Yao Zhang
Shanghai AI Laboratory, China
The Multi-modality Cell Segmentation Challenge: Towards Universal Solutions
Aug 10, 2023Abstract:Cell segmentation is a critical step for quantitative single-cell analysis in microscopy images. Existing cell segmentation methods are often tailored to specific modalities or require manual interventions to specify hyperparameters in different experimental settings. Here, we present a multi-modality cell segmentation benchmark, comprising over 1500 labeled images derived from more than 50 diverse biological experiments. The top participants developed a Transformer-based deep-learning algorithm that not only exceeds existing methods, but can also be applied to diverse microscopy images across imaging platforms and tissue types without manual parameter adjustments. This benchmark and the improved algorithm offer promising avenues for more accurate and versatile cell analysis in microscopy imaging.
Unleashing the Strengths of Unlabeled Data in Pan-cancer Abdominal Organ Quantification: the FLARE22 Challenge
Aug 10, 2023



Abstract:Quantitative organ assessment is an essential step in automated abdominal disease diagnosis and treatment planning. Artificial intelligence (AI) has shown great potential to automatize this process. However, most existing AI algorithms rely on many expert annotations and lack a comprehensive evaluation of accuracy and efficiency in real-world multinational settings. To overcome these limitations, we organized the FLARE 2022 Challenge, the largest abdominal organ analysis challenge to date, to benchmark fast, low-resource, accurate, annotation-efficient, and generalized AI algorithms. We constructed an intercontinental and multinational dataset from more than 50 medical groups, including Computed Tomography (CT) scans with different races, diseases, phases, and manufacturers. We independently validated that a set of AI algorithms achieved a median Dice Similarity Coefficient (DSC) of 90.0\% by using 50 labeled scans and 2000 unlabeled scans, which can significantly reduce annotation requirements. The best-performing algorithms successfully generalized to holdout external validation sets, achieving a median DSC of 89.5\%, 90.9\%, and 88.3\% on North American, European, and Asian cohorts, respectively. They also enabled automatic extraction of key organ biology features, which was labor-intensive with traditional manual measurements. This opens the potential to use unlabeled data to boost performance and alleviate annotation shortages for modern AI models.
Deep learning radiomics for assessment of gastroesophageal varices in people with compensated advanced chronic liver disease
Jun 13, 2023



Abstract:Objective: Bleeding from gastroesophageal varices (GEV) is a medical emergency associated with high mortality. We aim to construct an artificial intelligence-based model of two-dimensional shear wave elastography (2D-SWE) of the liver and spleen to precisely assess the risk of GEV and high-risk gastroesophageal varices (HRV). Design: A prospective multicenter study was conducted in patients with compensated advanced chronic liver disease. 305 patients were enrolled from 12 hospitals, and finally 265 patients were included, with 1136 liver stiffness measurement (LSM) images and 1042 spleen stiffness measurement (SSM) images generated by 2D-SWE. We leveraged deep learning methods to uncover associations between image features and patient risk, and thus conducted models to predict GEV and HRV. Results: A multi-modality Deep Learning Risk Prediction model (DLRP) was constructed to assess GEV and HRV, based on LSM and SSM images, and clinical information. Validation analysis revealed that the AUCs of DLRP were 0.91 for GEV (95% CI 0.90 to 0.93, p < 0.05) and 0.88 for HRV (95% CI 0.86 to 0.89, p < 0.01), which were significantly and robustly better than canonical risk indicators, including the value of LSM and SSM. Moreover, DLPR was better than the model using individual parameters, including LSM and SSM images. In HRV prediction, the 2D-SWE images of SSM outperform LSM (p < 0.01). Conclusion: DLRP shows excellent performance in predicting GEV and HRV over canonical risk indicators LSM and SSM. Additionally, the 2D-SWE images of SSM provided more information for better accuracy in predicting HRV than the LSM.
Chinese Intermediate English Learners outdid ChatGPT in deep cohesion: Evidence from English narrative writing
Mar 21, 2023



Abstract:ChatGPT is a publicly available chatbot that can quickly generate texts on given topics, but it is unknown whether the chatbot is really superior to human writers in all aspects of writing and whether its writing quality can be prominently improved on the basis of updating commands. Consequently, this study compared the writing performance on a narrative topic by ChatGPT and Chinese intermediate English (CIE) learners so as to reveal the chatbot's advantage and disadvantage in writing. The data were analyzed in terms of five discourse components using Coh-Metrix (a special instrument for analyzing language discourses), and the results revealed that ChatGPT performed better than human writers in narrativity, word concreteness, and referential cohesion, but worse in syntactic simplicity and deep cohesion in its initial version. After more revision commands were updated, while the resulting version was facilitated in syntactic simplicity, yet it is still lagged far behind CIE learners' writing in deep cohesion. In addition, the correlation analysis of the discourse components suggests that narrativity was correlated with referential cohesion in both ChatGPT and human writers, but the correlations varied within each group.
High speed free-space optical communication using standard fiber communication component without optical amplification
Feb 27, 2023



Abstract:Free-space optical communication (FSO) can achieve fast, secure and license-free communication without need for physical cables, making it a cost-effective, energy-efficient and flexible solution when the fiber connection is absent. To establish FSO connection on-demand, it is essential to build portable FSO devices with compact structure and light weight. Here, we develop a miniaturized FSO system and realize 9.16 Gbps FSO between two nodes that is 1 km apart, using a commercial fiber-coupled optical transceiver module with no optical amplification. Basing on the home-made compact 90 mm-diameter acquisition, pointing and tracking (APT) system with four-stage close-loop feedback, the link tracking error is controlled at 3 {\mu}rad and results an average coupling loss of 13.7 dB. Such loss is within the tolerance of the commercial optical communication modules, and without the need of optical amplifiers, which contributes to the low system weight and power consumption. As a result, a single FSO device weighs only about 12 kg, making it compact and portable for potential application in high-speed wireless communication. Our FSO link has been tested up to 4 km, with link loss of 18 dB in the foggy weather in Nanjing, that shows longer distances can be covered with optical amplification.
TIGER: Temporal Interaction Graph Embedding with Restarts
Feb 16, 2023



Abstract:Temporal interaction graphs (TIGs), consisting of sequences of timestamped interaction events, are prevalent in fields like e-commerce and social networks. To better learn dynamic node embeddings that vary over time, researchers have proposed a series of temporal graph neural networks for TIGs. However, due to the entangled temporal and structural dependencies, existing methods have to process the sequence of events chronologically and consecutively to ensure node representations are up-to-date. This prevents existing models from parallelization and reduces their flexibility in industrial applications. To tackle the above challenge, in this paper, we propose TIGER, a TIG embedding model that can restart at any timestamp. We introduce a restarter module that generates surrogate representations acting as the warm initialization of node representations. By restarting from multiple timestamps simultaneously, we divide the sequence into multiple chunks and naturally enable the parallelization of the model. Moreover, in contrast to previous models that utilize a single memory unit, we introduce a dual memory module to better exploit neighborhood information and alleviate the staleness problem. Extensive experiments on four public datasets and one industrial dataset are conducted, and the results verify both the effectiveness and the efficiency of our work.
ConsRec: Learning Consensus Behind Interactions for Group Recommendation
Feb 07, 2023
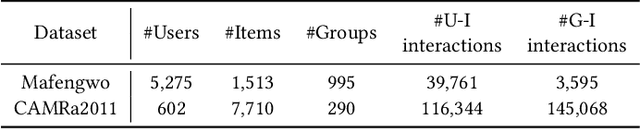


Abstract:Since group activities have become very common in daily life, there is an urgent demand for generating recommendations for a group of users, referred to as group recommendation task. Existing group recommendation methods usually infer groups' preferences via aggregating diverse members' interests. Actually, groups' ultimate choice involves compromises between members, and finally, an agreement can be reached. However, existing individual information aggregation lacks a holistic group-level consideration, failing to capture the consensus information. Besides, their specific aggregation strategies either suffer from high computational costs or become too coarse-grained to make precise predictions. To solve the aforementioned limitations, in this paper, we focus on exploring consensus behind group behavior data. To comprehensively capture the group consensus, we innovatively design three distinct views which provide mutually complementary information to enable multi-view learning, including member-level aggregation, item-level tastes, and group-level inherent preferences. To integrate and balance the multi-view information, an adaptive fusion component is further proposed. As to member-level aggregation, different from existing linear or attentive strategies, we design a novel hypergraph neural network that allows for efficient hypergraph convolutional operations to generate expressive member-level aggregation. We evaluate our ConsRec on two real-world datasets and experimental results show that our model outperforms state-of-the-art methods. An extensive case study also verifies the effectiveness of consensus modeling.
CL-CrossVQA: A Continual Learning Benchmark for Cross-Domain Visual Question Answering
Nov 19, 2022



Abstract:Visual Question Answering (VQA) is a multi-discipline research task. To produce the right answer, it requires an understanding of the visual content of images, the natural language questions, as well as commonsense reasoning over the information contained in the image and world knowledge. Recently, large-scale Vision-and-Language Pre-trained Models (VLPMs) have been the mainstream approach to VQA tasks due to their superior performance. The standard practice is to fine-tune large-scale VLPMs pre-trained on huge general-domain datasets using the domain-specific VQA datasets. However, in reality, the application domain can change over time, necessitating VLPMs to continually learn and adapt to new domains without forgetting previously acquired knowledge. Most existing continual learning (CL) research concentrates on unimodal tasks, whereas a more practical application scenario, i.e, CL on cross-domain VQA, has not been studied. Motivated by this, we introduce CL-CrossVQA, a rigorous Continual Learning benchmark for Cross-domain Visual Question Answering, through which we conduct extensive experiments on 4 VLPMs, 4 CL approaches, and 5 VQA datasets from different domains. In addition, by probing the forgetting phenomenon of the intermediate layers, we provide insights into how model architecture affects CL performance, why CL approaches can help mitigate forgetting in VLPMs to some extent, and how to design CL approaches suitable for VLPMs in this challenging continual learning environment. To facilitate future work on CL for cross-domain VQA, we will release our datasets and code.
Learning to Learn Domain-invariant Parameters for Domain Generalization
Nov 04, 2022



Abstract:Due to domain shift, deep neural networks (DNNs) usually fail to generalize well on unknown test data in practice. Domain generalization (DG) aims to overcome this issue by capturing domain-invariant representations from source domains. Motivated by the insight that only partial parameters of DNNs are optimized to extract domain-invariant representations, we expect a general model that is capable of well perceiving and emphatically updating such domain-invariant parameters. In this paper, we propose two modules of Domain Decoupling and Combination (DDC) and Domain-invariance-guided Backpropagation (DIGB), which can encourage such general model to focus on the parameters that have a unified optimization direction between pairs of contrastive samples. Our extensive experiments on two benchmarks have demonstrated that our proposed method has achieved state-of-the-art performance with strong generalization capability.
SAP-DETR: Bridging the Gap Between Salient Points and Queries-Based Transformer Detector for Fast Model Convergency
Nov 03, 2022Abstract:Recently, the dominant DETR-based approaches apply central-concept spatial prior to accelerate Transformer detector convergency. These methods gradually refine the reference points to the center of target objects and imbue object queries with the updated central reference information for spatially conditional attention. However, centralizing reference points may severely deteriorate queries' saliency and confuse detectors due to the indiscriminative spatial prior. To bridge the gap between the reference points of salient queries and Transformer detectors, we propose SAlient Point-based DETR (SAP-DETR) by treating object detection as a transformation from salient points to instance objects. In SAP-DETR, we explicitly initialize a query-specific reference point for each object query, gradually aggregate them into an instance object, and then predict the distance from each side of the bounding box to these points. By rapidly attending to query-specific reference region and other conditional extreme regions from the image features, SAP-DETR can effectively bridge the gap between the salient point and the query-based Transformer detector with a significant convergency speed. Our extensive experiments have demonstrated that SAP-DETR achieves 1.4 times convergency speed with competitive performance. Under the standard training scheme, SAP-DETR stably promotes the SOTA approaches by 1.0 AP. Based on ResNet-DC-101, SAP-DETR achieves 46.9 AP.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge