Nassir Navab
Computer Aided Medical Procedures, Technische Universit Munchen, Germany, Johns Hopkins University, Baltimore MD, USA
HouseCat6D -- A Large-Scale Multi-Modal Category Level 6D Object Pose Dataset with Household Objects in Realistic Scenarios
Dec 21, 2022Abstract:Estimating the 6D pose of objects is one of the major fields in 3D computer vision. Since the promising outcomes from instance-level pose estimation, the research trends are heading towards category-level pose estimation for more practical application scenarios. However, unlike well-established instance-level pose datasets, available category-level datasets lack annotation quality and provided pose quantity. We propose the new category level 6D pose dataset HouseCat6D featuring 1) Multi-modality of Polarimetric RGB+P and Depth, 2) Highly diverse 194 objects of 10 household object categories including 2 photometrically challenging categories, 3) High-quality pose annotation with an error range of only 1.35 mm to 1.74 mm, 4) 41 large scale scenes with extensive viewpoint coverage, 5) Checkerboard-free environment throughout the entire scene. We also provide benchmark results of state-of-the-art category-level pose estimation networks.
DisPositioNet: Disentangled Pose and Identity in Semantic Image Manipulation
Nov 10, 2022



Abstract:Graph representation of objects and their relations in a scene, known as a scene graph, provides a precise and discernible interface to manipulate a scene by modifying the nodes or the edges in the graph. Although existing works have shown promising results in modifying the placement and pose of objects, scene manipulation often leads to losing some visual characteristics like the appearance or identity of objects. In this work, we propose DisPositioNet, a model that learns a disentangled representation for each object for the task of image manipulation using scene graphs in a self-supervised manner. Our framework enables the disentanglement of the variational latent embeddings as well as the feature representation in the graph. In addition to producing more realistic images due to the decomposition of features like pose and identity, our method takes advantage of the probabilistic sampling in the intermediate features to generate more diverse images in object replacement or addition tasks. The results of our experiments show that disentangling the feature representations in the latent manifold of the model outperforms the previous works qualitatively and quantitatively on two public benchmarks. Project Page: https://scenegenie.github.io/DispositioNet/
Improved Techniques for the Conditional Generative Augmentation of Clinical Audio Data
Nov 05, 2022



Abstract:Data augmentation is a valuable tool for the design of deep learning systems to overcome data limitations and stabilize the training process. Especially in the medical domain, where the collection of large-scale data sets is challenging and expensive due to limited access to patient data, relevant environments, as well as strict regulations, community-curated large-scale public datasets, pretrained models, and advanced data augmentation methods are the main factors for developing reliable systems to improve patient care. However, for the development of medical acoustic sensing systems, an emerging field of research, the community lacks large-scale publicly available data sets and pretrained models. To address the problem of limited data, we propose a conditional generative adversarial neural network-based augmentation method which is able to synthesize mel spectrograms from a learned data distribution of a source data set. In contrast to previously proposed fully convolutional models, the proposed model implements residual Squeeze and Excitation modules in the generator architecture. We show that our method outperforms all classical audio augmentation techniques and previously published generative methods in terms of generated sample quality and a performance improvement of 2.84% of Macro F1-Score for a classifier trained on the augmented data set, an enhancement of $1.14\%$ in relation to previous work. By analyzing the correlation of intermediate feature spaces, we show that the residual Squeeze and Excitation modules help the model to reduce redundancy in the latent features. Therefore, the proposed model advances the state-of-the-art in the augmentation of clinical audio data and improves the data bottleneck for the design of clinical acoustic sensing systems.
What can we learn about a generated image corrupting its latent representation?
Oct 12, 2022Abstract:Generative adversarial networks (GANs) offer an effective solution to the image-to-image translation problem, thereby allowing for new possibilities in medical imaging. They can translate images from one imaging modality to another at a low cost. For unpaired datasets, they rely mostly on cycle loss. Despite its effectiveness in learning the underlying data distribution, it can lead to a discrepancy between input and output data. The purpose of this work is to investigate the hypothesis that we can predict image quality based on its latent representation in the GANs bottleneck. We achieve this by corrupting the latent representation with noise and generating multiple outputs. The degree of differences between them is interpreted as the strength of the representation: the more robust the latent representation, the fewer changes in the output image the corruption causes. Our results demonstrate that our proposed method has the ability to i) predict uncertain parts of synthesized images, and ii) identify samples that may not be reliable for downstream tasks, e.g., liver segmentation task.
MonoGraspNet: 6-DoF Grasping with a Single RGB Image
Sep 26, 2022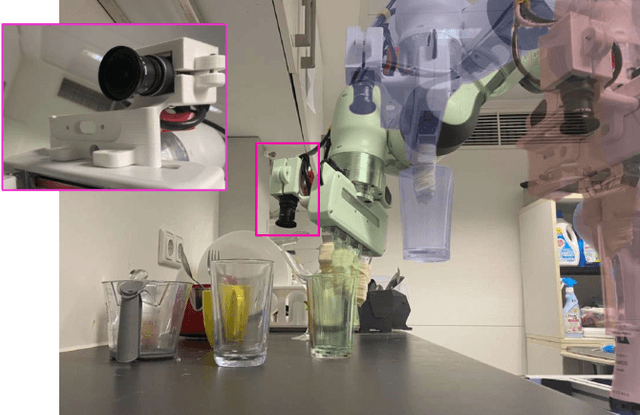
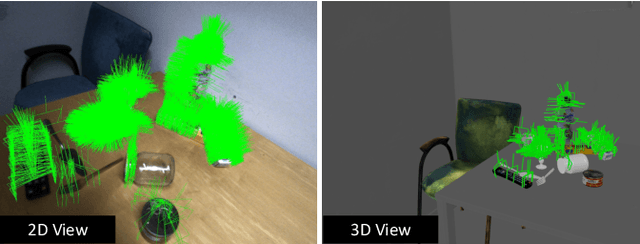
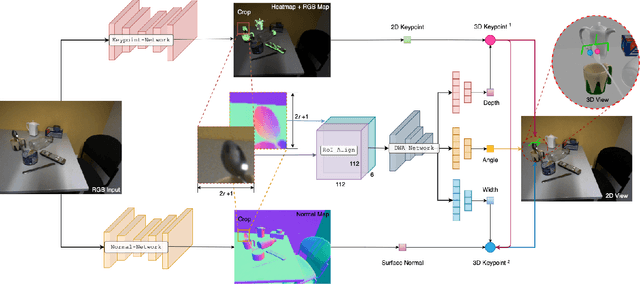
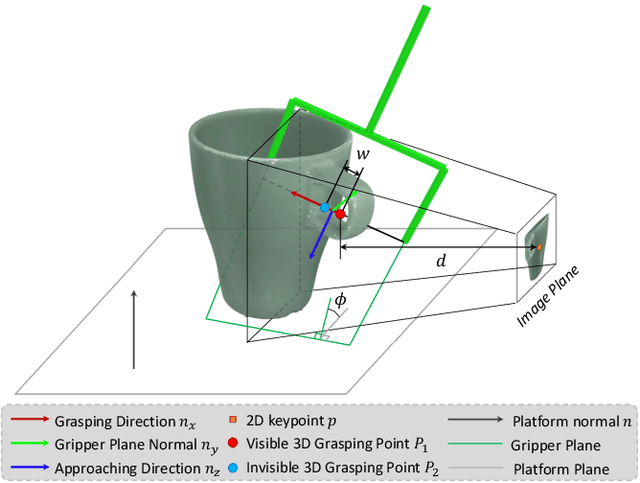

Abstract:6-DoF robotic grasping is a long-lasting but unsolved problem. Recent methods utilize strong 3D networks to extract geometric grasping representations from depth sensors, demonstrating superior accuracy on common objects but perform unsatisfactorily on photometrically challenging objects, e.g., objects in transparent or reflective materials. The bottleneck lies in that the surface of these objects can not reflect back accurate depth due to the absorption or refraction of light. In this paper, in contrast to exploiting the inaccurate depth data, we propose the first RGB-only 6-DoF grasping pipeline called MonoGraspNet that utilizes stable 2D features to simultaneously handle arbitrary object grasping and overcome the problems induced by photometrically challenging objects. MonoGraspNet leverages keypoint heatmap and normal map to recover the 6-DoF grasping poses represented by our novel representation parameterized with 2D keypoints with corresponding depth, grasping direction, grasping width, and angle. Extensive experiments in real scenes demonstrate that our method can achieve competitive results in grasping common objects and surpass the depth-based competitor by a large margin in grasping photometrically challenging objects. To further stimulate robotic manipulation research, we additionally annotate and open-source a multi-view and multi-scene real-world grasping dataset, containing 120 objects of mixed photometric complexity with 20M accurate grasping labels.
Holistic Segmentation
Sep 12, 2022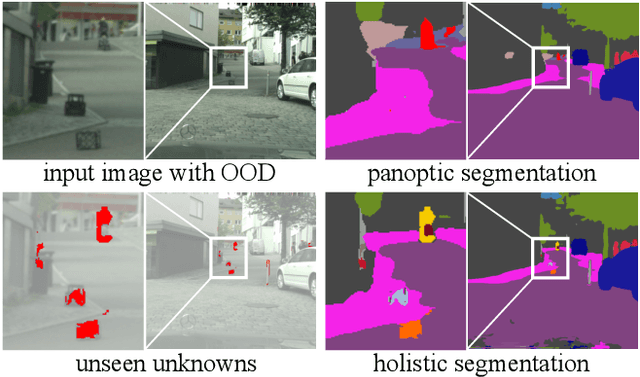
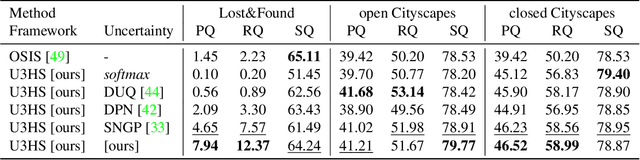
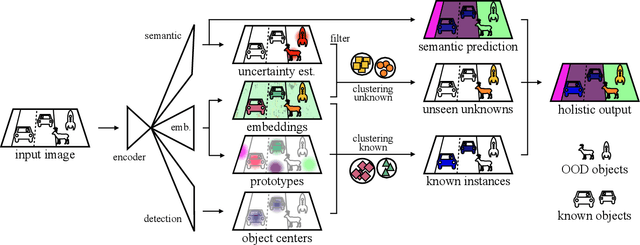
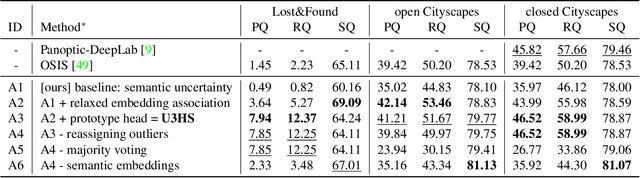
Abstract:As panoptic segmentation provides a prediction for every pixel in input, non-standard and unseen objects systematically lead to wrong outputs. However, in safety-critical settings, robustness against out-of-distribution samples and corner cases is crucial to avoid dangerous behaviors, such as ignoring an animal or a lost cargo on the road. Since driving datasets cannot contain enough data points to properly sample the long tail of the underlying distribution, a method must deal with unknown and unseen scenarios to be deployed safely. Previous methods targeted part of this issue, by re-identifying already seen unlabeled objects. In this work, we broaden the scope proposing holistic segmentation: a task to identify and separate unseen unknown objects into instances, without learning from unknowns, while performing panoptic segmentation of known classes. We tackle this new problem with U3HS, which first finds unknowns as highly uncertain regions, then clusters the corresponding instance-aware embeddings into individual objects. By doing so, for the first time in panoptic segmentation with unknown objects, our U3HS is not trained with unknown data, thus leaving the settings unconstrained with respect to the type of objects and allowing for a holistic scene understanding. Extensive experiments and comparisons on two public datasets, namely Cityscapes and Lost&Found as a transfer, demonstrate the effectiveness of U3HS in the challenging task of holistic segmentation, with competitive closed-set panoptic segmentation performance.
STTAR: Surgical Tool Tracking using off-the-shelf Augmented Reality Head-Mounted Displays
Aug 17, 2022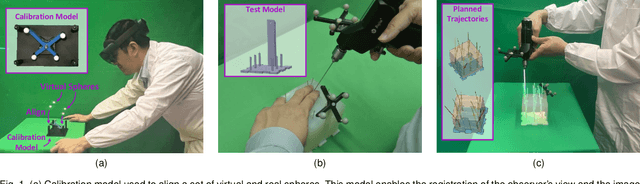
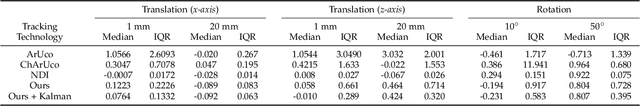
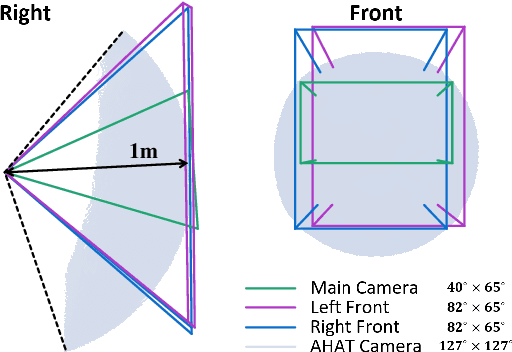
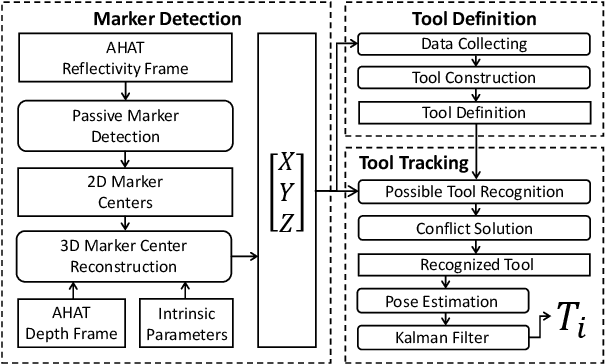
Abstract:The use of Augmented Reality (AR) for navigation purposes has shown beneficial in assisting physicians during the performance of surgical procedures. These applications commonly require knowing the pose of surgical tools and patients to provide visual information that surgeons can use during the task performance. Existing medical-grade tracking systems use infrared cameras placed inside the Operating Room (OR) to identify retro-reflective markers attached to objects of interest and compute their pose. Some commercially available AR Head-Mounted Displays (HMDs) use similar cameras for self-localization, hand tracking, and estimating the objects' depth. This work presents a framework that uses the built-in cameras of AR HMDs to enable accurate tracking of retro-reflective markers, such as those used in surgical procedures, without the need to integrate any additional components. This framework is also capable of simultaneously tracking multiple tools. Our results show that the tracking and detection of the markers can be achieved with an accuracy of 0.09 +- 0.06 mm on lateral translation, 0.42 +- 0.32 mm on longitudinal translation, and 0.80 +- 0.39 deg for rotations around the vertical axis. Furthermore, to showcase the relevance of the proposed framework, we evaluate the system's performance in the context of surgical procedures. This use case was designed to replicate the scenarios of k-wire insertions in orthopedic procedures. For evaluation, two surgeons and one biomedical researcher were provided with visual navigation, each performing 21 injections. Results from this use case provide comparable accuracy to those reported in the literature for AR-based navigation procedures.
Towards Autonomous Atlas-based Ultrasound Acquisitions in Presence of Articulated Motion
Aug 10, 2022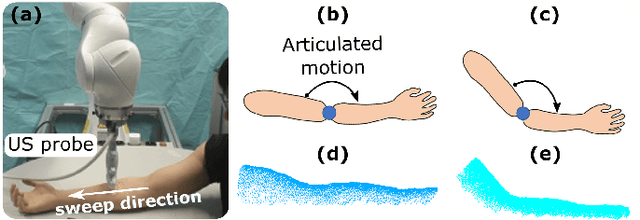
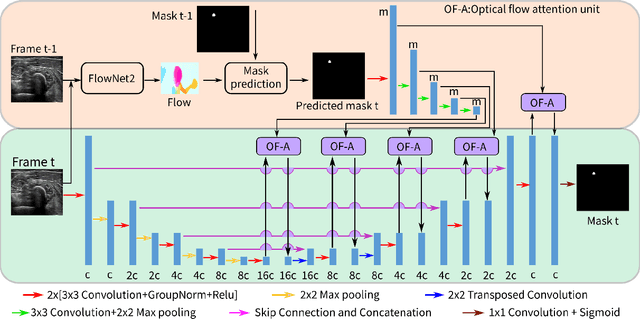
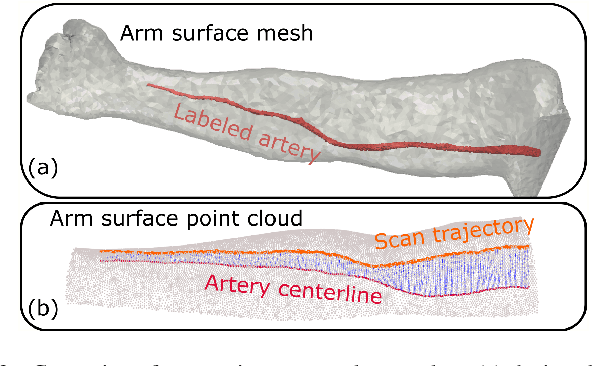
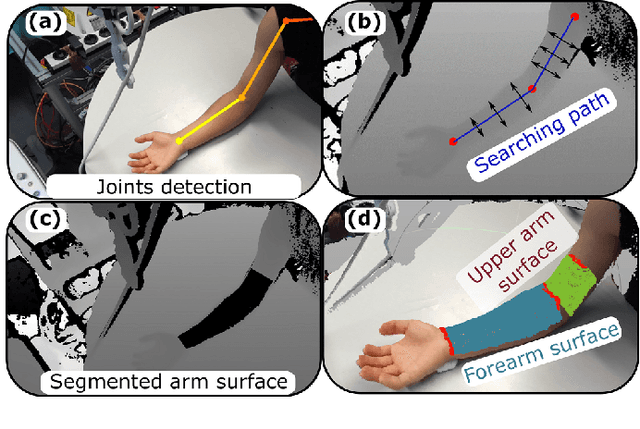
Abstract:Robotic ultrasound (US) imaging aims at overcoming some of the limitations of free-hand US examinations, e.g. difficulty in guaranteeing intra- and inter-operator repeatability. However, due to anatomical and physiological variations between patients and relative movement of anatomical substructures, it is challenging to robustly generate optimal trajectories to examine the anatomies of interest, in particular, when they comprise articulated joints. To address this challenge, this paper proposes a vision-based approach allowing autonomous robotic US limb scanning. To this end, an atlas MRI template of a human arm with annotated vascular structures is used to generate trajectories and register and project them onto patients' skin surfaces for robotic US acquisition. To effectively segment and accurately reconstruct the targeted 3D vessel, we make use of spatial continuity in consecutive US frames by incorporating channel attention modules into a U-Net-type neural network. The automatic trajectory generation method is evaluated on six volunteers with various articulated joint angles. In all cases, the system can successfully acquire the planned vascular structure on volunteers' limbs. For one volunteer the MRI scan was also available, which allows the evaluation of the average radius of the scanned artery from US images, resulting in a radius estimation ($1.2\pm0.05~mm$) comparable to the MRI ground truth ($1.2\pm0.04~mm$).
Precise Repositioning of Robotic Ultrasound: Improving Registration-based Motion Compensation using Ultrasound Confidence Optimization
Aug 10, 2022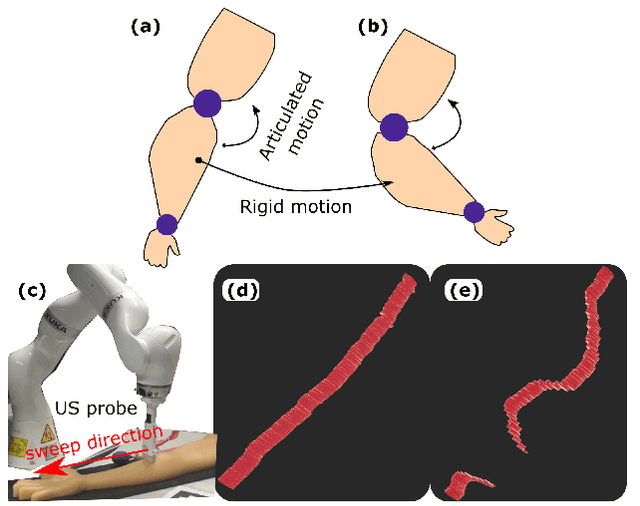
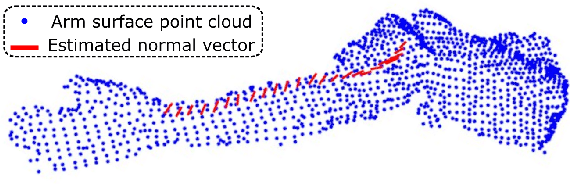
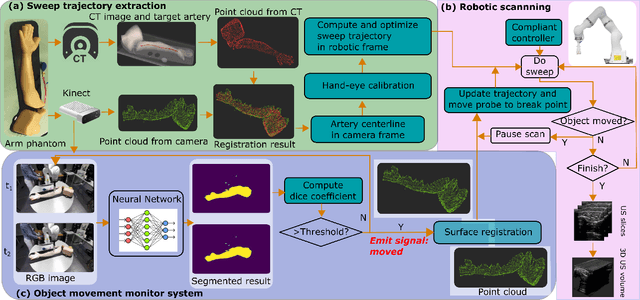
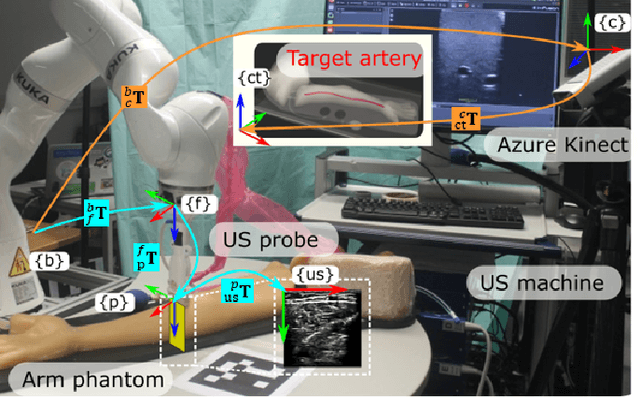

Abstract:Robotic ultrasound (US) imaging has been seen as a promising solution to overcome the limitations of free-hand US examinations, i.e., inter-operator variability. \revision{However, the fact that robotic US systems cannot react to subject movements during scans limits their clinical acceptance.} Regarding human sonographers, they often react to patient movements by repositioning the probe or even restarting the acquisition, in particular for the scans of anatomies with long structures like limb arteries. To realize this characteristic, we proposed a vision-based system to monitor the subject's movement and automatically update the scan trajectory thus seamlessly obtaining a complete 3D image of the target anatomy. The motion monitoring module is developed using the segmented object masks from RGB images. Once the subject is moved, the robot will stop and recompute a suitable trajectory by registering the surface point clouds of the object obtained before and after the movement using the iterative closest point algorithm. Afterward, to ensure optimal contact conditions after repositioning US probe, a confidence-based fine-tuning process is used to avoid potential gaps between the probe and contact surface. Finally, the whole system is validated on a human-like arm phantom with an uneven surface, while the object segmentation network is also validated on volunteers. The results demonstrate that the presented system can react to object movements and reliably provide accurate 3D images.
CloudAttention: Efficient Multi-Scale Attention Scheme For 3D Point Cloud Learning
Jul 31, 2022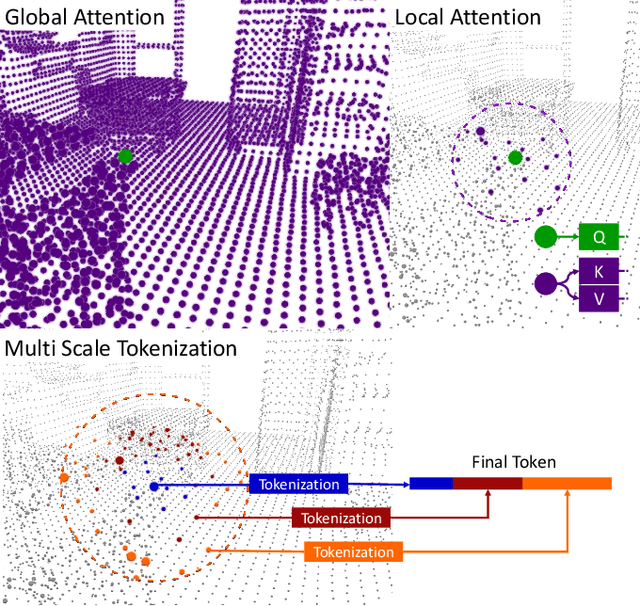
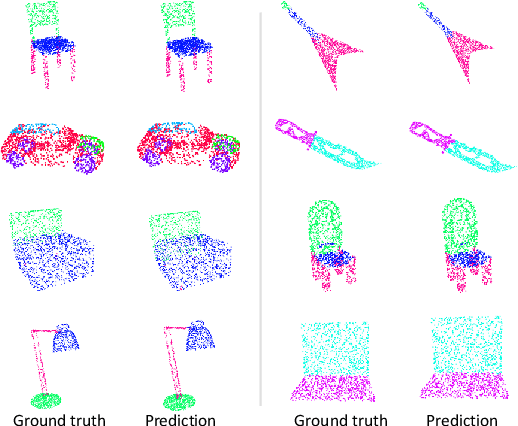
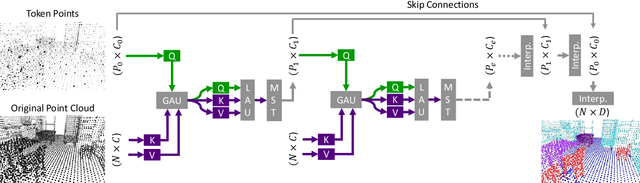
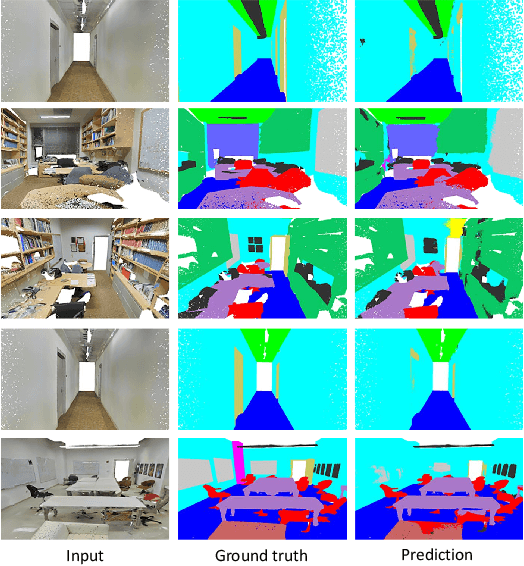

Abstract:Processing 3D data efficiently has always been a challenge. Spatial operations on large-scale point clouds, stored as sparse data, require extra cost. Attracted by the success of transformers, researchers are using multi-head attention for vision tasks. However, attention calculations in transformers come with quadratic complexity in the number of inputs and miss spatial intuition on sets like point clouds. We redesign set transformers in this work and incorporate them into a hierarchical framework for shape classification and part and scene segmentation. We propose our local attention unit, which captures features in a spatial neighborhood. We also compute efficient and dynamic global cross attentions by leveraging sampling and grouping at each iteration. Finally, to mitigate the non-heterogeneity of point clouds, we propose an efficient Multi-Scale Tokenization (MST), which extracts scale-invariant tokens for attention operations. The proposed hierarchical model achieves state-of-the-art shape classification in mean accuracy and yields results on par with the previous segmentation methods while requiring significantly fewer computations. Our proposed architecture predicts segmentation labels with around half the latency and parameter count of the previous most efficient method with comparable performance. The code is available at https://github.com/YigeWang-WHU/CloudAttention.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge