Guang Yang
Shanghai Key Laboratory of Magnetic Resonance, School of Physics and Electronic Science, East China Normal University, Shanghai, China, Institute of Magnetic Resonance and Molecular Imaging in Medicine, East China Normal University, Shanghai, China
Large Language Models Meet Virtual Cell: A Survey
Oct 09, 2025Abstract:Large language models (LLMs) are transforming cellular biology by enabling the development of "virtual cells"--computational systems that represent, predict, and reason about cellular states and behaviors. This work provides a comprehensive review of LLMs for virtual cell modeling. We propose a unified taxonomy that organizes existing methods into two paradigms: LLMs as Oracles, for direct cellular modeling, and LLMs as Agents, for orchestrating complex scientific tasks. We identify three core tasks--cellular representation, perturbation prediction, and gene regulation inference--and review their associated models, datasets, evaluation benchmarks, as well as the critical challenges in scalability, generalizability, and interpretability.
First-order State Space Model for Lightweight Image Super-resolution
Sep 10, 2025



Abstract:State space models (SSMs), particularly Mamba, have shown promise in NLP tasks and are increasingly applied to vision tasks. However, most Mamba-based vision models focus on network architecture and scan paths, with little attention to the SSM module. In order to explore the potential of SSMs, we modified the calculation process of SSM without increasing the number of parameters to improve the performance on lightweight super-resolution tasks. In this paper, we introduce the First-order State Space Model (FSSM) to improve the original Mamba module, enhancing performance by incorporating token correlations. We apply a first-order hold condition in SSMs, derive the new discretized form, and analyzed cumulative error. Extensive experimental results demonstrate that FSSM improves the performance of MambaIR on five benchmark datasets without additionally increasing the number of parameters, and surpasses current lightweight SR methods, achieving state-of-the-art results.
* Accept by ICASSP 2025 (Oral)
Forewarned is Forearmed: Pre-Synthesizing Jailbreak-like Instructions to Enhance LLM Safety Guardrail to Potential Attacks
Aug 28, 2025Abstract:Despite advances in improving large language model (LLM) to refuse to answer malicious instructions, widely used LLMs remain vulnerable to jailbreak attacks where attackers generate instructions with distributions differing from safety alignment corpora. New attacks expose LLMs' inability to recognize unseen malicious instructions, highlighting a critical distributional mismatch between training data and real-world attacks that forces developers into reactive patching cycles. To tackle this challenge, we propose IMAGINE, a synthesis framework that leverages embedding space distribution analysis to generate jailbreak-like instructions. This approach effectively fills the distributional gap between authentic jailbreak patterns and safety alignment corpora. IMAGINE follows an iterative optimization process that dynamically evolves text generation distributions across iterations, thereby augmenting the coverage of safety alignment data distributions through synthesized data examples. Based on the safety-aligned corpus enhanced through IMAGINE, our framework demonstrates significant decreases in attack success rate on Qwen2.5, Llama3.1, and Llama3.2 without compromising their utility.
A Physics-Driven Neural Network with Parameter Embedding for Generating Quantitative MR Maps from Weighted Images
Aug 11, 2025Abstract:We propose a deep learning-based approach that integrates MRI sequence parameters to improve the accuracy and generalizability of quantitative image synthesis from clinical weighted MRI. Our physics-driven neural network embeds MRI sequence parameters -- repetition time (TR), echo time (TE), and inversion time (TI) -- directly into the model via parameter embedding, enabling the network to learn the underlying physical principles of MRI signal formation. The model takes conventional T1-weighted, T2-weighted, and T2-FLAIR images as input and synthesizes T1, T2, and proton density (PD) quantitative maps. Trained on healthy brain MR images, it was evaluated on both internal and external test datasets. The proposed method achieved high performance with PSNR values exceeding 34 dB and SSIM values above 0.92 for all synthesized parameter maps. It outperformed conventional deep learning models in accuracy and robustness, including data with previously unseen brain structures and lesions. Notably, our model accurately synthesized quantitative maps for these unseen pathological regions, highlighting its superior generalization capability. Incorporating MRI sequence parameters via parameter embedding allows the neural network to better learn the physical characteristics of MR signals, significantly enhancing the performance and reliability of quantitative MRI synthesis. This method shows great potential for accelerating qMRI and improving its clinical utility.
Conformal Sets in Multiple-Choice Question Answering under Black-Box Settings with Provable Coverage Guarantees
Aug 07, 2025Abstract:Large Language Models (LLMs) have shown remarkable progress in multiple-choice question answering (MCQA), but their inherent unreliability, such as hallucination and overconfidence, limits their application in high-risk domains. To address this, we propose a frequency-based uncertainty quantification method under black-box settings, leveraging conformal prediction (CP) to ensure provable coverage guarantees. Our approach involves multiple independent samplings of the model's output distribution for each input, with the most frequent sample serving as a reference to calculate predictive entropy (PE). Experimental evaluations across six LLMs and four datasets (MedMCQA, MedQA, MMLU, MMLU-Pro) demonstrate that frequency-based PE outperforms logit-based PE in distinguishing between correct and incorrect predictions, as measured by AUROC. Furthermore, the method effectively controls the empirical miscoverage rate under user-specified risk levels, validating that sampling frequency can serve as a viable substitute for logit-based probabilities in black-box scenarios. This work provides a distribution-free model-agnostic framework for reliable uncertainty quantification in MCQA with guaranteed coverage, enhancing the trustworthiness of LLMs in practical applications.
AsyncFlow: An Asynchronous Streaming RL Framework for Efficient LLM Post-Training
Jul 02, 2025Abstract:Reinforcement learning (RL) has become a pivotal technology in the post-training phase of large language models (LLMs). Traditional task-colocated RL frameworks suffer from significant scalability bottlenecks, while task-separated RL frameworks face challenges in complex dataflows and the corresponding resource idling and workload imbalance. Moreover, most existing frameworks are tightly coupled with LLM training or inference engines, making it difficult to support custom-designed engines. To address these challenges, we propose AsyncFlow, an asynchronous streaming RL framework for efficient post-training. Specifically, we introduce a distributed data storage and transfer module that provides a unified data management and fine-grained scheduling capability in a fully streamed manner. This architecture inherently facilitates automated pipeline overlapping among RL tasks and dynamic load balancing. Moreover, we propose a producer-consumer-based asynchronous workflow engineered to minimize computational idleness by strategically deferring parameter update process within staleness thresholds. Finally, the core capability of AsynFlow is architecturally decoupled from underlying training and inference engines and encapsulated by service-oriented user interfaces, offering a modular and customizable user experience. Extensive experiments demonstrate an average of 1.59 throughput improvement compared with state-of-the-art baseline. The presented architecture in this work provides actionable insights for next-generation RL training system designs.
Memory-Augmented Incomplete Multimodal Survival Prediction via Cross-Slide and Gene-Attentive Hypergraph Learning
Jun 24, 2025Abstract:Multimodal pathology-genomic analysis is critical for cancer survival prediction. However, existing approaches predominantly integrate formalin-fixed paraffin-embedded (FFPE) slides with genomic data, while neglecting the availability of other preservation slides, such as Fresh Froze (FF) slides. Moreover, as the high-resolution spatial nature of pathology data tends to dominate the cross-modality fusion process, it hinders effective multimodal fusion and leads to modality imbalance challenges between pathology and genomics. These methods also typically require complete data modalities, limiting their clinical applicability with incomplete modalities, such as missing either pathology or genomic data. In this paper, we propose a multimodal survival prediction framework that leverages hypergraph learning to effectively integrate multi-WSI information and cross-modality interactions between pathology slides and genomics data while addressing modality imbalance. In addition, we introduce a memory mechanism that stores previously learned paired pathology-genomic features and dynamically compensates for incomplete modalities. Experiments on five TCGA datasets demonstrate that our model outperforms advanced methods by over 2.3% in C-Index. Under incomplete modality scenarios, our approach surpasses pathology-only (3.3%) and gene-only models (7.9%). Code: https://github.com/MCPathology/M2Surv
Surgery-R1: Advancing Surgical-VQLA with Reasoning Multimodal Large Language Model via Reinforcement Learning
Jun 24, 2025Abstract:In recent years, significant progress has been made in the field of surgical scene understanding, particularly in the task of Visual Question Localized-Answering in robotic surgery (Surgical-VQLA). However, existing Surgical-VQLA models lack deep reasoning capabilities and interpretability in surgical scenes, which limits their reliability and potential for development in clinical applications. To address this issue, inspired by the development of Reasoning Multimodal Large Language Models (MLLMs), we first build the Surgery-R1-54k dataset, including paired data for Visual-QA, Grounding-QA, and Chain-of-Thought (CoT). Then, we propose the first Reasoning MLLM for Surgical-VQLA (Surgery-R1). In our Surgery-R1, we design a two-stage fine-tuning mechanism to enable the basic MLLM with complex reasoning abilities by utilizing supervised fine-tuning (SFT) and reinforcement fine-tuning (RFT). Furthermore, for an efficient and high-quality rule-based reward system in our RFT, we design a Multimodal Coherence reward mechanism to mitigate positional illusions that may arise in surgical scenarios. Experiment results demonstrate that Surgery-R1 outperforms other existing state-of-the-art (SOTA) models in the Surgical-VQLA task and widely-used MLLMs, while also validating its reasoning capabilities and the effectiveness of our approach. The code and dataset will be organized in https://github.com/FiFi-HAO467/Surgery-R1.
LEGATO: Large-scale End-to-end Generalizable Approach to Typeset OMR
Jun 23, 2025Abstract:We propose Legato, a new end-to-end transformer model for optical music recognition (OMR). Legato is the first large-scale pretrained OMR model capable of recognizing full-page or multi-page typeset music scores and the first to generate documents in ABC notation, a concise, human-readable format for symbolic music. Bringing together a pretrained vision encoder with an ABC decoder trained on a dataset of more than 214K images, our model exhibits the strong ability to generalize across various typeset scores. We conduct experiments on a range of datasets and demonstrate that our model achieves state-of-the-art performance. Given the lack of a standardized evaluation for end-to-end OMR, we comprehensively compare our model against the previous state of the art using a diverse set of metrics.
An LLM-as-Judge Metric for Bridging the Gap with Human Evaluation in SE Tasks
May 27, 2025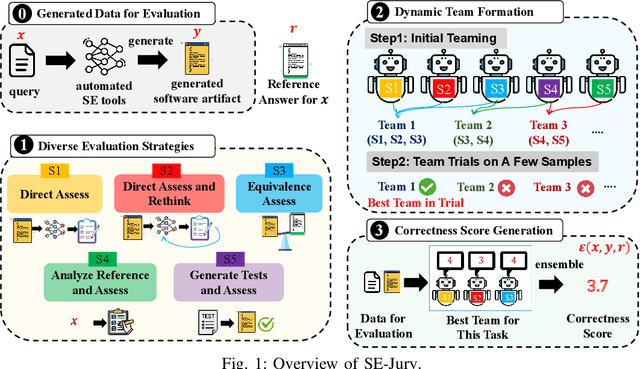
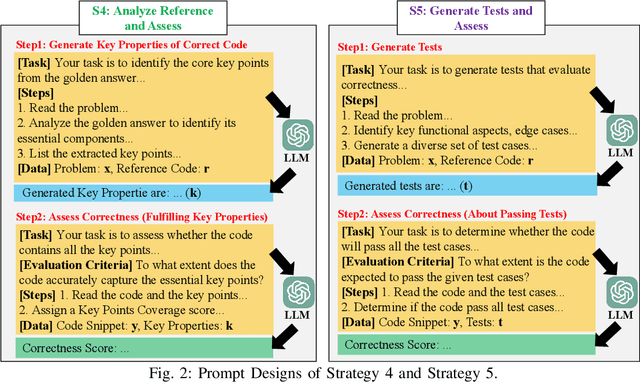
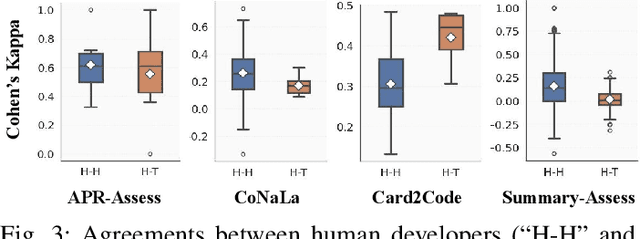
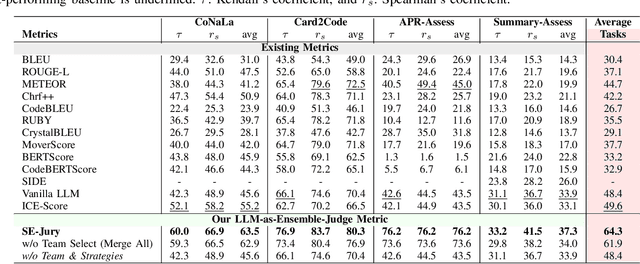
Abstract:Large Language Models (LLMs) and other automated techniques have been increasingly used to support software developers by generating software artifacts such as code snippets, patches, and comments. However, accurately assessing the correctness of these generated artifacts remains a significant challenge. On one hand, human evaluation provides high accuracy but is labor-intensive and lacks scalability. On the other hand, other existing automatic evaluation metrics are scalable and require minimal human effort, but they often fail to accurately reflect the actual correctness of generated software artifacts. In this paper, we present SWE-Judge, the first evaluation metric for LLM-as-Ensemble-Judge specifically designed to accurately assess the correctness of generated software artifacts. SWE-Judge first defines five distinct evaluation strategies, each implemented as an independent judge. A dynamic team selection mechanism then identifies the most appropriate subset of judges to produce a final correctness score through ensembling. We evaluate SWE-Judge across a diverse set of software engineering (SE) benchmarks, including CoNaLa, Card2Code, HumanEval-X, APPS, APR-Assess, and Summary-Assess. These benchmarks span three SE tasks: code generation, automated program repair, and code summarization. Experimental results demonstrate that SWE-Judge consistently achieves a higher correlation with human judgments, with improvements ranging from 5.9% to 183.8% over existing automatic metrics. Furthermore, SWE-Judge reaches agreement levels with human annotators that are comparable to inter-annotator agreement in code generation and program repair tasks. These findings underscore SWE-Judge's potential as a scalable and reliable alternative to human evaluation.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge