Unsupervised Image Registration Towards Enhancing Performance and Explainability in Cardiac And Brain Image Analysis
Mar 07, 2022Chengjia Wang, Guang Yang, Giorgos Papanastasiou
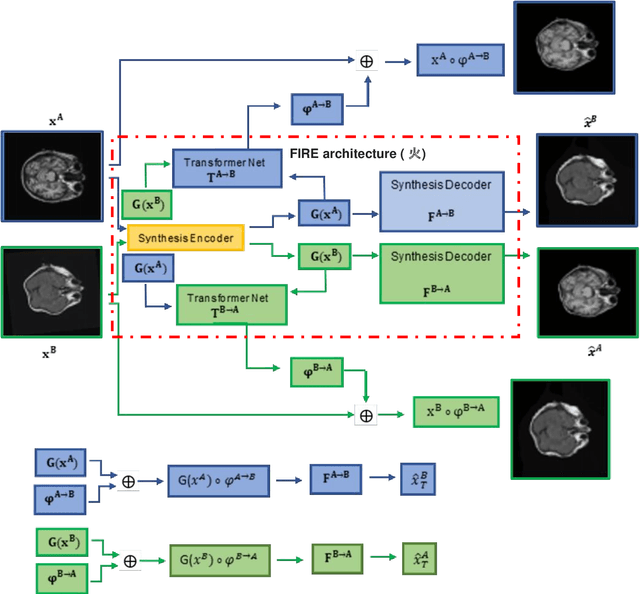
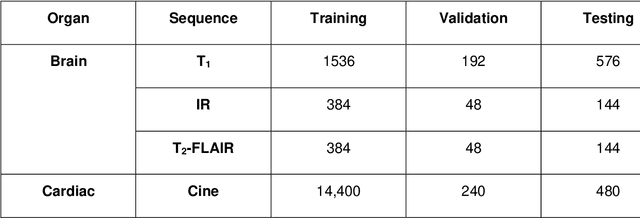
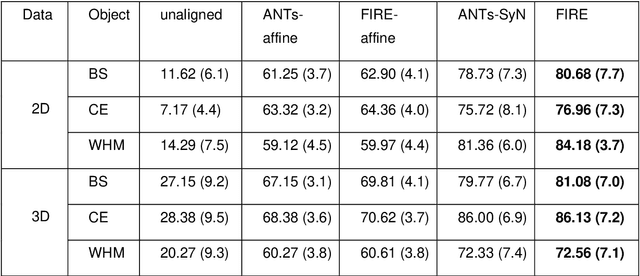
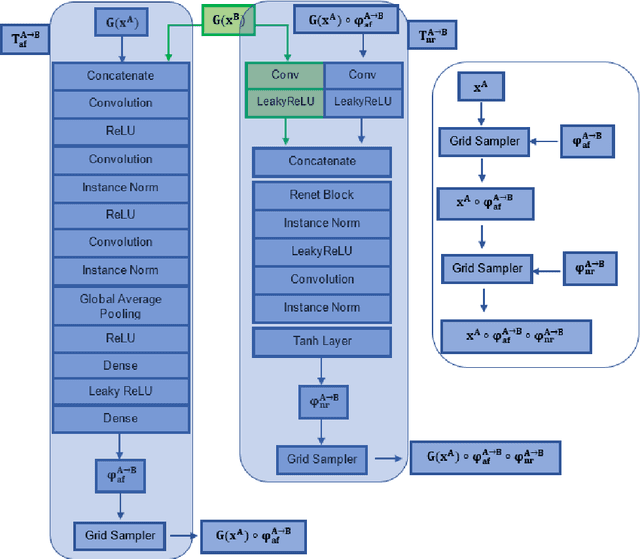
Magnetic Resonance Imaging (MRI) typically recruits multiple sequences (defined here as "modalities"). As each modality is designed to offer different anatomical and functional clinical information, there are evident disparities in the imaging content across modalities. Inter- and intra-modality affine and non-rigid image registration is an essential medical image analysis process in clinical imaging, as for example before imaging biomarkers need to be derived and clinically evaluated across different MRI modalities, time phases and slices. Although commonly needed in real clinical scenarios, affine and non-rigid image registration is not extensively investigated using a single unsupervised model architecture. In our work, we present an un-supervised deep learning registration methodology which can accurately model affine and non-rigid trans-formations, simultaneously. Moreover, inverse-consistency is a fundamental inter-modality registration property that is not considered in deep learning registration algorithms. To address inverse-consistency, our methodology performs bi-directional cross-modality image synthesis to learn modality-invariant latent rep-resentations, while involves two factorised transformation networks and an inverse-consistency loss to learn topology-preserving anatomical transformations. Overall, our model (named "FIRE") shows improved performances against the reference standard baseline method on multi-modality brain 2D and 3D MRI and intra-modality cardiac 4D MRI data experiments.
Explainable COVID-19 Infections Identification and Delineation Using Calibrated Pseudo Labels
Feb 11, 2022Ming Li, Yingying Fang, Zeyu Tang, Chibudom Onuorah, Jun Xia, Javier Del Ser, Simon Walsh, Guang Yang
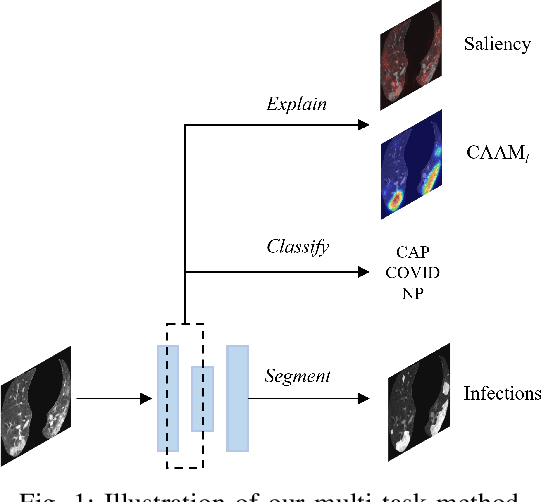
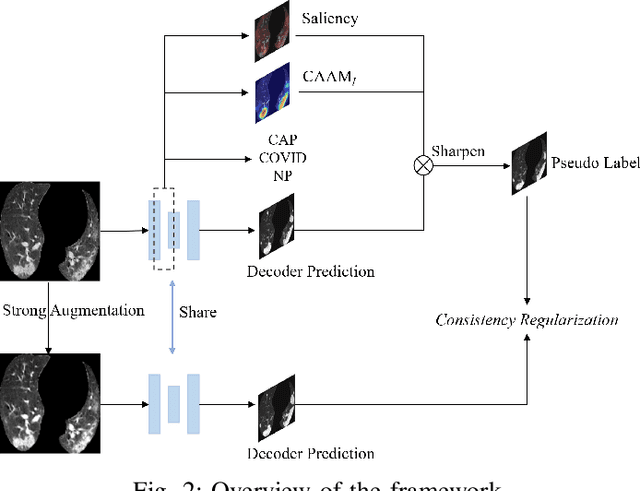
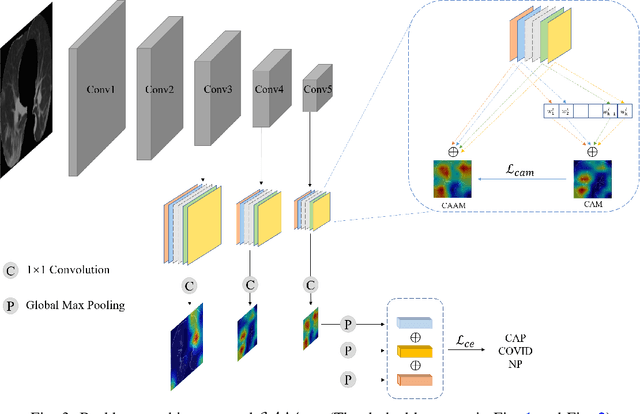
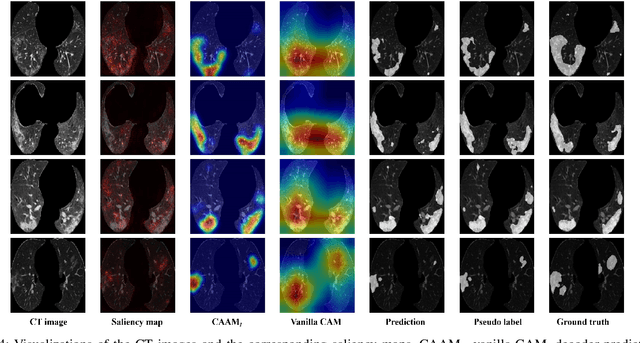
The upheaval brought by the arrival of the COVID-19 pandemic has continued to bring fresh challenges over the past two years. During this COVID-19 pandemic, there has been a need for rapid identification of infected patients and specific delineation of infection areas in computed tomography (CT) images. Although deep supervised learning methods have been established quickly, the scarcity of both image-level and pixellevel labels as well as the lack of explainable transparency still hinder the applicability of AI. Can we identify infected patients and delineate the infections with extreme minimal supervision? Semi-supervised learning (SSL) has demonstrated promising performance under limited labelled data and sufficient unlabelled data. Inspired by SSL, we propose a model-agnostic calibrated pseudo-labelling strategy and apply it under a consistency regularization framework to generate explainable identification and delineation results. We demonstrate the effectiveness of our model with the combination of limited labelled data and sufficient unlabelled data or weakly-labelled data. Extensive experiments have shown that our model can efficiently utilize limited labelled data and provide explainable classification and segmentation results for decision-making in clinical routine.
DocBed: A Multi-Stage OCR Solution for Documents with Complex Layouts
Feb 03, 2022Wenzhen Zhu, Negin Sokhandan, Guang Yang, Sujitha Martin, Suchitra Sathyanarayana
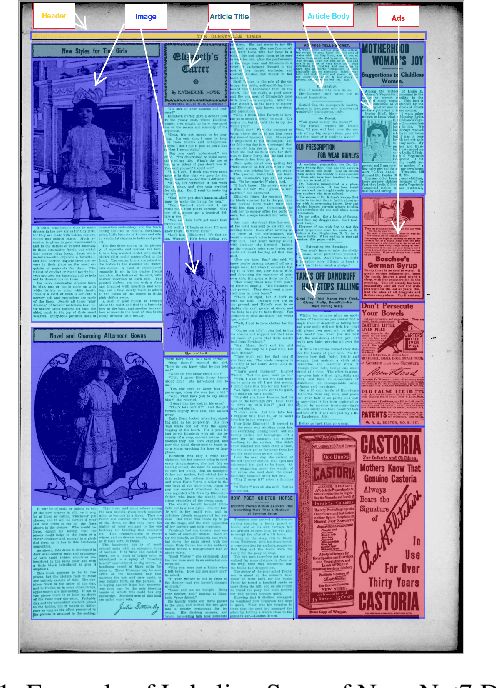

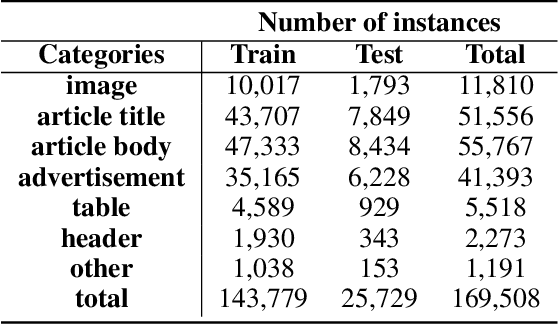
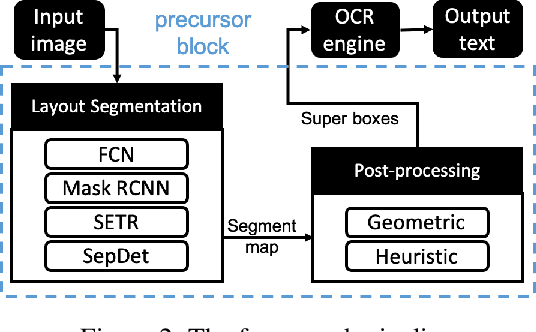
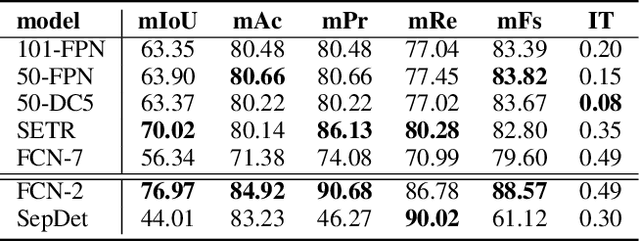
Digitization of newspapers is of interest for many reasons including preservation of history, accessibility and search ability, etc. While digitization of documents such as scientific articles and magazines is prevalent in literature, one of the main challenges for digitization of newspaper lies in its complex layout (e.g. articles spanning multiple columns, text interrupted by images) analysis, which is necessary to preserve human read-order. This work provides a major breakthrough in the digitization of newspapers on three fronts: first, releasing a dataset of 3000 fully-annotated, real-world newspaper images from 21 different U.S. states representing an extensive variety of complex layouts for document layout analysis; second, proposing layout segmentation as a precursor to existing optical character recognition (OCR) engines, where multiple state-of-the-art image segmentation models and several post-processing methods are explored for document layout segmentation; third, providing a thorough and structured evaluation protocol for isolated layout segmentation and end-to-end OCR.
AI-based Medical e-Diagnosis for Fast and Automatic Ventricular Volume Measurement in the Patients with Normal Pressure Hydrocephalus
Jan 31, 2022Xi Zhou, Qinghao Ye, Xiaolin Yang, Jiakuan Chen, Haiqin Ma, Jun Xia, Javier Del Ser, Guang Yang
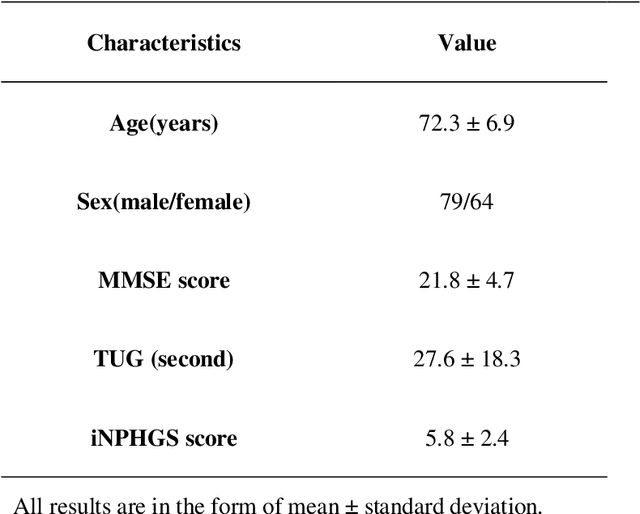

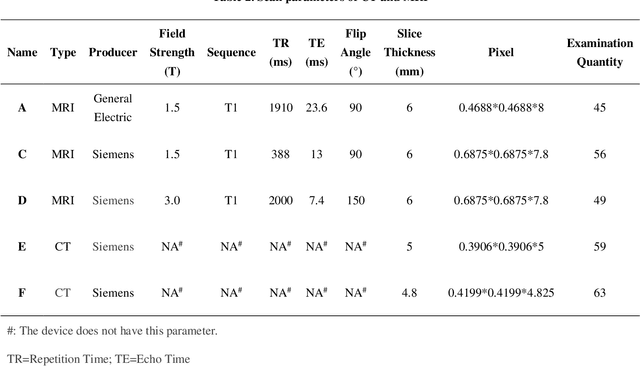
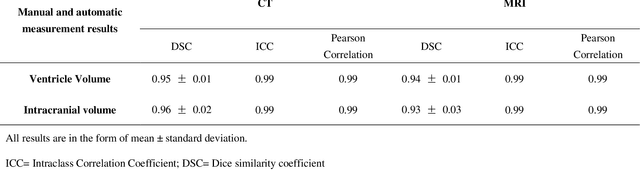
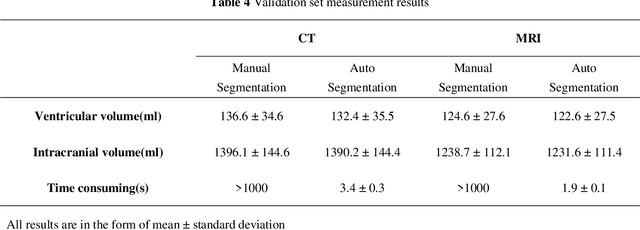
Based on CT and MRI images acquired from normal pressure hydrocephalus (NPH) patients, using machine learning methods, we aim to establish a multi-modal and high-performance automatic ventricle segmentation method to achieve efficient and accurate automatic measurement of the ventricular volume. First, we extract the brain CT and MRI images of 143 definite NPH patients. Second, we manually label the ventricular volume (VV) and intracranial volume (ICV). Then, we use machine learning method to extract features and establish automatic ventricle segmentation model. Finally, we verify the reliability of the model and achieved automatic measurement of VV and ICV. In CT images, the Dice similarity coefficient (DSC), Intraclass Correlation Coefficient (ICC), Pearson correlation, and Bland-Altman analysis of the automatic and manual segmentation result of the VV were 0.95, 0.99, 0.99, and 4.2$\pm$2.6 respectively. The results of ICV were 0.96, 0.99, 0.99, and 6.0$\pm$3.8 respectively. The whole process takes 3.4$\pm$0.3 seconds. In MRI images, the DSC, ICC, Pearson correlation, and Bland-Altman analysis of the automatic and manual segmentation result of the VV were 0.94, 0.99, 0.99, and 2.0$\pm$0.6 respectively. The results of ICV were 0.93, 0.99, 0.99, and 7.9$\pm$3.8 respectively. The whole process took 1.9$\pm$0.1 seconds. We have established a multi-modal and high-performance automatic ventricle segmentation method to achieve efficient and accurate automatic measurement of the ventricular volume of NPH patients. This can help clinicians quickly and accurately understand the situation of NPH patient's ventricles.
Fast MRI Reconstruction: How Powerful Transformers Are?
Jan 23, 2022Jiahao Huang, Yinzhe Wu, Huanjun Wu, Guang Yang
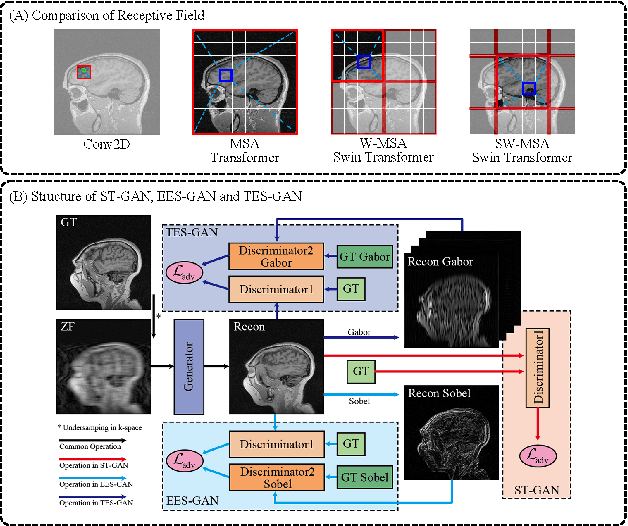
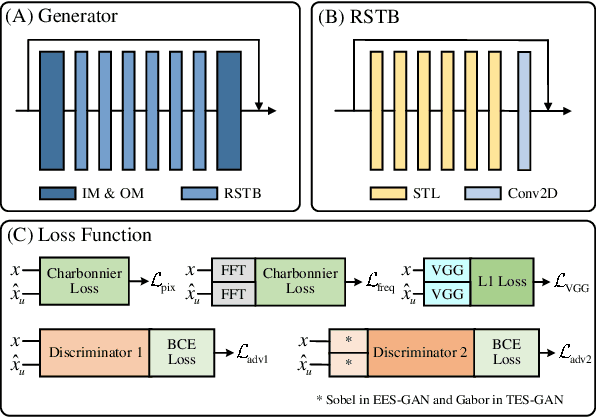
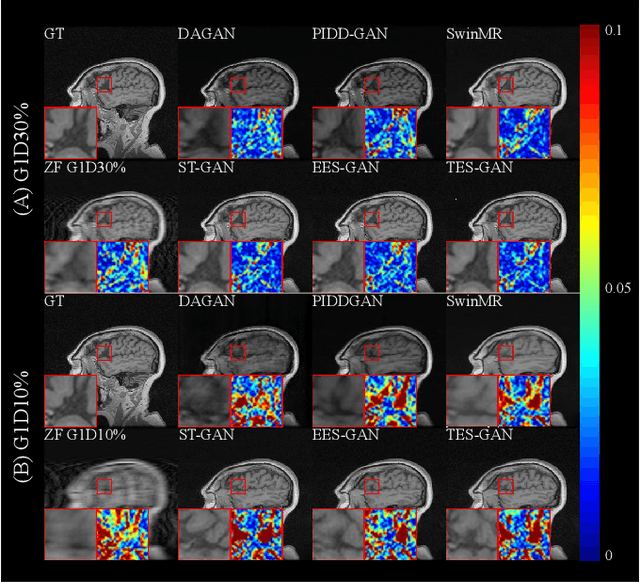
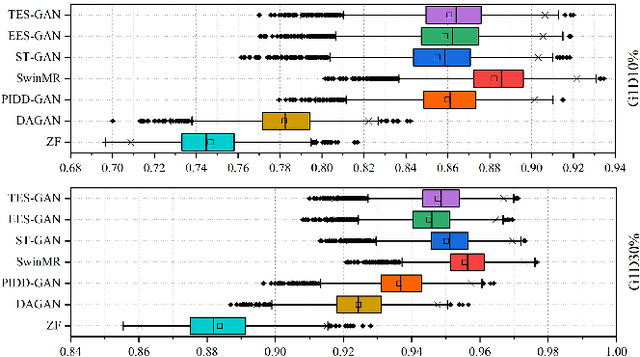
Magnetic resonance imaging (MRI) is a widely used non-radiative and non-invasive method for clinical interrogation of organ structures and metabolism, with an inherently long scanning time. Methods by k-space undersampling and deep learning based reconstruction have been popularised to accelerate the scanning process. This work focuses on investigating how powerful transformers are for fast MRI by exploiting and comparing different novel network architectures. In particular, a generative adversarial network (GAN) based Swin transformer (ST-GAN) was introduced for the fast MRI reconstruction. To further preserve the edge and texture information, edge enhanced GAN based Swin transformer (EESGAN) and texture enhanced GAN based Swin transformer (TES-GAN) were also developed, where a dual-discriminator GAN structure was applied. We compared our proposed GAN based transformers, standalone Swin transformer and other convolutional neural networks based based GAN model in terms of the evaluation metrics PSNR, SSIM and FID. We showed that transformers work well for the MRI reconstruction from different undersampling conditions. The utilisation of GAN's adversarial structure improves the quality of images reconstructed when undersampled for 30% or higher.
Online Attentive Kernel-Based Temporal Difference Learning
Jan 22, 2022Guang Yang, Xingguo Chen, Shangdong Yang, Huihui Wang, Shaokang Dong, Yang Gao
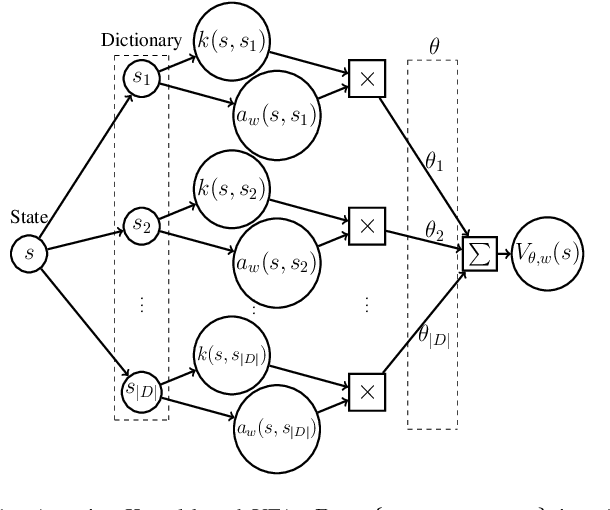
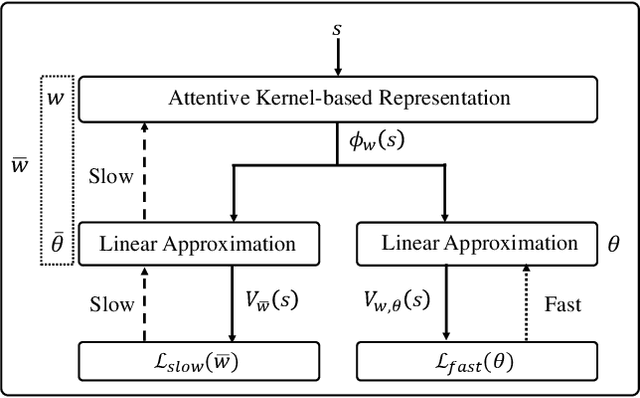
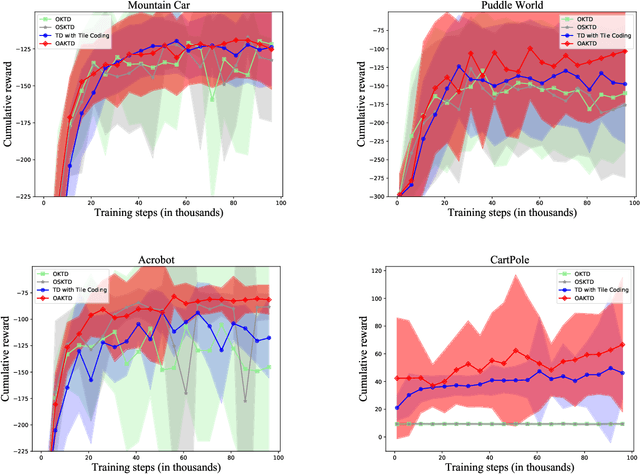
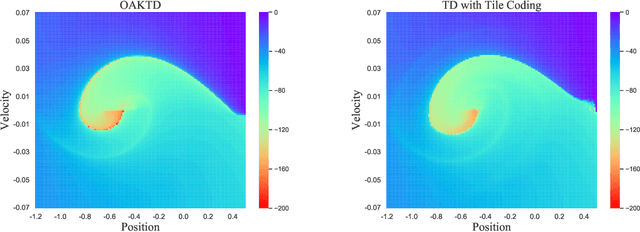
With rising uncertainty in the real world, online Reinforcement Learning (RL) has been receiving increasing attention due to its fast learning capability and improving data efficiency. However, online RL often suffers from complex Value Function Approximation (VFA) and catastrophic interference, creating difficulty for the deep neural network to be applied to an online RL algorithm in a fully online setting. Therefore, a simpler and more adaptive approach is introduced to evaluate value function with the kernel-based model. Sparse representations are superior at handling interference, indicating that competitive sparse representations should be learnable, non-prior, non-truncated and explicit when compared with current sparse representation methods. Moreover, in learning sparse representations, attention mechanisms are utilized to represent the degree of sparsification, and a smooth attentive function is introduced into the kernel-based VFA. In this paper, we propose an Online Attentive Kernel-Based Temporal Difference (OAKTD) algorithm using two-timescale optimization and provide convergence analysis of our proposed algorithm. Experimental evaluations showed that OAKTD outperformed several Online Kernel-based Temporal Difference (OKTD) learning algorithms in addition to the Temporal Difference (TD) learning algorithm with Tile Coding on public Mountain Car, Acrobot, CartPole and Puddle World tasks.
Data Harmonisation for Information Fusion in Digital Healthcare: A State-of-the-Art Systematic Review, Meta-Analysis and Future Research Directions
Jan 17, 2022Yang Nan, Javier Del Ser, Simon Walsh, Carola Schönlieb, Michael Roberts, Ian Selby, Kit Howard, John Owen, Jon Neville, Julien Guiot, Benoit Ernst, Ana Pastor, Angel Alberich-Bayarri, Marion I. Menzel, Sean Walsh, Wim Vos, Nina Flerin, Jean-Paul Charbonnier, Eva van Rikxoort, Avishek Chatterjee, Henry Woodruff, Philippe Lambin, Leonor Cerdá-Alberich, Luis Martí-Bonmatí, Francisco Herrera, Guang Yang
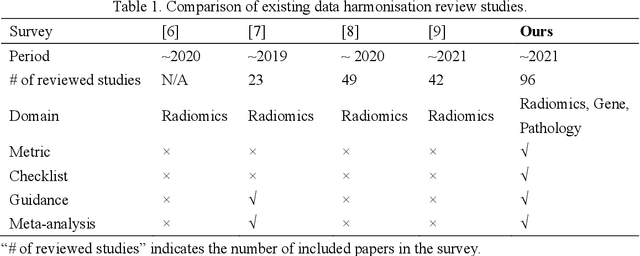
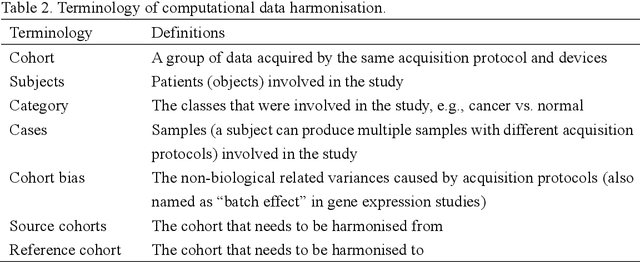



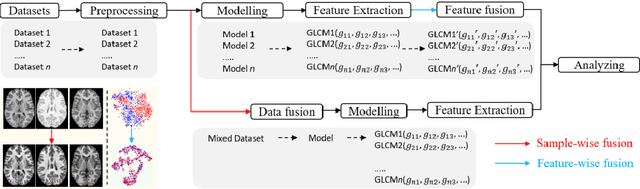
Removing the bias and variance of multicentre data has always been a challenge in large scale digital healthcare studies, which requires the ability to integrate clinical features extracted from data acquired by different scanners and protocols to improve stability and robustness. Previous studies have described various computational approaches to fuse single modality multicentre datasets. However, these surveys rarely focused on evaluation metrics and lacked a checklist for computational data harmonisation studies. In this systematic review, we summarise the computational data harmonisation approaches for multi-modality data in the digital healthcare field, including harmonisation strategies and evaluation metrics based on different theories. In addition, a comprehensive checklist that summarises common practices for data harmonisation studies is proposed to guide researchers to report their research findings more effectively. Last but not least, flowcharts presenting possible ways for methodology and metric selection are proposed and the limitations of different methods have been surveyed for future research.
Swin Transformer for Fast MRI
Jan 10, 2022Jiahao Huang, Yingying Fang, Yinzhe Wu, Huanjun Wu, Zhifan Gao, Yang Li, Javier Del Ser, Jun Xia, Guang Yang
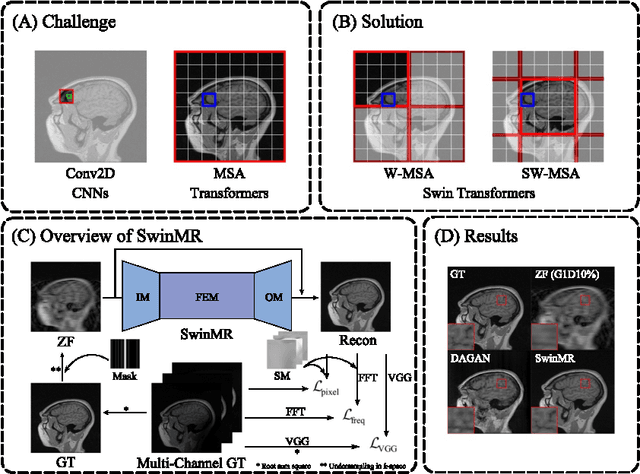
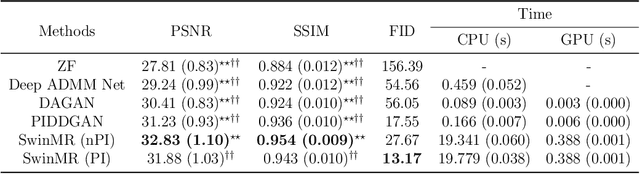
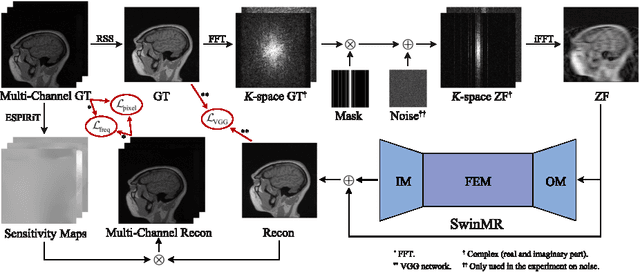
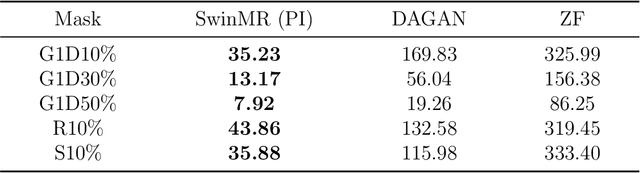
Magnetic resonance imaging (MRI) is an important non-invasive clinical tool that can produce high-resolution and reproducible images. However, a long scanning time is required for high-quality MR images, which leads to exhaustion and discomfort of patients, inducing more artefacts due to voluntary movements of the patients and involuntary physiological movements. To accelerate the scanning process, methods by k-space undersampling and deep learning based reconstruction have been popularised. This work introduced SwinMR, a novel Swin transformer based method for fast MRI reconstruction. The whole network consisted of an input module (IM), a feature extraction module (FEM) and an output module (OM). The IM and OM were 2D convolutional layers and the FEM was composed of a cascaded of residual Swin transformer blocks (RSTBs) and 2D convolutional layers. The RSTB consisted of a series of Swin transformer layers (STLs). The shifted windows multi-head self-attention (W-MSA/SW-MSA) of STL was performed in shifted windows rather than the multi-head self-attention (MSA) of the original transformer in the whole image space. A novel multi-channel loss was proposed by using the sensitivity maps, which was proved to reserve more textures and details. We performed a series of comparative studies and ablation studies in the Calgary-Campinas public brain MR dataset and conducted a downstream segmentation experiment in the Multi-modal Brain Tumour Segmentation Challenge 2017 dataset. The results demonstrate our SwinMR achieved high-quality reconstruction compared with other benchmark methods, and it shows great robustness with different undersampling masks, under noise interruption and on different datasets. The code is publicly available at https://github.com/ayanglab/SwinMR.
AI-based Reconstruction for Fast MRI -- A Systematic Review and Meta-analysis
Dec 23, 2021Yutong Chen, Carola-Bibiane Schönlieb, Pietro Liò, Tim Leiner, Pier Luigi Dragotti, Ge Wang, Daniel Rueckert, David Firmin, Guang Yang
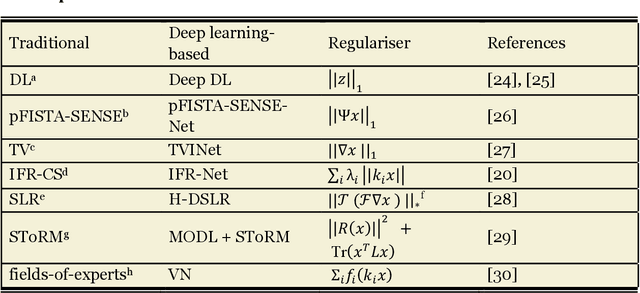

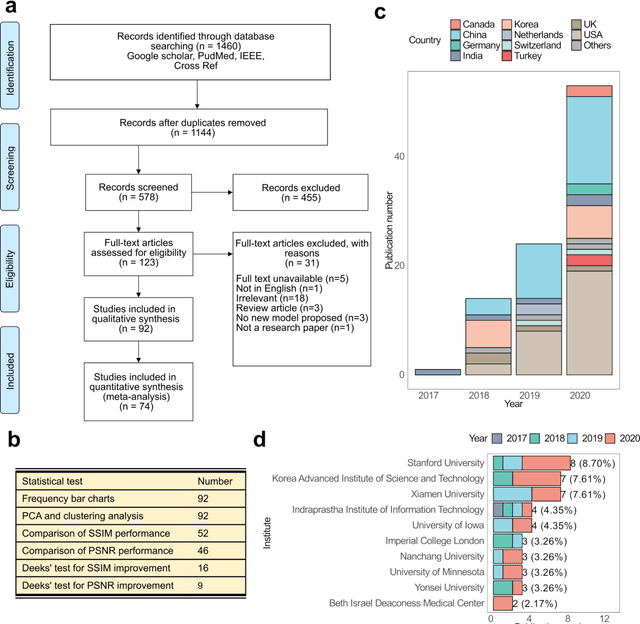
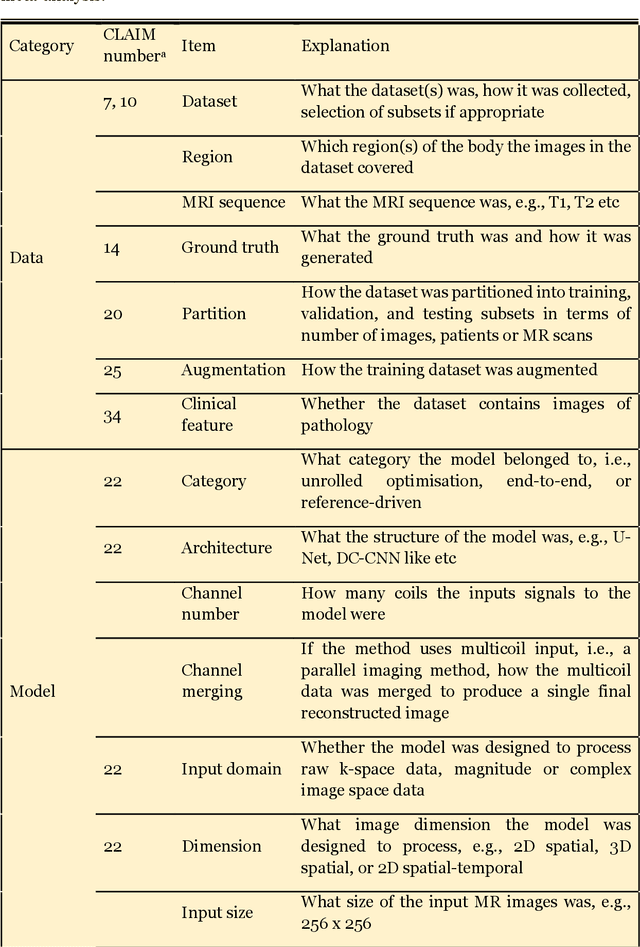
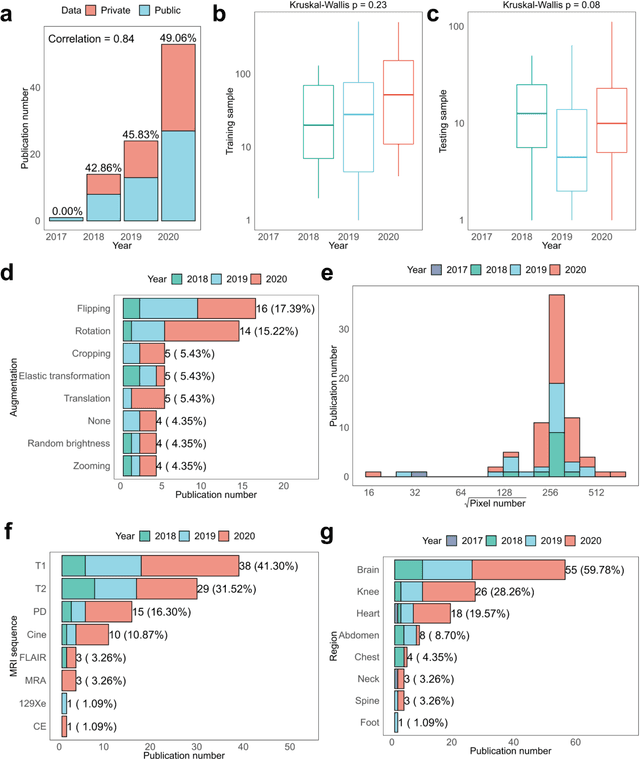
Compressed sensing (CS) has been playing a key role in accelerating the magnetic resonance imaging (MRI) acquisition process. With the resurgence of artificial intelligence, deep neural networks and CS algorithms are being integrated to redefine the state of the art of fast MRI. The past several years have witnessed substantial growth in the complexity, diversity, and performance of deep learning-based CS techniques that are dedicated to fast MRI. In this meta-analysis, we systematically review the deep learning-based CS techniques for fast MRI, describe key model designs, highlight breakthroughs, and discuss promising directions. We have also introduced a comprehensive analysis framework and a classification system to assess the pivotal role of deep learning in CS-based acceleration for MRI.
Edge-Enhanced Dual Discriminator Generative Adversarial Network for Fast MRI with Parallel Imaging Using Multi-view Information
Dec 10, 2021Jiahao Huang, Weiping Ding, Jun Lv, Jingwen Yang, Hao Dong, Javier Del Ser, Jun Xia, Tiaojuan Ren, Stephen Wong, Guang Yang
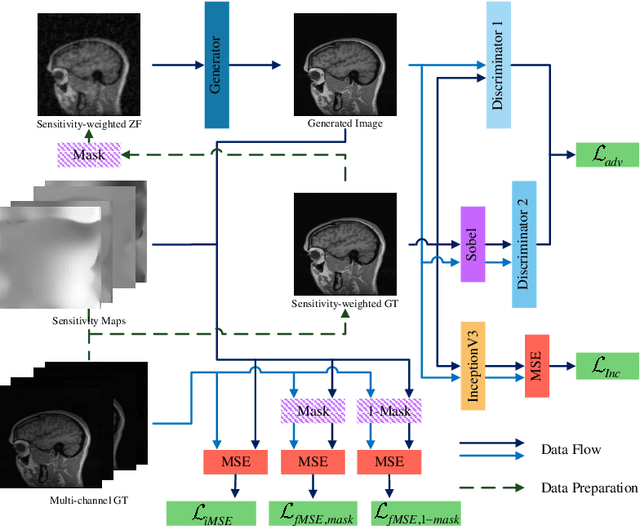
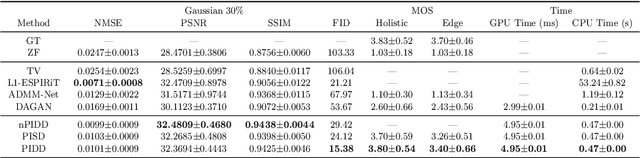
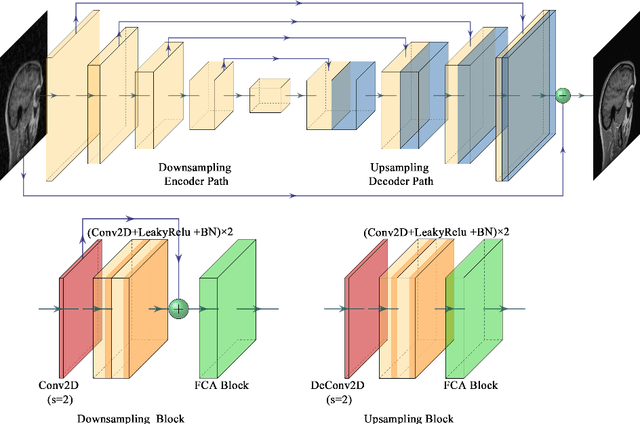
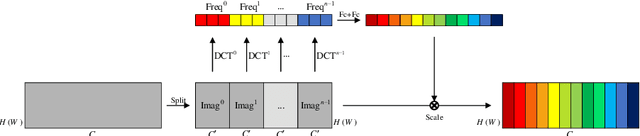
In clinical medicine, magnetic resonance imaging (MRI) is one of the most important tools for diagnosis, triage, prognosis, and treatment planning. However, MRI suffers from an inherent slow data acquisition process because data is collected sequentially in k-space. In recent years, most MRI reconstruction methods proposed in the literature focus on holistic image reconstruction rather than enhancing the edge information. This work steps aside this general trend by elaborating on the enhancement of edge information. Specifically, we introduce a novel parallel imaging coupled dual discriminator generative adversarial network (PIDD-GAN) for fast multi-channel MRI reconstruction by incorporating multi-view information. The dual discriminator design aims to improve the edge information in MRI reconstruction. One discriminator is used for holistic image reconstruction, whereas the other one is responsible for enhancing edge information. An improved U-Net with local and global residual learning is proposed for the generator. Frequency channel attention blocks (FCA Blocks) are embedded in the generator for incorporating attention mechanisms. Content loss is introduced to train the generator for better reconstruction quality. We performed comprehensive experiments on Calgary-Campinas public brain MR dataset and compared our method with state-of-the-art MRI reconstruction methods. Ablation studies of residual learning were conducted on the MICCAI13 dataset to validate the proposed modules. Results show that our PIDD-GAN provides high-quality reconstructed MR images, with well-preserved edge information. The time of single-image reconstruction is below 5ms, which meets the demand of faster processing.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge