Jianan Chen
Teaching pathology foundation models to accurately predict gene expression with parameter efficient knowledge transfer
Apr 09, 2025



Abstract:Gene expression profiling provides critical insights into cellular heterogeneity, biological processes and disease mechanisms. There has been an increasing interest in computational approaches that can predict gene expression directly from digitalized histopathology images. While image foundation models have shown promise in a variety of pathology downstream analysis, their performances on gene-expression prediction are still limited. Explicitly incorporating information from the transcriptomic models can help image models to address domain shift, yet the fine-tuning and alignment of foundation models can be expensive. In the work, we propose Parameter Efficient Knowledge trAnsfer (PEKA), a novel framework that leverages Block-Affine Adaptation and integrates knowledge distillation and structure alignment losses for cross-modal knowledge transfer. We evaluated PEKA for gene expression prediction using multiple spatial transcriptomics datasets (comprising 206,123 image tiles with matched gene expression profiles) that encompassed various types of tissue. PEKA achieved at least 5\% performance improvement over baseline foundation models while also outperforming alternative parameter-efficient fine-tuning strategies. We will release the code, datasets and aligned models after peer-review to facilitate broader adoption and further development for parameter efficient model alignment.
Upcycling Noise for Federated Unlearning
Dec 07, 2024



Abstract:In Federated Learning (FL), multiple clients collaboratively train a model without sharing raw data. This paradigm can be further enhanced by Differential Privacy (DP) to protect local data from information inference attacks and is thus termed DPFL. An emerging privacy requirement, ``the right to be forgotten'' for clients, poses new challenges to DPFL but remains largely unexplored. Despite numerous studies on federated unlearning (FU), they are inapplicable to DPFL because the noise introduced by the DP mechanism compromises their effectiveness and efficiency. In this paper, we propose Federated Unlearning with Indistinguishability (FUI) to unlearn the local data of a target client in DPFL for the first time. FUI consists of two main steps: local model retraction and global noise calibration, resulting in an unlearning model that is statistically indistinguishable from the retrained model. Specifically, we demonstrate that the noise added in DPFL can endow the unlearning model with a certain level of indistinguishability after local model retraction, and then fortify the degree of unlearning through global noise calibration. Additionally, for the efficient and consistent implementation of the proposed FUI, we formulate a two-stage Stackelberg game to derive optimal unlearning strategies for both the server and the target client. Privacy and convergence analyses confirm theoretical guarantees, while experimental results based on four real-world datasets illustrate that our proposed FUI achieves superior model performance and higher efficiency compared to mainstream FU schemes. Simulation results further verify the optimality of the derived unlearning strategies.
A Unified Platform for At-Home Post-Stroke Rehabilitation Enabled by Wearable Technologies and Artificial Intelligence
Nov 28, 2024



Abstract:At-home rehabilitation for post-stroke patients presents significant challenges, as continuous, personalized care is often limited outside clinical settings. Additionally, the absence of comprehensive solutions addressing diverse rehabilitation needs in home environments complicates recovery efforts. Here, we introduce a smart home platform that integrates wearable sensors, ambient monitoring, and large language model (LLM)-powered assistance to provide seamless health monitoring and intelligent support. The system leverages machine learning enabled plantar pressure arrays for motor recovery assessment (94% classification accuracy), a wearable eye-tracking module for cognitive evaluation, and ambient sensors for precise smart home control (100% operational success, <1 s latency). Additionally, the LLM-powered agent, Auto-Care, offers real-time interventions, such as health reminders and environmental adjustments, enhancing user satisfaction by 29%. This work establishes a fully integrated platform for long-term, personalized rehabilitation, offering new possibilities for managing chronic conditions and supporting aging populations.
Airport Delay Prediction with Temporal Fusion Transformers
May 14, 2024Abstract:Since flight delay hurts passengers, airlines, and airports, its prediction becomes crucial for the decision-making of all stakeholders in the aviation industry and thus has been attempted by various previous research. However, previous delay predictions are often categorical and at a highly aggregated level. To improve that, this study proposes to apply the novel Temporal Fusion Transformer model and predict numerical airport arrival delays at quarter hour level for U.S. top 30 airports. Inputs to our model include airport demand and capacity forecasts, historic airport operation efficiency information, airport wind and visibility conditions, as well as enroute weather and traffic conditions. The results show that our model achieves satisfactory performance measured by small prediction errors on the test set. In addition, the interpretability analysis of the model outputs identifies the important input factors for delay prediction.
Cross-Validation Is All You Need: A Statistical Approach To Label Noise Estimation
Jun 24, 2023Abstract:Label noise is prevalent in machine learning datasets. It is crucial to identify and remove label noise because models trained on noisy data can have substantially reduced accuracy and generalizability. Most existing label noise detection approaches are designed for classification tasks, and data cleaning for outcome prediction analysis is relatively unexplored. Inspired by the fluctuations in performance across different folds in cross-validation, we propose Repeated Cross-Validations for label noise estimation (ReCoV) to address this gap. ReCoV constructs a noise histogram that ranks the noise level of samples based on a large number of cross-validations by recording sample IDs in each worst-performing fold. We further propose three approaches for identifying noisy samples based on noise histograms to address increasingly complex noise distributions. We show that ReCoV outperforms state-of-the-art algorithms for label cleaning in a classification task benchmark. More importantly, we show that removing ReCoV-identified noisy samples in two medical imaging outcome prediction datasets significantly improves model performance on test sets. As a statistical approach that does not rely on hyperparameters, noise distributions, or model structures, ReCoV is compatible with any machine learning analysis.
Intra-Modal Constraint Loss For Image-Text Retrieval
Jul 13, 2022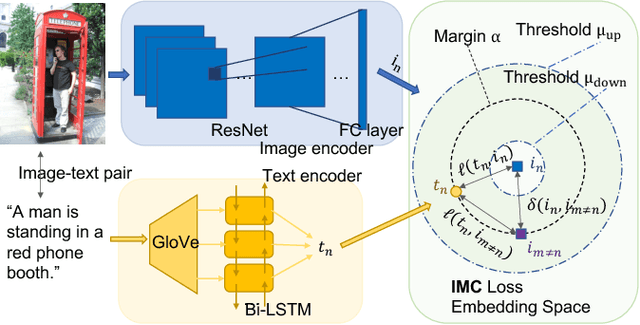
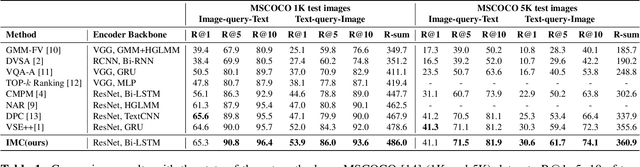
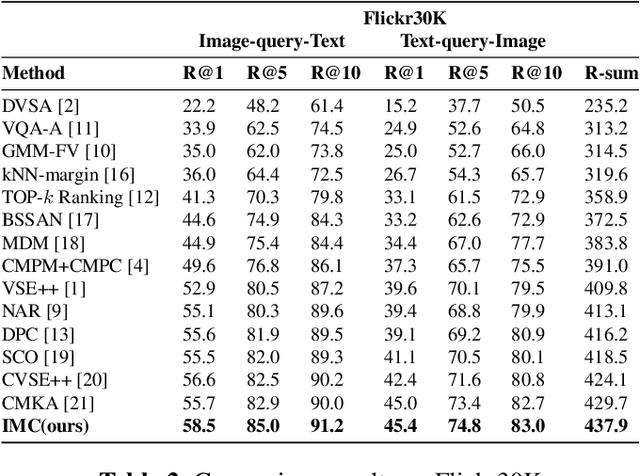
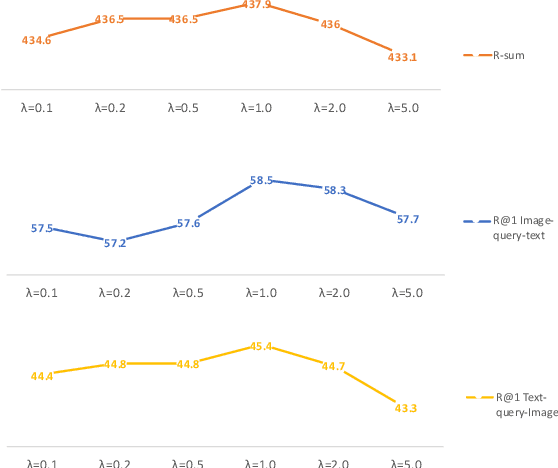
Abstract:Cross-modal retrieval has drawn much attention in both computer vision and natural language processing domains. With the development of convolutional and recurrent neural networks, the bottleneck of retrieval across image-text modalities is no longer the extraction of image and text features but an efficient loss function learning in embedding space. Many loss functions try to closer pairwise features from heterogeneous modalities. This paper proposes a method for learning joint embedding of images and texts using an intra-modal constraint loss function to reduce the violation of negative pairs from the same homogeneous modality. Experimental results show that our approach outperforms state-of-the-art bi-directional image-text retrieval methods on Flickr30K and Microsoft COCO datasets. Our code is publicly available: https://github.com/CanonChen/IMC.
Metastatic Cancer Outcome Prediction with Injective Multiple Instance Pooling
Mar 09, 2022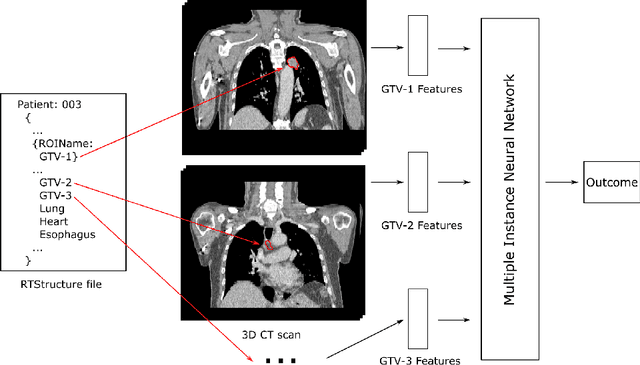
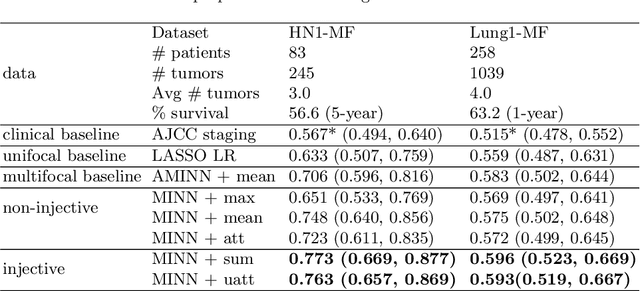
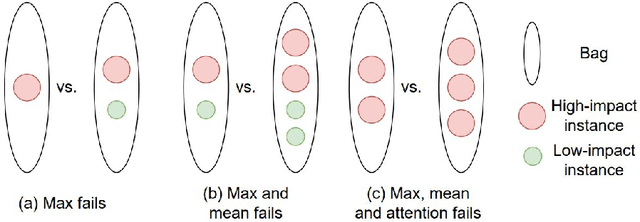
Abstract:Cancer stage is a large determinant of patient prognosis and management in many cancer types, and is often assessed using medical imaging modalities, such as CT and MRI. These medical images contain rich information that can be explored to stratify patients within each stage group to further improve prognostic algorithms. Although the majority of cancer deaths result from metastatic and multifocal disease, building imaging biomarkers for patients with multiple tumors has been a challenging task due to the lack of annotated datasets and standard study framework. In this paper, we process two public datasets to set up a benchmark cohort of 341 patient in total for studying outcome prediction of multifocal metastatic cancer. We identify the lack of expressiveness in common multiple instance classification networks and propose two injective multiple instance pooling functions that are better suited to outcome prediction. Our results show that multiple instance learning with injective pooling functions can achieve state-of-the-art performance in the non-small-cell lung cancer CT and head and neck CT outcome prediction benchmarking tasks. We will release the processed multifocal datasets, our code and the intermediate files i.e. extracted radiomic features to support further transparent and reproducible research.
The MICCAI Hackathon on reproducibility, diversity, and selection of papers at the MICCAI conference
Mar 04, 2021
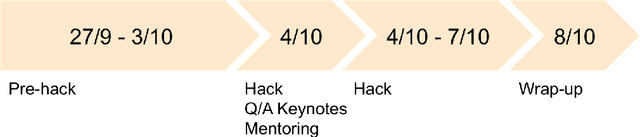
Abstract:The MICCAI conference has encountered tremendous growth over the last years in terms of the size of the community, as well as the number of contributions and their technical success. With this growth, however, come new challenges for the community. Methods are more difficult to reproduce and the ever-increasing number of paper submissions to the MICCAI conference poses new questions regarding the selection process and the diversity of topics. To exchange, discuss, and find novel and creative solutions to these challenges, a new format of a hackathon was initiated as a satellite event at the MICCAI 2020 conference: The MICCAI Hackathon. The first edition of the MICCAI Hackathon covered the topics reproducibility, diversity, and selection of MICCAI papers. In the manner of a small think-tank, participants collaborated to find solutions to these challenges. In this report, we summarize the insights from the MICCAI Hackathon into immediate and long-term measures to address these challenges. The proposed measures can be seen as starting points and guidelines for discussions and actions to possibly improve the MICCAI conference with regards to reproducibility, diversity, and selection of papers.
AMINN: Autoencoder-based Multiple Instance Neural Network for Outcome Prediction of Multifocal Liver Metastases
Dec 12, 2020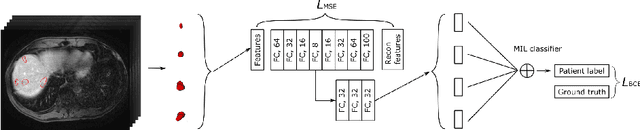
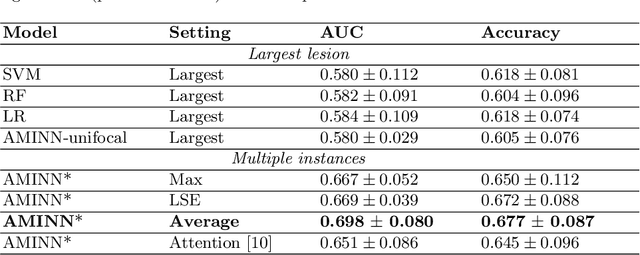
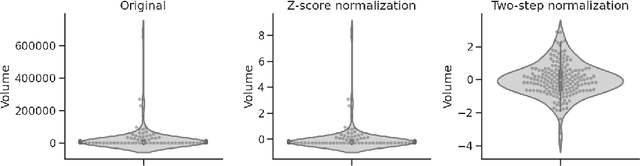
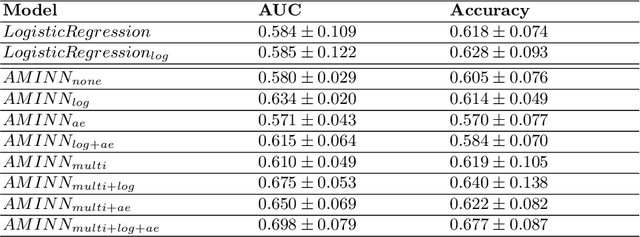
Abstract:Colorectal cancer is one of the most common and lethal cancers and colorectal cancer liver metastases (CRLM) is the major cause of death in patients with colorectal cancer. Multifocality occurs frequently in CRLM, but is relatively unexplored in CRLM outcome prediction. Most existing clinical and imaging biomarkers do not take the imaging features of all multifocal lesions into account. In this paper, we present an end-to-end autoencoder-based multiple instance neural network (AMINN) for the prediction of survival outcomes in multifocal CRLM patients using radiomic features extracted from contrast-enhanced MRIs. Specifically, we jointly train an autoencoder to reconstruct input features and a multiple instance network to make predictions by aggregating information from all tumour lesions of a patient. In addition, we incorporate a two-step normalization technique to improve the training of deep neural networks, built on the observation that the distributions of radiomic features are almost always severely skewed. Experimental results empirically validated our hypothesis that incorporating imaging features of all lesions improves outcome prediction for multifocal cancer. The proposed ADMINN framework achieved an area under the ROC curve (AUC) of 0.70, which is 19.5% higher than baseline methods. We built a risk score based on the outputs of our network and compared it to other clinical and imaging biomarkers. Our risk score is the only one that achieved statistical significance in univariate and multivariate cox proportional hazard modeling in our cohort of multifocal CRLM patients. The effectiveness of incorporating all lesions and applying two-step normalization is demonstrated by a series of ablation studies. Our code will be released after the peer-review process.
AIBench: An Industry Standard AI Benchmark Suite from Internet Services
Apr 30, 2020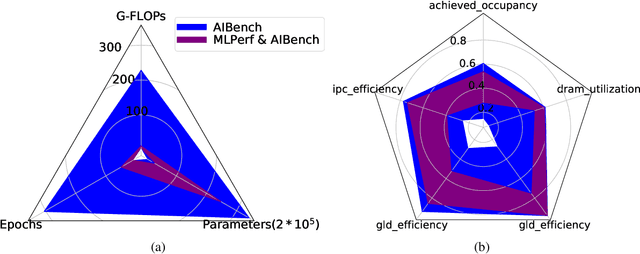
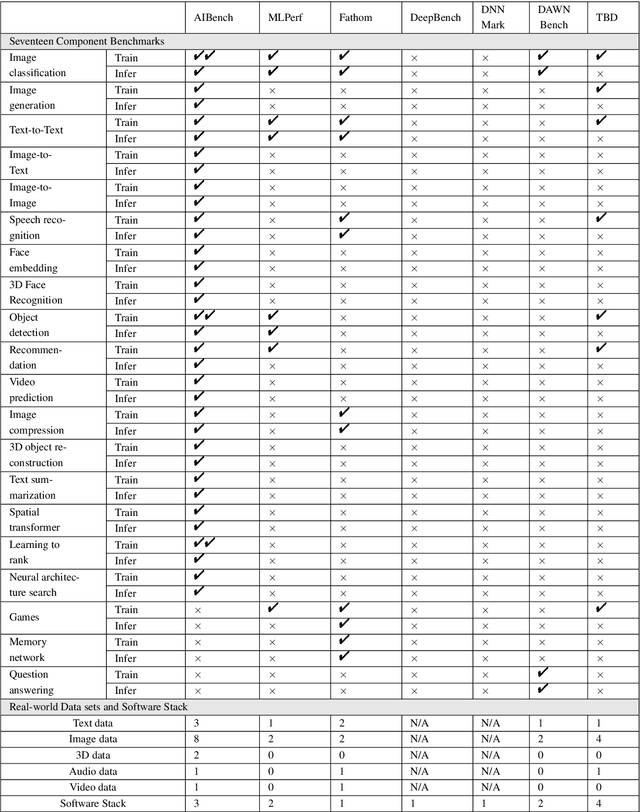
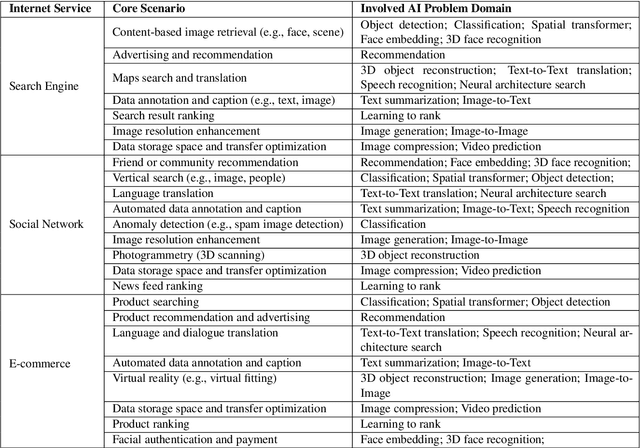
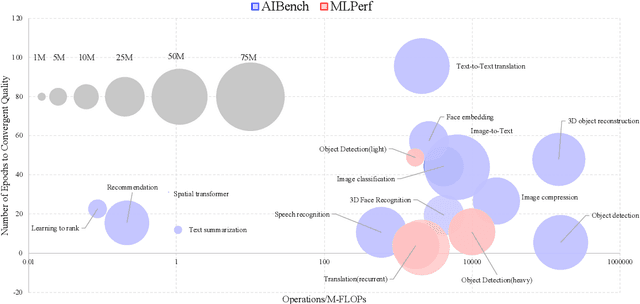
Abstract:The booming successes of machine learning in different domains boost industry-scale deployments of innovative AI algorithms, systems, and architectures, and thus the importance of benchmarking grows. However, the confidential nature of the workloads, the paramount importance of the representativeness and diversity of benchmarks, and the prohibitive cost of training a state-of-the-art model mutually aggravate the AI benchmarking challenges. In this paper, we present a balanced AI benchmarking methodology for meeting the subtly different requirements of different stages in developing a new system/architecture and ranking/purchasing commercial off-the-shelf ones. Performing an exhaustive survey on the most important AI domain-Internet services with seventeen industry partners, we identify and include seventeen representative AI tasks to guarantee the representativeness and diversity of the benchmarks. Meanwhile, for reducing the benchmarking cost, we select a benchmark subset to a minimum-three tasks-according to the criteria: diversity of model complexity, computational cost, and convergence rate, repeatability, and having widely-accepted metrics or not. We contribute by far the most comprehensive AI benchmark suite-AIBench. The evaluations show AIBench outperforms MLPerf in terms of the diversity and representativeness of model complexity, computational cost, convergent rate, computation and memory access patterns, and hotspot functions. With respect to the AIBench full benchmarks, its subset shortens the benchmarking cost by 41%, while maintaining the primary workload characteristics. The specifications, source code, and performance numbers are publicly available from the web site http://www.benchcouncil.org/AIBench/index.html.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge