular 3D object detection--recovering the extent, location, and orientation of objects from an input RGB image. To be practical in the open world, such a detector must generalize beyond closed-set categories, support diverse prompt modalities, and leverage geometric cues when available. Progress is hampered by two bottlenecks: existing methods are designed for a single prompt type and lack a mechanism to incorporate additional geometric cues, and current 3D datasets cover only narrow categories in controlled environments, limiting open-world transfer. In this work we address both gaps. First, we introduce WildDet3D, a unified geometry-aware architecture that natively accepts text, point, and box prompts and can incorporate auxiliary depth signals at inference time. Second, we present WildDet3D-Data, the largest open 3D detection dataset to date, constructed by generating candidate 3D boxes from existing 2D annotations and retaining only human-verified ones, yielding over 1M images across 13.5K categories in diverse real-world scenes. WildDet3D establishes a new state-of-the-art across multiple benchmarks and settings. In the open-world setting, it achieves 22.6/24.8 AP3D on our newly introduced WildDet3D-Bench with text and box prompts. On Omni3D, it reaches 34.2/36.4 AP3D with text and box prompts, respectively. In zero-shot evaluation, it achieves 40.3/48.9 ODS on Argoverse 2 and ScanNet. Notably, incorporating depth cues at inference time yields substantial additional gains (+20.7 AP on average across settings).
TrajTok: Learning Trajectory Tokens enables better Video Understanding
Feb 26, 2026Abstract:Tokenization in video models, typically through patchification, generates an excessive and redundant number of tokens. This severely limits video efficiency and scalability. While recent trajectory-based tokenizers offer a promising solution by decoupling video duration from token count, they rely on complex external segmentation and tracking pipelines that are slow and task-agnostic. We propose TrajTok, an end-to-end video tokenizer module that is fully integrated and co-trained with video models for a downstream objective, dynamically adapting its token granularity to semantic complexity, independent of video duration. TrajTok contains a unified segmenter that performs implicit clustering over pixels in both space and time to directly produce object trajectories in a single forward pass. By prioritizing downstream adaptability over pixel-perfect segmentation fidelity, TrajTok is lightweight and efficient, yet empirically improves video understanding performance. With TrajTok, we implement a video CLIP model trained from scratch (TrajViT2). It achieves the best accuracy at scale across both classification and retrieval benchmarks, while maintaining efficiency comparable to the best token-merging methods. TrajTok also proves to be a versatile component beyond its role as a tokenizer. We show that it can be seamlessly integrated as either a probing head for pretrained visual features (TrajAdapter) or an alignment connector in vision-language models (TrajVLM) with especially strong performance in long-video reasoning.
Molmo2: Open Weights and Data for Vision-Language Models with Video Understanding and Grounding
Jan 15, 2026Abstract:Today's strongest video-language models (VLMs) remain proprietary. The strongest open-weight models either rely on synthetic data from proprietary VLMs, effectively distilling from them, or do not disclose their training data or recipe. As a result, the open-source community lacks the foundations needed to improve on the state-of-the-art video (and image) language models. Crucially, many downstream applications require more than just high-level video understanding; they require grounding -- either by pointing or by tracking in pixels. Even proprietary models lack this capability. We present Molmo2, a new family of VLMs that are state-of-the-art among open-source models and demonstrate exceptional new capabilities in point-driven grounding in single image, multi-image, and video tasks. Our key contribution is a collection of 7 new video datasets and 2 multi-image datasets, including a dataset of highly detailed video captions for pre-training, a free-form video Q&A dataset for fine-tuning, a new object tracking dataset with complex queries, and an innovative new video pointing dataset, all collected without the use of closed VLMs. We also present a training recipe for this data utilizing an efficient packing and message-tree encoding scheme, and show bi-directional attention on vision tokens and a novel token-weight strategy improves performance. Our best-in-class 8B model outperforms others in the class of open weight and data models on short videos, counting, and captioning, and is competitive on long-videos. On video-grounding Molmo2 significantly outperforms existing open-weight models like Qwen3-VL (35.5 vs 29.6 accuracy on video counting) and surpasses proprietary models like Gemini 3 Pro on some tasks (38.4 vs 20.0 F1 on video pointing and 56.2 vs 41.1 J&F on video tracking).
Generate Any Scene: Evaluating and Improving Text-to-Vision Generation with Scene Graph Programming
Dec 11, 2024



Abstract:DALL-E and Sora have gained attention by producing implausible images, such as "astronauts riding a horse in space." Despite the proliferation of text-to-vision models that have inundated the internet with synthetic visuals, from images to 3D assets, current benchmarks predominantly evaluate these models on real-world scenes paired with captions. We introduce Generate Any Scene, a framework that systematically enumerates scene graphs representing a vast array of visual scenes, spanning realistic to imaginative compositions. Generate Any Scene leverages 'scene graph programming', a method for dynamically constructing scene graphs of varying complexity from a structured taxonomy of visual elements. This taxonomy includes numerous objects, attributes, and relations, enabling the synthesis of an almost infinite variety of scene graphs. Using these structured representations, Generate Any Scene translates each scene graph into a caption, enabling scalable evaluation of text-to-vision models through standard metrics. We conduct extensive evaluations across multiple text-to-image, text-to-video, and text-to-3D models, presenting key findings on model performance. We find that DiT-backbone text-to-image models align more closely with input captions than UNet-backbone models. Text-to-video models struggle with balancing dynamics and consistency, while both text-to-video and text-to-3D models show notable gaps in human preference alignment. We demonstrate the effectiveness of Generate Any Scene by conducting three practical applications leveraging captions generated by Generate Any Scene: 1) a self-improving framework where models iteratively enhance their performance using generated data, 2) a distillation process to transfer specific strengths from proprietary models to open-source counterparts, and 3) improvements in content moderation by identifying and generating challenging synthetic data.
ProVision: Programmatically Scaling Vision-centric Instruction Data for Multimodal Language Models
Dec 09, 2024



Abstract:With the rise of multimodal applications, instruction data has become critical for training multimodal language models capable of understanding complex image-based queries. Existing practices rely on powerful but costly large language models (LLMs) or multimodal language models (MLMs) to produce instruction data. These are often prone to hallucinations, licensing issues and the generation process is often hard to scale and interpret. In this work, we present a programmatic approach that employs scene graphs as symbolic representations of images and human-written programs to systematically synthesize vision-centric instruction data. Our approach ensures the interpretability and controllability of the data generation process and scales efficiently while maintaining factual accuracy. By implementing a suite of 24 single-image, 14 multi-image instruction generators, and a scene graph generation pipeline, we build a scalable, cost-effective system: ProVision which produces diverse question-answer pairs concerning objects, attributes, relations, depth, etc., for any given image. Applied to Visual Genome and DataComp datasets, we generate over 10 million instruction data points, ProVision-10M, and leverage them in both pretraining and instruction tuning stages of MLMs. When adopted in the instruction tuning stage, our single-image instruction data yields up to a 7% improvement on the 2D split and 8% on the 3D split of CVBench, along with a 3% increase in performance on QBench2, RealWorldQA, and MMMU. Our multi-image instruction data leads to an 8% improvement on Mantis-Eval. Incorporation of our data in both pre-training and fine-tuning stages of xGen-MM-4B leads to an averaged improvement of 1.6% across 11 benchmarks.
Task Me Anything
Jun 17, 2024



Abstract:Benchmarks for large multimodal language models (MLMs) now serve to simultaneously assess the general capabilities of models instead of evaluating for a specific capability. As a result, when a developer wants to identify which models to use for their application, they are overwhelmed by the number of benchmarks and remain uncertain about which benchmark's results are most reflective of their specific use case. This paper introduces Task-Me-Anything, a benchmark generation engine which produces a benchmark tailored to a user's needs. Task-Me-Anything maintains an extendable taxonomy of visual assets and can programmatically generate a vast number of task instances. Additionally, it algorithmically addresses user queries regarding MLM performance efficiently within a computational budget. It contains 113K images, 10K videos, 2K 3D object assets, over 365 object categories, 655 attributes, and 335 relationships. It can generate 750M image/video question-answering pairs, which focus on evaluating MLM perceptual capabilities. Task-Me-Anything reveals critical insights: open-source MLMs excel in object and attribute recognition but lack spatial and temporal understanding; each model exhibits unique strengths and weaknesses; larger models generally perform better, though exceptions exist; and GPT4o demonstrates challenges in recognizing rotating/moving objects and distinguishing colors.
m&m's: A Benchmark to Evaluate Tool-Use for multi-step multi-modal Tasks
Mar 21, 2024Abstract:Real-world multi-modal problems are rarely solved by a single machine learning model, and often require multi-step computational plans that involve stitching several models. Tool-augmented LLMs hold tremendous promise for automating the generation of such computational plans. However, the lack of standardized benchmarks for evaluating LLMs as planners for multi-step multi-modal tasks has prevented a systematic study of planner design decisions. Should LLMs generate a full plan in a single shot or step-by-step? Should they invoke tools directly with Python code or through structured data formats like JSON? Does feedback improve planning? To answer these questions and more, we introduce m&m's: a benchmark containing 4K+ multi-step multi-modal tasks involving 33 tools that include multi-modal models, (free) public APIs, and image processing modules. For each of these task queries, we provide automatically generated plans using this realistic toolset. We further provide a high-quality subset of 1,565 task plans that are human-verified and correctly executable. With m&m's, we evaluate 6 popular LLMs with 2 planning strategies (multi-step vs. step-by-step planning), 2 plan formats (JSON vs. code), and 3 types of feedback (parsing/verification/execution). Finally, we summarize takeaways from our extensive experiments. Our dataset and code are available on HuggingFace (https://huggingface.co/datasets/zixianma/mnms) and Github (https://github.com/RAIVNLab/mnms).
LesionPaste: One-Shot Anomaly Detection for Medical Images
Mar 12, 2022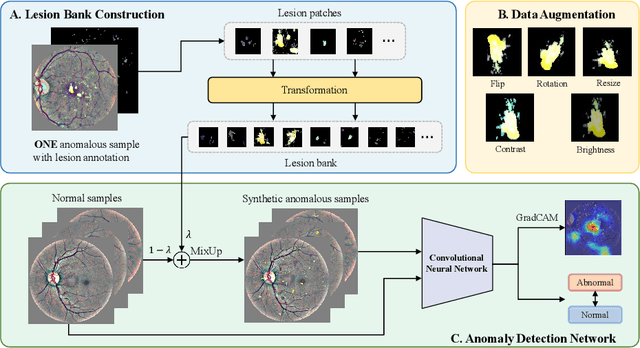
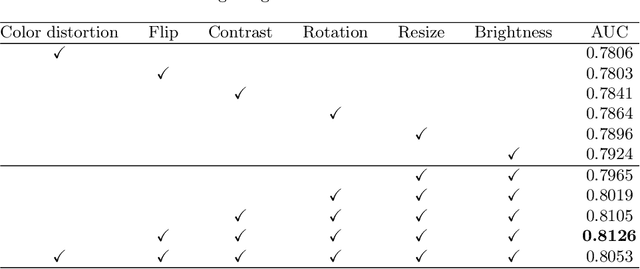
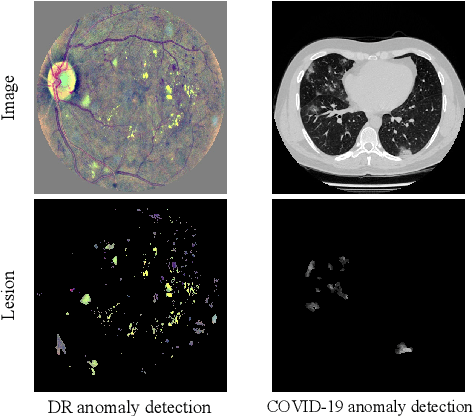
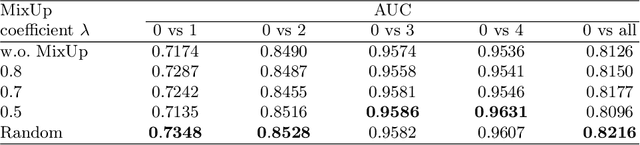
Abstract:Due to the high cost of manually annotating medical images, especially for large-scale datasets, anomaly detection has been explored through training models with only normal data. Lacking prior knowledge of true anomalies is the main reason for the limited application of previous anomaly detection methods, especially in the medical image analysis realm. In this work, we propose a one-shot anomaly detection framework, namely LesionPaste, that utilizes true anomalies from a single annotated sample and synthesizes artificial anomalous samples for anomaly detection. First, a lesion bank is constructed by applying augmentation to randomly selected lesion patches. Then, MixUp is adopted to paste patches from the lesion bank at random positions in normal images to synthesize anomalous samples for training. Finally, a classification network is trained using the synthetic abnormal samples and the true normal data. Extensive experiments are conducted on two publicly-available medical image datasets with different types of abnormalities. On both datasets, our proposed LesionPaste largely outperforms several state-of-the-art unsupervised and semi-supervised anomaly detection methods, and is on a par with the fully-supervised counterpart. To note, LesionPaste is even better than the fully-supervised method in detecting early-stage diabetic retinopathy.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge