Le Lu
Med-Query: Steerable Parsing of 9-DoF Medical Anatomies with Query Embedding
Dec 05, 2022



Abstract:Automatic parsing of human anatomies at instance-level from 3D computed tomography (CT) scans is a prerequisite step for many clinical applications. The presence of pathologies, broken structures or limited field-of-view (FOV) all can make anatomy parsing algorithms vulnerable. In this work, we explore how to exploit and conduct the prosperous detection-then-segmentation paradigm in 3D medical data, and propose a steerable, robust, and efficient computing framework for detection, identification, and segmentation of anatomies in CT scans. Considering complicated shapes, sizes and orientations of anatomies, without lose of generality, we present the nine degrees-of-freedom (9-DoF) pose estimation solution in full 3D space using a novel single-stage, non-hierarchical forward representation. Our whole framework is executed in a steerable manner where any anatomy of interest can be directly retrieved to further boost the inference efficiency. We have validated the proposed method on three medical imaging parsing tasks of ribs, spine, and abdominal organs. For rib parsing, CT scans have been annotated at the rib instance-level for quantitative evaluation, similarly for spine vertebrae and abdominal organs. Extensive experiments on 9-DoF box detection and rib instance segmentation demonstrate the effectiveness of our framework (with the identification rate of 97.0% and the segmentation Dice score of 90.9%) in high efficiency, compared favorably against several strong baselines (e.g., CenterNet, FCOS, and nnU-Net). For spine identification and segmentation, our method achieves a new state-of-the-art result on the public CTSpine1K dataset. Last, we report highly competitive results in multi-organ segmentation at FLARE22 competition. Our annotations, code and models will be made publicly available at: https://github.com/alibaba-damo-academy/Med_Query.
A New Probabilistic V-Net Model with Hierarchical Spatial Feature Transform for Efficient Abdominal Multi-Organ Segmentation
Aug 02, 2022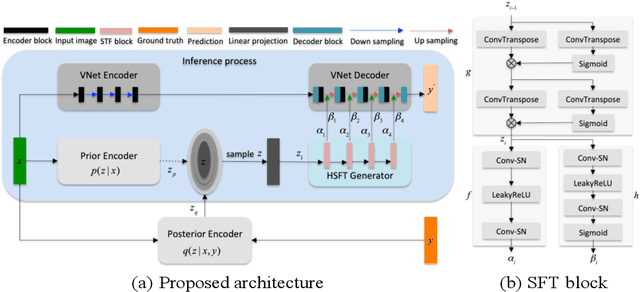
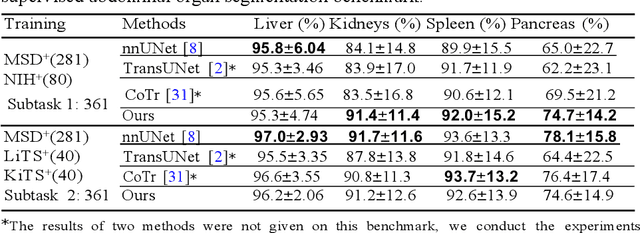
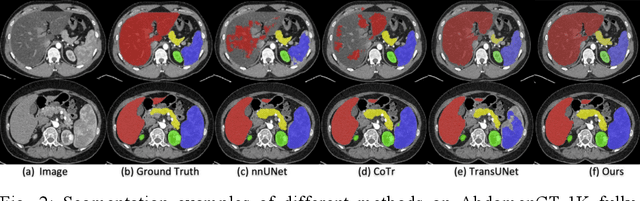
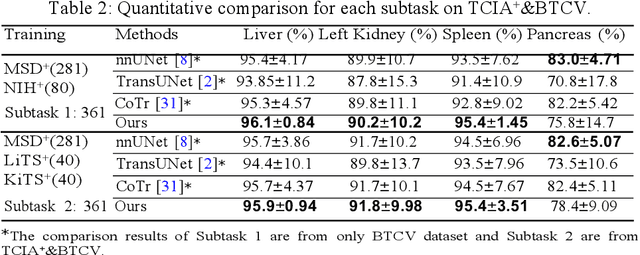
Abstract:Accurate and robust abdominal multi-organ segmentation from CT imaging of different modalities is a challenging task due to complex inter- and intra-organ shape and appearance variations among abdominal organs. In this paper, we propose a probabilistic multi-organ segmentation network with hierarchical spatial-wise feature modulation to capture flexible organ semantic variants and inject the learnt variants into different scales of feature maps for guiding segmentation. More specifically, we design an input decomposition module via a conditional variational auto-encoder to learn organ-specific distributions on the low dimensional latent space and model richer organ semantic variations that is conditioned on input images.Then by integrating these learned variations into the V-Net decoder hierarchically via spatial feature transformation, which has the ability to convert the variations into conditional Affine transformation parameters for spatial-wise feature maps modulating and guiding the fine-scale segmentation. The proposed method is trained on the publicly available AbdomenCT-1K dataset and evaluated on two other open datasets, i.e., 100 challenging/pathological testing patient cases from AbdomenCT-1K fully-supervised abdominal organ segmentation benchmark and 90 cases from TCIA+&BTCV dataset. Highly competitive or superior quantitative segmentation results have been achieved using these datasets for four abdominal organs of liver, kidney, spleen and pancreas with reported Dice scores improved by 7.3% for kidneys and 9.7% for pancreas, while being ~7 times faster than two strong baseline segmentation methods(nnUNet and CoTr).
LViT: Language meets Vision Transformer in Medical Image Segmentation
Jun 29, 2022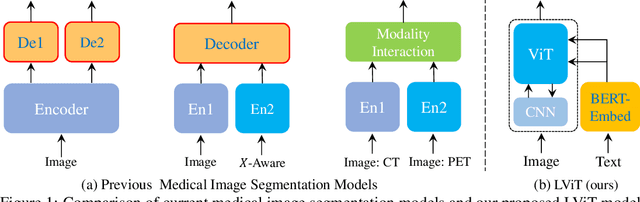
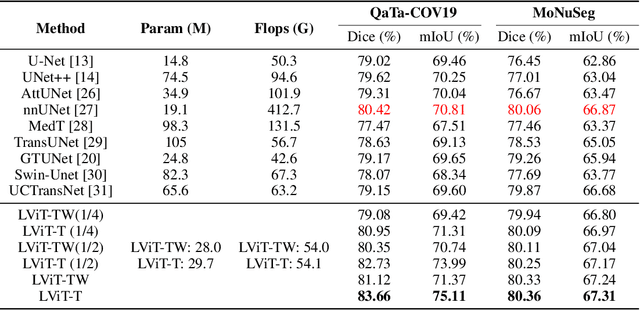
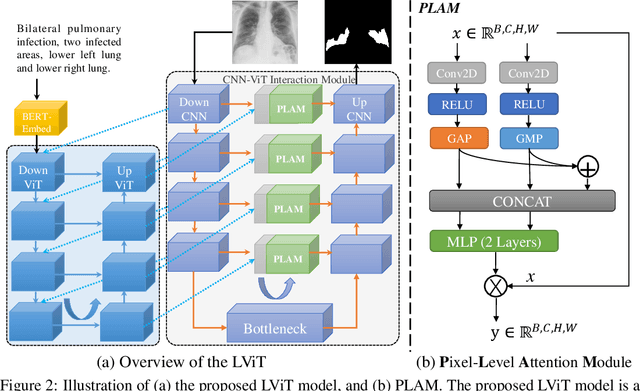
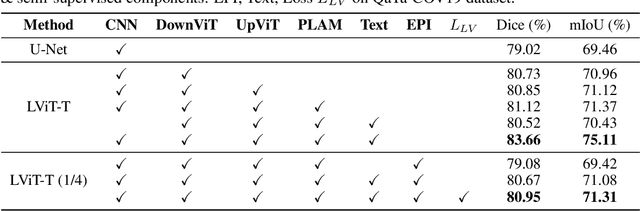
Abstract:Deep learning has been widely used in medical image segmentation and other aspects. However, the performance of existing medical image segmentation models has been limited by the challenge of obtaining sufficient number of high-quality data with the high cost of data annotation. To overcome the limitation, we propose a new vision-language medical image segmentation model LViT (Language meets Vision Transformer). In our model, medical text annotation is introduced to compensate for the quality deficiency in image data. In addition, the text information can guide the generation of pseudo labels to a certain extent and further guarantee the quality of pseudo labels in semi-supervised learning. We also propose the Exponential Pseudo label Iteration mechanism (EPI) to help extend the semi-supervised version of LViT and the Pixel-Level Attention Module (PLAM) to preserve local features of images. In our model, LV (Language-Vision) loss is designed to supervise the training of unlabeled images using text information directly. To validate the performance of LViT, we construct multimodal medical segmentation datasets (image + text) containing pathological images, X-rays,etc. Experimental results show that our proposed LViT has better segmentation performance in both fully and semi-supervised conditions. Code and datasets are available at https://github.com/HUANGLIZI/LViT.
Localized Adversarial Domain Generalization
May 09, 2022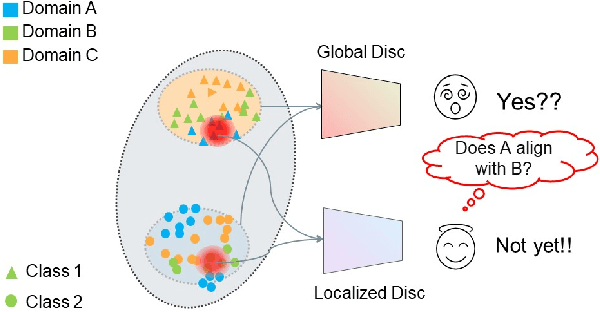
Abstract:Deep learning methods can struggle to handle domain shifts not seen in training data, which can cause them to not generalize well to unseen domains. This has led to research attention on domain generalization (DG), which aims to the model's generalization ability to out-of-distribution. Adversarial domain generalization is a popular approach to DG, but conventional approaches (1) struggle to sufficiently align features so that local neighborhoods are mixed across domains; and (2) can suffer from feature space over collapse which can threaten generalization performance. To address these limitations, we propose localized adversarial domain generalization with space compactness maintenance~(LADG) which constitutes two major contributions. First, we propose an adversarial localized classifier as the domain discriminator, along with a principled primary branch. This constructs a min-max game whereby the aim of the featurizer is to produce locally mixed domains. Second, we propose to use a coding-rate loss to alleviate feature space over collapse. We conduct comprehensive experiments on the Wilds DG benchmark to validate our approach, where LADG outperforms leading competitors on most datasets.
Lumbar Bone Mineral Density Estimation from Chest X-ray Images: Anatomy-aware Attentive Multi-ROI Modeling
Jan 05, 2022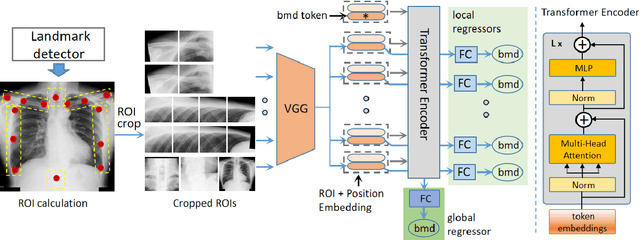
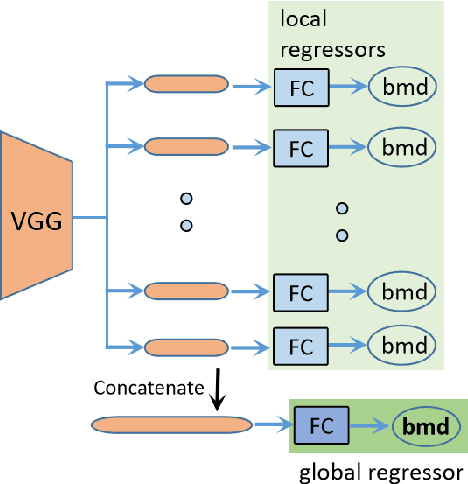
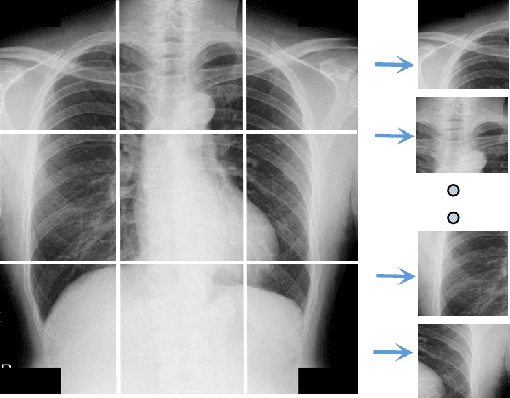
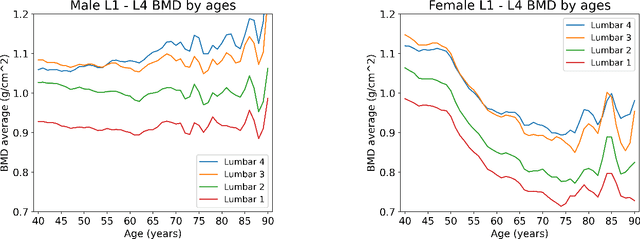
Abstract:Osteoporosis is a common chronic metabolic bone disease that is often under-diagnosed and under-treated due to the limited access to bone mineral density (BMD) examinations, e.g. via Dual-energy X-ray Absorptiometry (DXA). In this paper, we propose a method to predict BMD from Chest X-ray (CXR), one of the most commonly accessible and low-cost medical imaging examinations. Our method first automatically detects Regions of Interest (ROIs) of local and global bone structures from the CXR. Then a multi-ROI deep model with transformer encoder is developed to exploit both local and global information in the chest X-ray image for accurate BMD estimation. Our method is evaluated on 13719 CXR patient cases with their ground truth BMD scores measured by gold-standard DXA. The model predicted BMD has a strong correlation with the ground truth (Pearson correlation coefficient 0.889 on lumbar 1). When applied for osteoporosis screening, it achieves a high classification performance (AUC 0.963 on lumbar 1). As the first effort in the field using CXR scans to predict the BMD, the proposed algorithm holds strong potential in early osteoporosis screening and public health promotion.
Coherence Learning using Keypoint-based Pooling Network for Accurately Assessing Radiographic Knee Osteoarthritis
Dec 16, 2021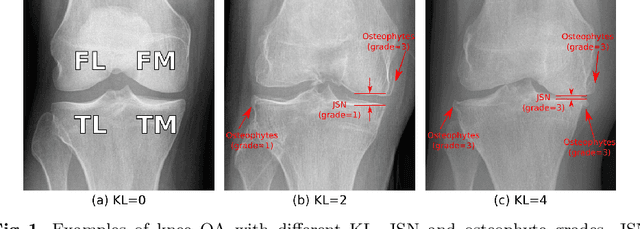
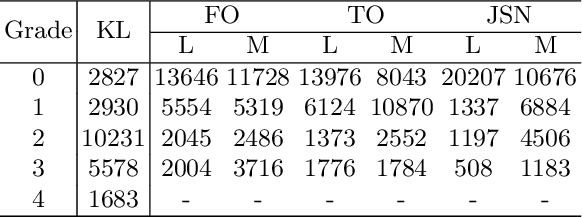
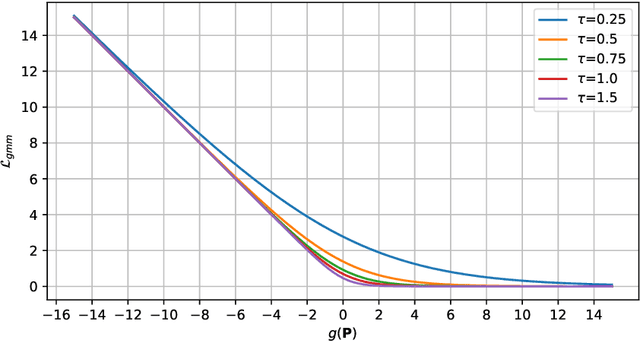
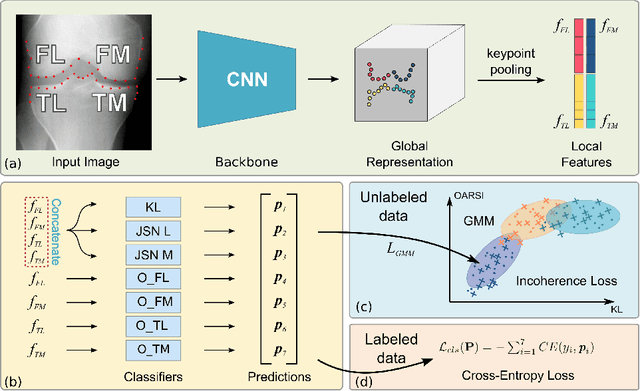
Abstract:Knee osteoarthritis (OA) is a common degenerate joint disorder that affects a large population of elderly people worldwide. Accurate radiographic assessment of knee OA severity plays a critical role in chronic patient management. Current clinically-adopted knee OA grading systems are observer subjective and suffer from inter-rater disagreements. In this work, we propose a computer-aided diagnosis approach to provide more accurate and consistent assessments of both composite and fine-grained OA grades simultaneously. A novel semi-supervised learning method is presented to exploit the underlying coherence in the composite and fine-grained OA grades by learning from unlabeled data. By representing the grade coherence using the log-probability of a pre-trained Gaussian Mixture Model, we formulate an incoherence loss to incorporate unlabeled data in training. The proposed method also describes a keypoint-based pooling network, where deep image features are pooled from the disease-targeted keypoints (extracted along the knee joint) to provide more aligned and pathologically informative feature representations, for accurate OA grade assessments. The proposed method is comprehensively evaluated on the public Osteoarthritis Initiative (OAI) data, a multi-center ten-year observational study on 4,796 subjects. Experimental results demonstrate that our method leads to significant improvements over previous strong whole image-based deep classification network baselines (like ResNet-50).
Comprehensive and Clinically Accurate Head and Neck Organs at Risk Delineation via Stratified Deep Learning: A Large-scale Multi-Institutional Study
Nov 01, 2021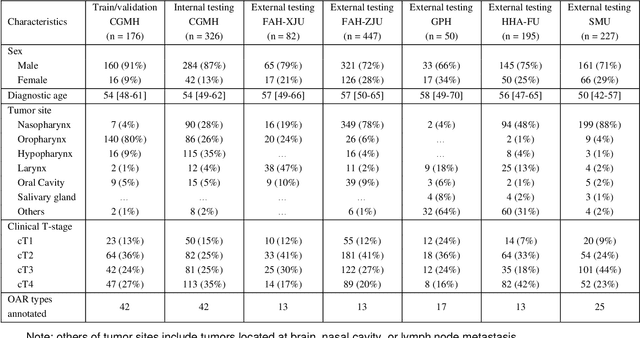
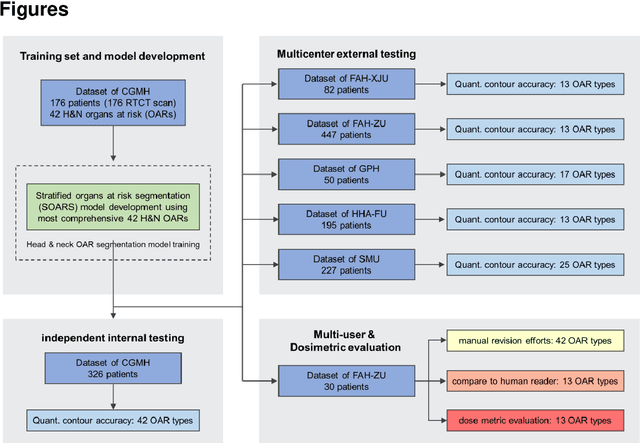
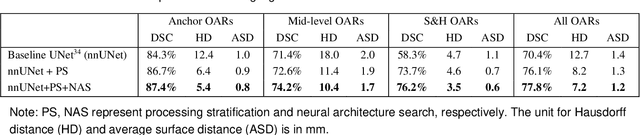
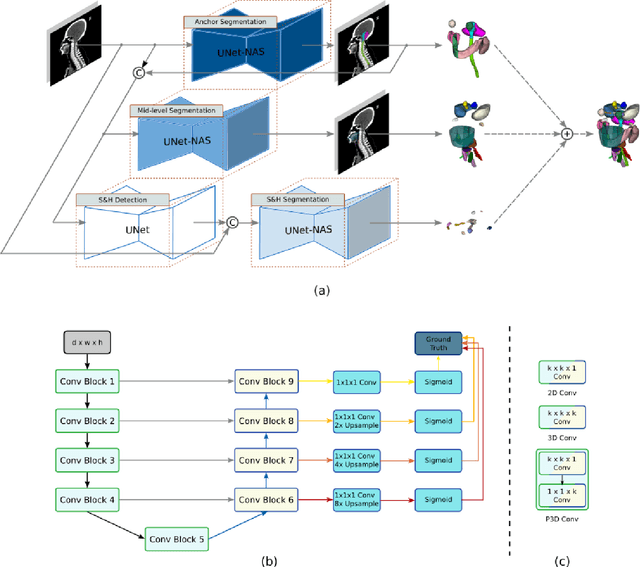
Abstract:Accurate organ at risk (OAR) segmentation is critical to reduce the radiotherapy post-treatment complications. Consensus guidelines recommend a set of more than 40 OARs in the head and neck (H&N) region, however, due to the predictable prohibitive labor-cost of this task, most institutions choose a substantially simplified protocol by delineating a smaller subset of OARs and neglecting the dose distributions associated with other OARs. In this work we propose a novel, automated and highly effective stratified OAR segmentation (SOARS) system using deep learning to precisely delineate a comprehensive set of 42 H&N OARs. SOARS stratifies 42 OARs into anchor, mid-level, and small & hard subcategories, with specifically derived neural network architectures for each category by neural architecture search (NAS) principles. We built SOARS models using 176 training patients in an internal institution and independently evaluated on 1327 external patients across six different institutions. It consistently outperformed other state-of-the-art methods by at least 3-5% in Dice score for each institutional evaluation (up to 36% relative error reduction in other metrics). More importantly, extensive multi-user studies evidently demonstrated that 98% of the SOARS predictions need only very minor or no revisions for direct clinical acceptance (saving 90% radiation oncologists workload), and their segmentation and dosimetric accuracy are within or smaller than the inter-user variation. These findings confirmed the strong clinical applicability of SOARS for the OAR delineation process in H&N cancer radiotherapy workflows, with improved efficiency, comprehensiveness, and quality.
Circle Representation for Medical Object Detection
Oct 22, 2021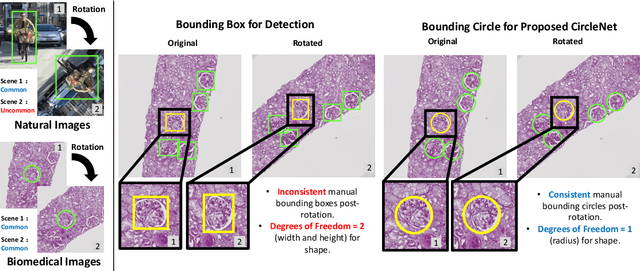
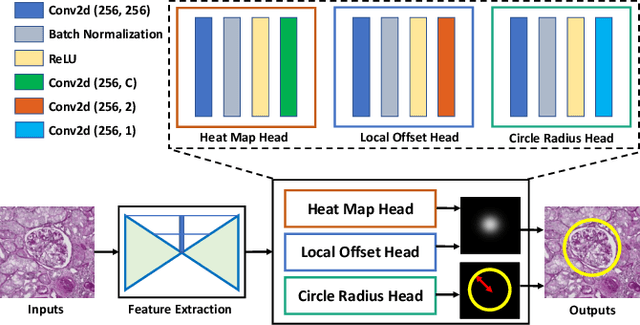
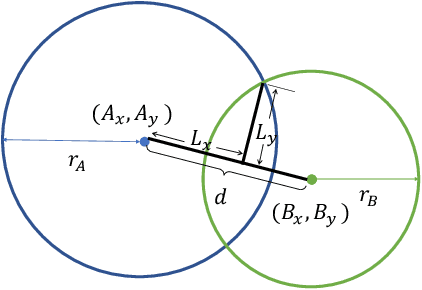
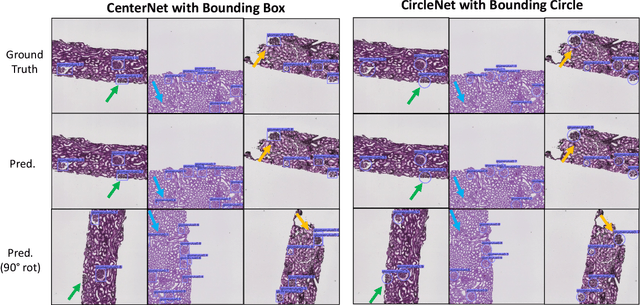
Abstract:Box representation has been extensively used for object detection in computer vision. Such representation is efficacious but not necessarily optimized for biomedical objects (e.g., glomeruli), which play an essential role in renal pathology. In this paper, we propose a simple circle representation for medical object detection and introduce CircleNet, an anchor-free detection framework. Compared with the conventional bounding box representation, the proposed bounding circle representation innovates in three-fold: (1) it is optimized for ball-shaped biomedical objects; (2) The circle representation reduced the degree of freedom compared with box representation; (3) It is naturally more rotation invariant. When detecting glomeruli and nuclei on pathological images, the proposed circle representation achieved superior detection performance and be more rotation-invariant, compared with the bounding box. The code has been made publicly available: https://github.com/hrlblab/CircleNet
A deep learning pipeline for localization, differentiation, and uncertainty estimation of liver lesions using multi-phasic and multi-sequence MRI
Oct 17, 2021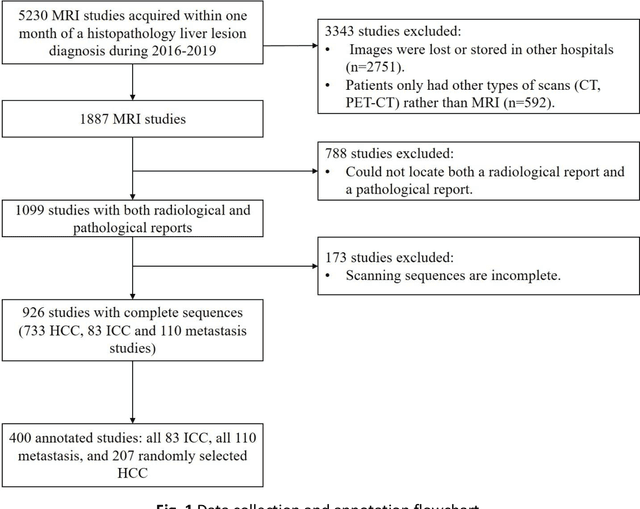
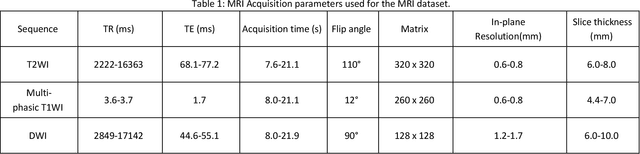
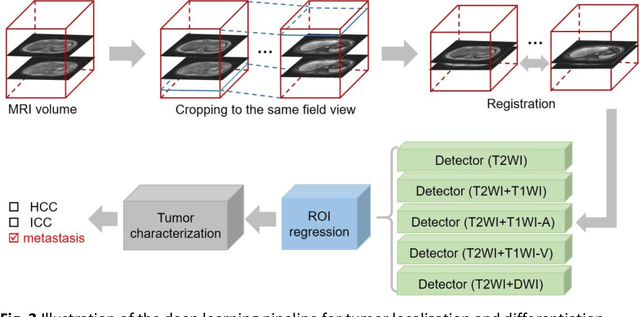
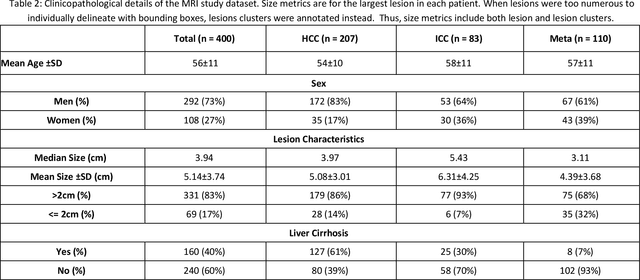
Abstract:Objectives: to propose a fully-automatic computer-aided diagnosis (CAD) solution for liver lesion characterization, with uncertainty estimation. Methods: we enrolled 400 patients who had either liver resection or a biopsy and was diagnosed with either hepatocellular carcinoma (HCC), intrahepatic cholangiocarcinoma, or secondary metastasis, from 2006 to 2019. Each patient was scanned with T1WI, T2WI, T1WI venous phase (T2WI-V), T1WI arterial phase (T1WI-A), and DWI MRI sequences. We propose a fully-automatic deep CAD pipeline that localizes lesions from 3D MRI studies using key-slice parsing and provides a confidence measure for its diagnoses. We evaluate using five-fold cross validation and compare performance against three radiologists, including a senior hepatology radiologist, a junior hepatology radiologist and an abdominal radiologist. Results: the proposed CAD solution achieves a mean F1 score of 0.62, outperforming the abdominal radiologist (0.47), matching the junior hepatology radiologist (0.61), and underperforming the senior hepatology radiologist (0.68). The CAD system can informatively assess its diagnostic confidence, i.e., when only evaluating on the 70% most confident cases the mean f1 score and sensitivity at 80% specificity for HCC vs. others are boosted from 0.62 to 0.71 and 0.84 to 0.92, respectively. Conclusion: the proposed fully-automatic CAD solution can provide good diagnostic performance with informative confidence assessments in finding and discriminating liver lesions from MRI studies.
Accurate and Generalizable Quantitative Scoring of Liver Steatosis from Ultrasound Images via Scalable Deep Learning
Oct 12, 2021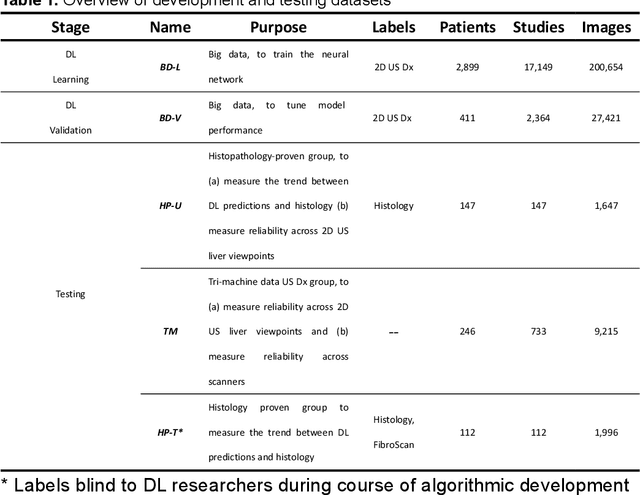
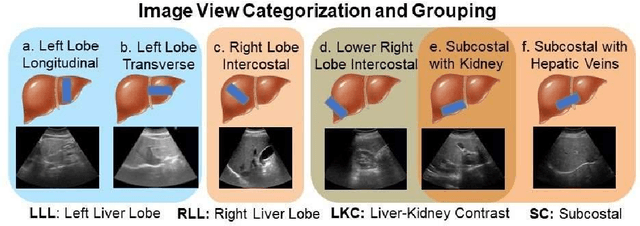
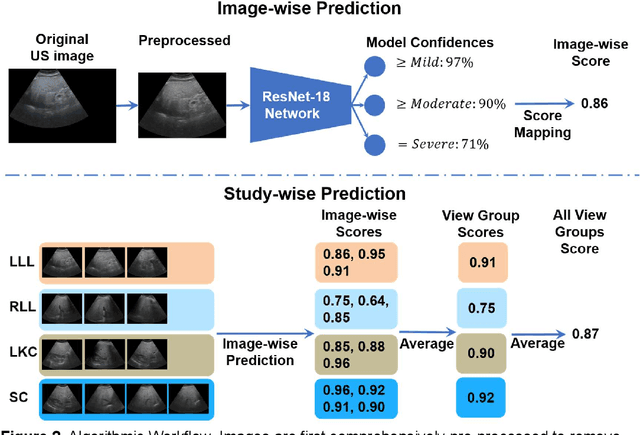
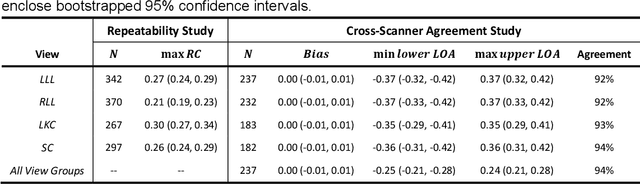
Abstract:Background & Aims: Hepatic steatosis is a major cause of chronic liver disease. 2D ultrasound is the most widely used non-invasive tool for screening and monitoring, but associated diagnoses are highly subjective. We developed a scalable deep learning (DL) algorithm for quantitative scoring of liver steatosis from 2D ultrasound images. Approach & Results: Using retrospectively collected multi-view ultrasound data from 3,310 patients, 19,513 studies, and 228,075 images, we trained a DL algorithm to diagnose steatosis stages (healthy, mild, moderate, or severe) from ultrasound diagnoses. Performance was validated on two multi-scanner unblinded and blinded (initially to DL developer) histology-proven cohorts (147 and 112 patients) with histopathology fatty cell percentage diagnoses, and a subset with FibroScan diagnoses. We also quantified reliability across scanners and viewpoints. Results were evaluated using Bland-Altman and receiver operating characteristic (ROC) analysis. The DL algorithm demonstrates repeatable measurements with a moderate number of images (3 for each viewpoint) and high agreement across 3 premium ultrasound scanners. High diagnostic performance was observed across all viewpoints: area under the curves of the ROC to classify >=mild, >=moderate, =severe steatosis grades were 0.85, 0.90, and 0.93, respectively. The DL algorithm outperformed or performed at least comparably to FibroScan with statistically significant improvements for all levels on the unblinded histology-proven cohort, and for =severe steatosis on the blinded histology-proven cohort. Conclusions: The DL algorithm provides a reliable quantitative steatosis assessment across view and scanners on two multi-scanner cohorts. Diagnostic performance was high with comparable or better performance than FibroScan.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge