Qianni Zhang
GT U-Net: A U-Net Like Group Transformer Network for Tooth Root Segmentation
Sep 30, 2021
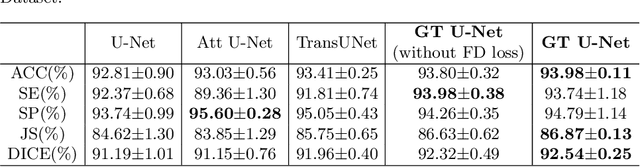
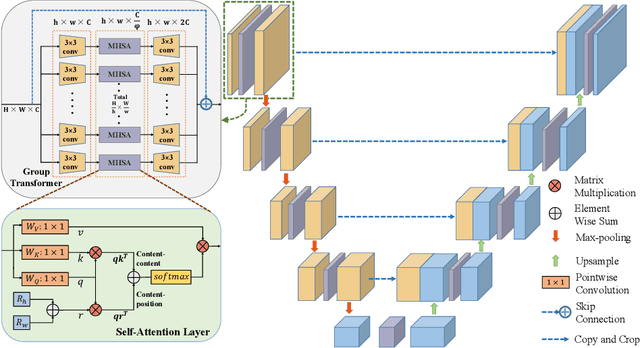
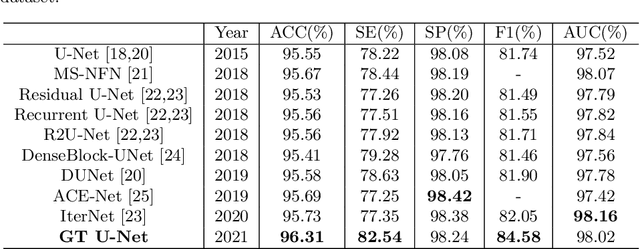
Abstract:To achieve an accurate assessment of root canal therapy, a fundamental step is to perform tooth root segmentation on oral X-ray images, in that the position of tooth root boundary is significant anatomy information in root canal therapy evaluation. However, the fuzzy boundary makes the tooth root segmentation very challenging. In this paper, we propose a novel end-to-end U-Net like Group Transformer Network (GT U-Net) for the tooth root segmentation. The proposed network retains the essential structure of U-Net but each of the encoders and decoders is replaced by a group Transformer, which significantly reduces the computational cost of traditional Transformer architectures by using the grouping structure and the bottleneck structure. In addition, the proposed GT U-Net is composed of a hybrid structure of convolution and Transformer, which makes it independent of pre-training weights. For optimization, we also propose a shape-sensitive Fourier Descriptor (FD) loss function to make use of shape prior knowledge. Experimental results show that our proposed network achieves the state-of-the-art performance on our collected tooth root segmentation dataset and the public retina dataset DRIVE. Code has been released at https://github.com/Kent0n-Li/GT-U-Net.
Structure-aware scale-adaptive networks for cancer segmentation in whole-slide images
Sep 26, 2021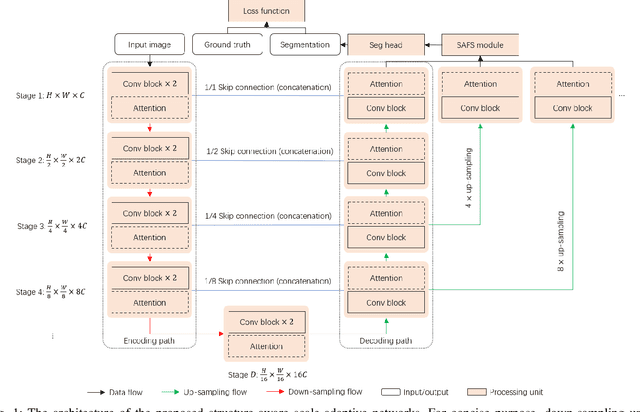
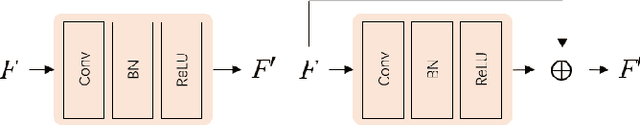
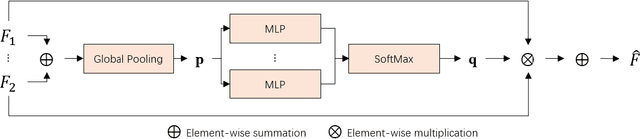
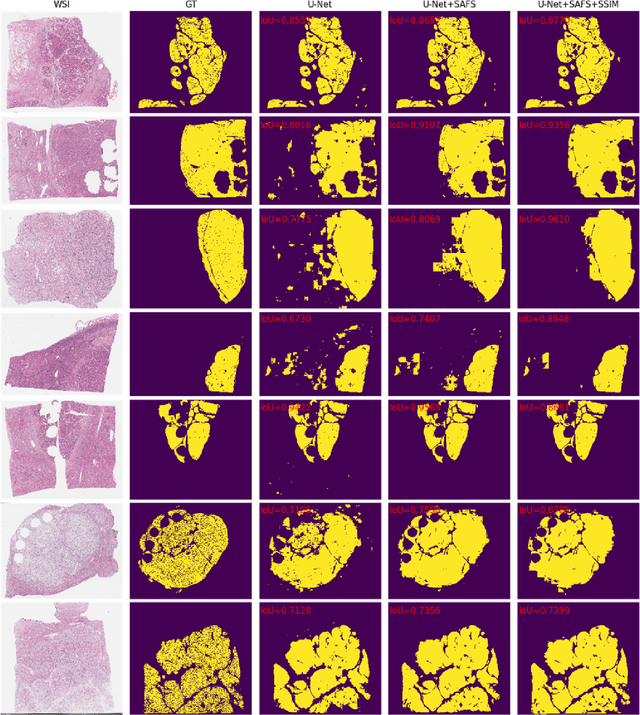
Abstract:Cancer segmentation in whole-slide images is a fundamental step for viable tumour burden estimation, which is of great value for cancer assessment. However, factors like vague boundaries or small regions dissociated from viable tumour areas make it a challenging task. Considering the usefulness of multi-scale features in various vision-related tasks, we present a structure-aware scale-adaptive feature selection method for efficient and accurate cancer segmentation. Based on a segmentation network with a popular encoder-decoder architecture, a scale-adaptive module is proposed for selecting more robust features to represent the vague, non-rigid boundaries. Furthermore, a structural similarity metric is proposed for better tissue structure awareness to deal with small region segmentation. In addition, advanced designs including several attention mechanisms and the selective-kernel convolutions are applied to the baseline network for comparative study purposes. Extensive experimental results show that the proposed structure-aware scale-adaptive networks achieve outstanding performance on liver cancer segmentation when compared to top ten submitted results in the challenge of PAIP 2019. Further evaluation on colorectal cancer segmentation shows that the scale-adaptive module improves the baseline network or outperforms the other excellent designs of attention mechanisms when considering the tradeoff between efficiency and accuracy.
Magnification-independent Histopathological Image Classification with Similarity-based Multi-scale Embeddings
Jul 02, 2021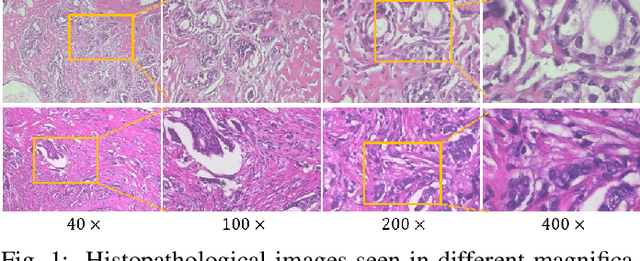
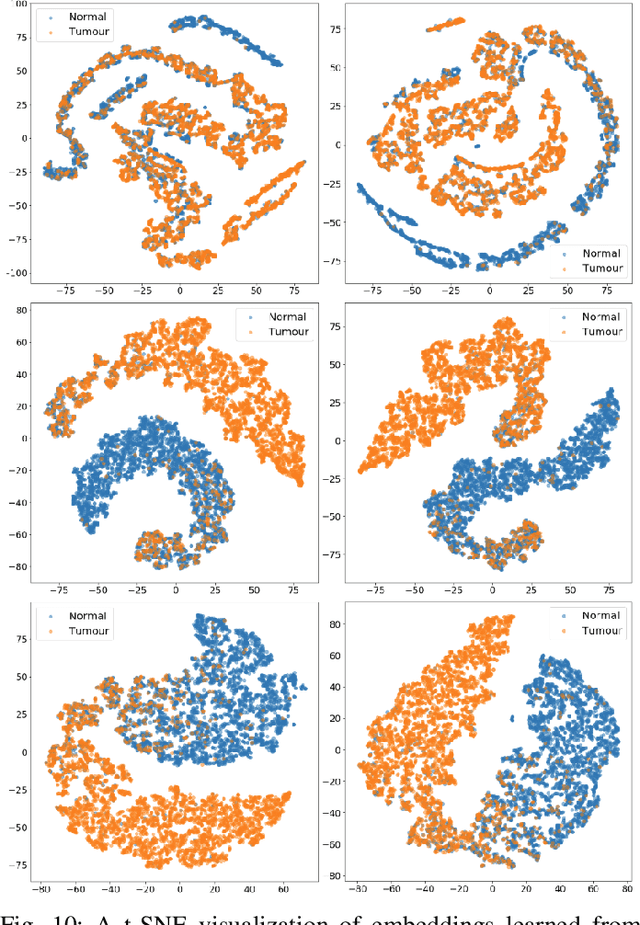
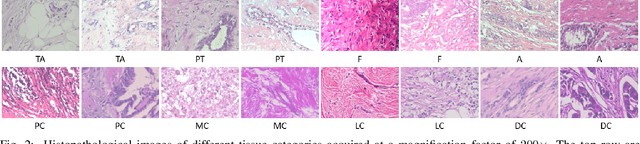
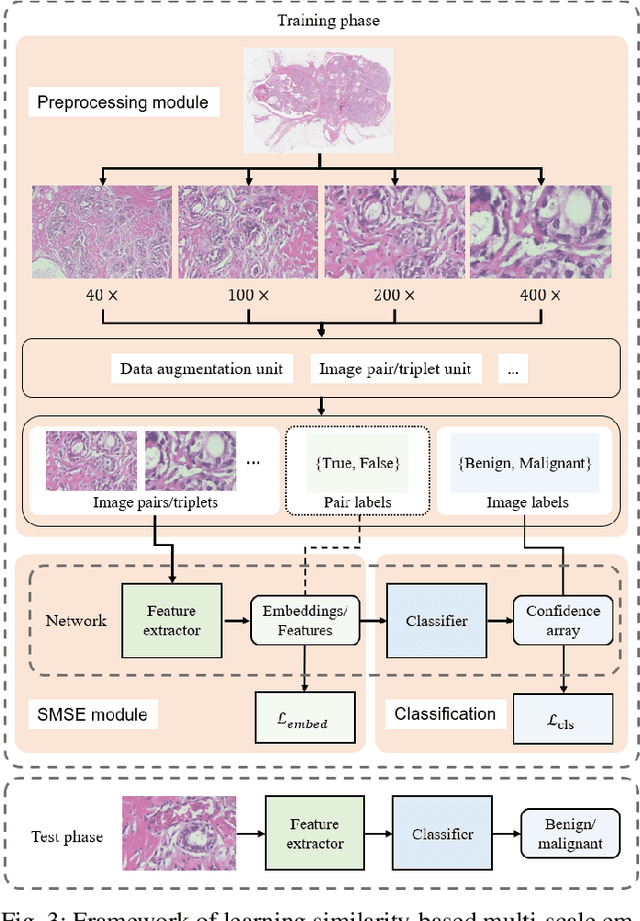
Abstract:The classification of histopathological images is of great value in both cancer diagnosis and pathological studies. However, multiple reasons, such as variations caused by magnification factors and class imbalance, make it a challenging task where conventional methods that learn from image-label datasets perform unsatisfactorily in many cases. We observe that tumours of the same class often share common morphological patterns. To exploit this fact, we propose an approach that learns similarity-based multi-scale embeddings (SMSE) for magnification-independent histopathological image classification. In particular, a pair loss and a triplet loss are leveraged to learn similarity-based embeddings from image pairs or image triplets. The learned embeddings provide accurate measurements of similarities between images, which are regarded as a more effective form of representation for histopathological morphology than normal image features. Furthermore, in order to ensure the generated models are magnification-independent, images acquired at different magnification factors are simultaneously fed to networks during training for learning multi-scale embeddings. In addition to the SMSE, to eliminate the impact of class imbalance, instead of using the hard sample mining strategy that intuitively discards some easy samples, we introduce a new reinforced focal loss to simultaneously punish hard misclassified samples while suppressing easy well-classified samples. Experimental results show that the SMSE improves the performance for histopathological image classification tasks for both breast and liver cancers by a large margin compared to previous methods. In particular, the SMSE achieves the best performance on the BreakHis benchmark with an improvement ranging from 5% to 18% compared to previous methods using traditional features.
SRPN: similarity-based region proposal networks for nuclei and cells detection in histology images
Jun 25, 2021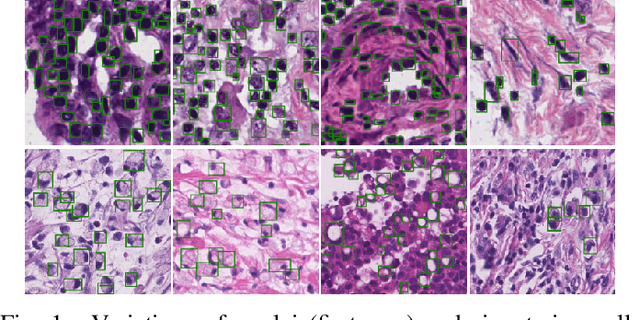
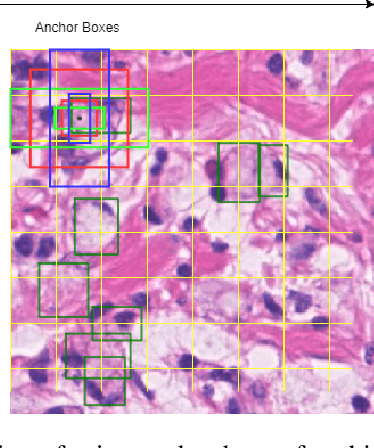
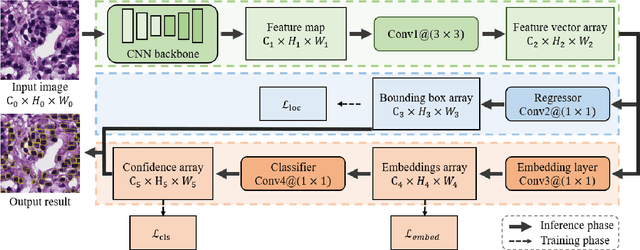
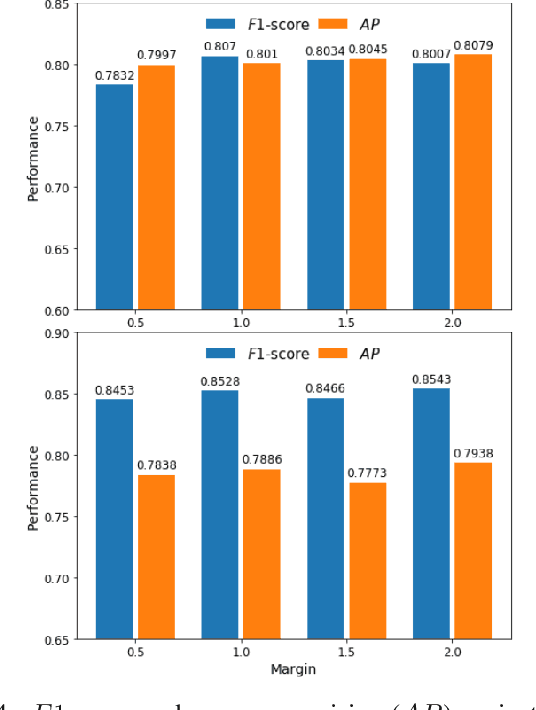
Abstract:The detection of nuclei and cells in histology images is of great value in both clinical practice and pathological studies. However, multiple reasons such as morphological variations of nuclei or cells make it a challenging task where conventional object detection methods cannot obtain satisfactory performance in many cases. A detection task consists of two sub-tasks, classification and localization. Under the condition of dense object detection, classification is a key to boost the detection performance. Considering this, we propose similarity based region proposal networks (SRPN) for nuclei and cells detection in histology images. In particular, a customized convolution layer termed as embedding layer is designed for network building. The embedding layer is added into the region proposal networks, enabling the networks to learn discriminative features based on similarity learning. Features obtained by similarity learning can significantly boost the classification performance compared to conventional methods. SRPN can be easily integrated into standard convolutional neural networks architectures such as the Faster R-CNN and RetinaNet. We test the proposed approach on tasks of multi-organ nuclei detection and signet ring cells detection in histological images. Experimental results show that networks applying similarity learning achieved superior performance on both tasks when compared to their counterparts. In particular, the proposed SRPN achieve state-of-the-art performance on the MoNuSeg benchmark for nuclei segmentation and detection while compared to previous methods, and on the signet ring cell detection benchmark when compared with baselines. The sourcecode is publicly available at: https://github.com/sigma10010/nuclei_cells_det.
* Accepted by Medical Image Analysis for publication
Anatomy-Guided Parallel Bottleneck Transformer Network for Automated Evaluation of Root Canal Therapy
May 02, 2021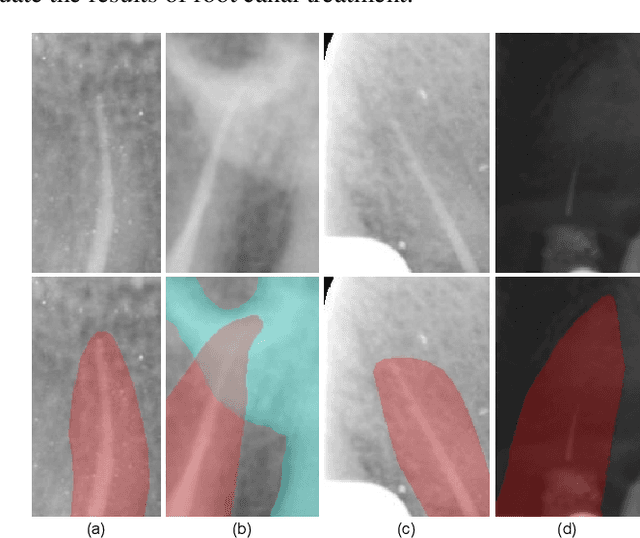
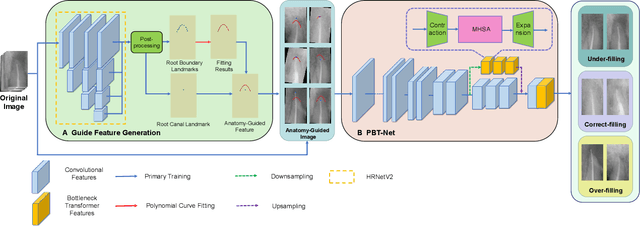
Abstract:Objective: Accurate evaluation of the root canal filling result in X-ray image is a significant step for the root canal therapy, which is based on the relative position between the apical area boundary of tooth root and the top of filled gutta-percha in root canal as well as the shape of the tooth root and so on to classify the result as correct-filling, under-filling or over-filling. Methods: We propose a novel anatomy-guided Transformer diagnosis network. For obtaining accurate anatomy-guided features, a polynomial curve fitting segmentation is proposed to segment the fuzzy boundary. And a Parallel Bottleneck Transformer network (PBT-Net) is introduced as the classification network for the final evaluation. Results, and conclusion: Our numerical experiments show that our anatomy-guided PBT-Net improves the accuracy from 40\% to 85\% relative to the baseline classification network. Comparing with the SOTA segmentation network indicates that the ASD is significantly reduced by 30.3\% through our fitting segmentation. Significance: Polynomial curve fitting segmentation has a great segmentation effect for extremely fuzzy boundaries. The prior knowledge guided classification network is suitable for the evaluation of root canal therapy greatly. And the new proposed Parallel Bottleneck Transformer for realizing self-attention is general in design, facilitating a broad use in most backbone networks.
High-Resolution Segmentation of Tooth Root Fuzzy Edge Based on Polynomial Curve Fitting with Landmark Detection
Mar 07, 2021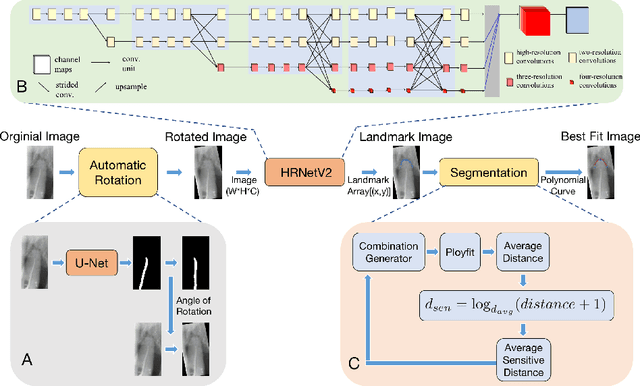
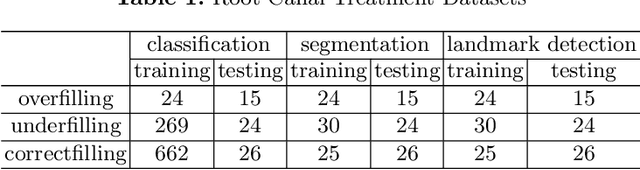
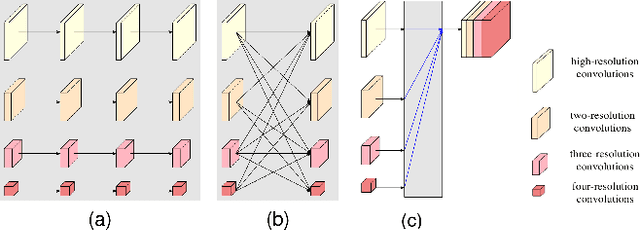
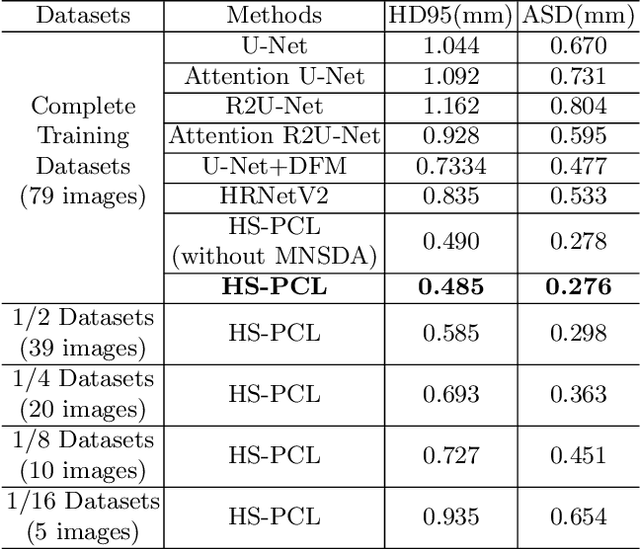
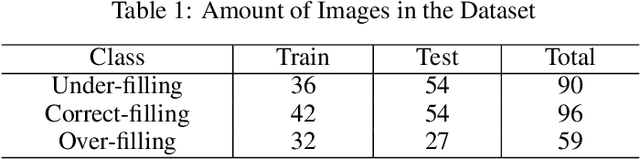
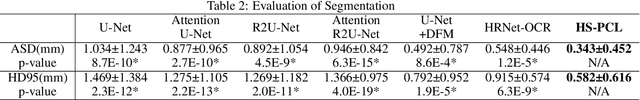
Abstract:As the most economical and routine auxiliary examination in the diagnosis of root canal treatment, oral X-ray has been widely used by stomatologists. It is still challenging to segment the tooth root with a blurry boundary for the traditional image segmentation method. To this end, we propose a model for high-resolution segmentation based on polynomial curve fitting with landmark detection (HS-PCL). It is based on detecting multiple landmarks evenly distributed on the edge of the tooth root to fit a smooth polynomial curve as the segmentation of the tooth root, thereby solving the problem of fuzzy edge. In our model, a maximum number of the shortest distances algorithm (MNSDA) is proposed to automatically reduce the negative influence of the wrong landmarks which are detected incorrectly and deviate from the tooth root on the fitting result. Our numerical experiments demonstrate that the proposed approach not only reduces Hausdorff95 (HD95) by 33.9% and Average Surface Distance (ASD) by 42.1% compared with the state-of-the-art method, but it also achieves excellent results on the minute quantity of datasets, which greatly improves the feasibility of automatic root canal therapy evaluation by medical image computing.
Multimodal Gait Recognition for Neurodegenerative Diseases
Jan 07, 2021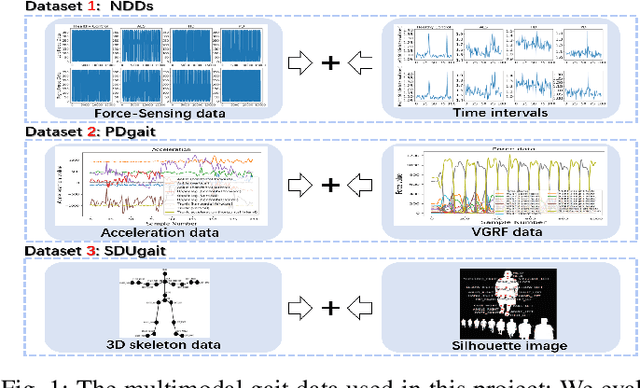
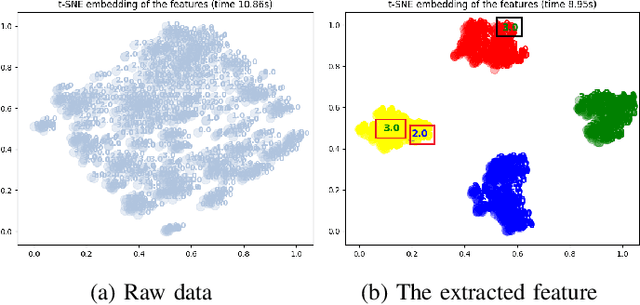
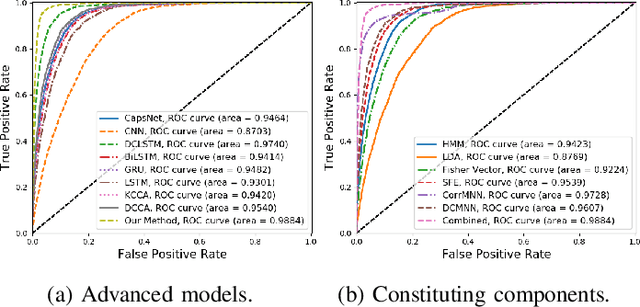
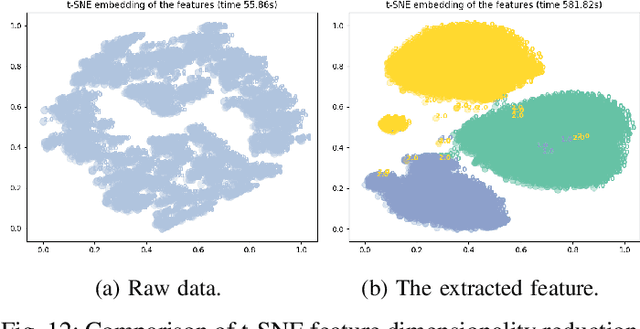
Abstract:In recent years, single modality based gait recognition has been extensively explored in the analysis of medical images or other sensory data, and it is recognised that each of the established approaches has different strengths and weaknesses. As an important motor symptom, gait disturbance is usually used for diagnosis and evaluation of diseases; moreover, the use of multi-modality analysis of the patient's walking pattern compensates for the one-sidedness of single modality gait recognition methods that only learn gait changes in a single measurement dimension. The fusion of multiple measurement resources has demonstrated promising performance in the identification of gait patterns associated with individual diseases. In this paper, as a useful tool, we propose a novel hybrid model to learn the gait differences between three neurodegenerative diseases, between patients with different severity levels of Parkinson's disease and between healthy individuals and patients, by fusing and aggregating data from multiple sensors. A spatial feature extractor (SFE) is applied to generating representative features of images or signals. In order to capture temporal information from the two modality data, a new correlative memory neural network (CorrMNN) architecture is designed for extracting temporal features. Afterwards, we embed a multi-switch discriminator to associate the observations with individual state estimations. Compared with several state-of-the-art techniques, our proposed framework shows more accurate classification results.
Perceptual underwater image enhancement with deep learning and physical priors
Sep 26, 2020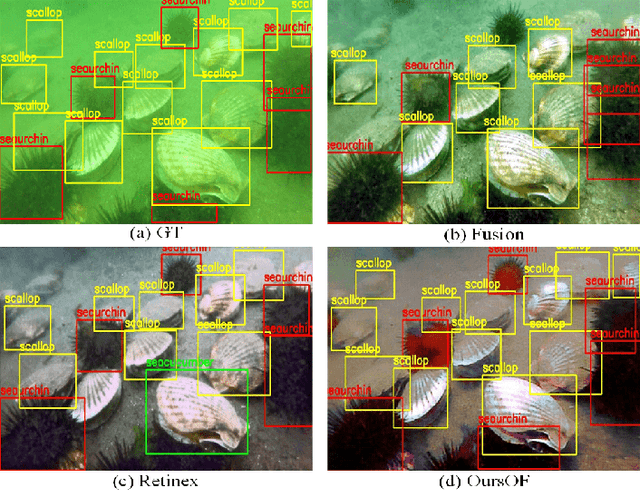
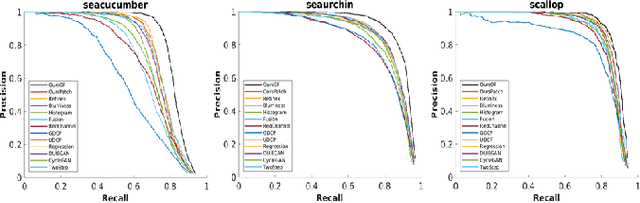
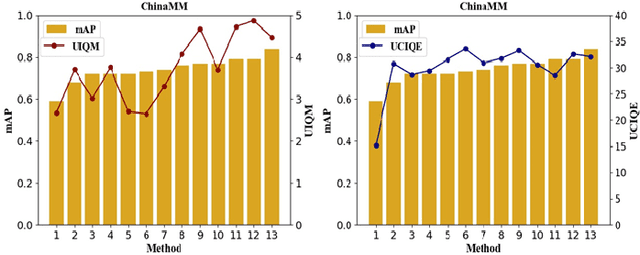
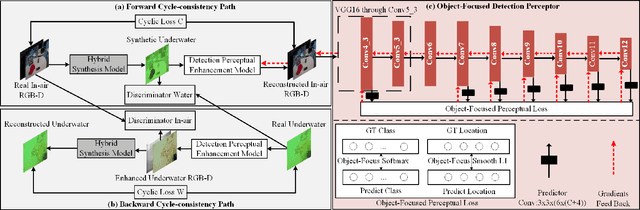
Abstract:Underwater image enhancement, as a pre-processing step to improve the accuracy of the following object detection task, has drawn considerable attention in the field of underwater navigation and ocean exploration. However, most of the existing underwater image enhancement strategies tend to consider enhancement and detection as two independent modules with no interaction, and the practice of separate optimization does not always help the underwater object detection task. In this paper, we propose two perceptual enhancement models, each of which uses a deep enhancement model with a detection perceptor. The detection perceptor provides coherent information in the form of gradients to the enhancement model, guiding the enhancement model to generate patch level visually pleasing images or detection favourable images. In addition, due to the lack of training data, a hybrid underwater image synthesis model, which fuses physical priors and data-driven cues, is proposed to synthesize training data and generalise our enhancement model for real-world underwater images. Experimental results show the superiority of our proposed method over several state-of-the-art methods on both real-world and synthetic underwater datasets.
Deep Learning Methods for Lung Cancer Segmentation in Whole-slide Histopathology Images -- the ACDC@LungHP Challenge 2019
Aug 21, 2020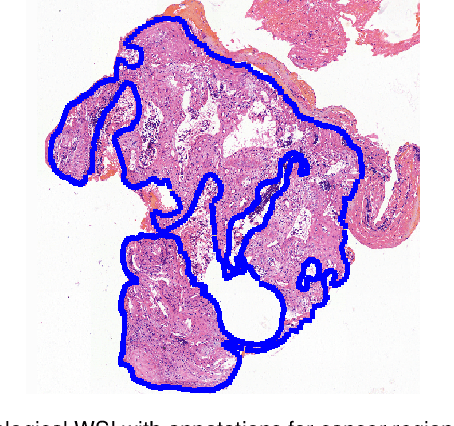
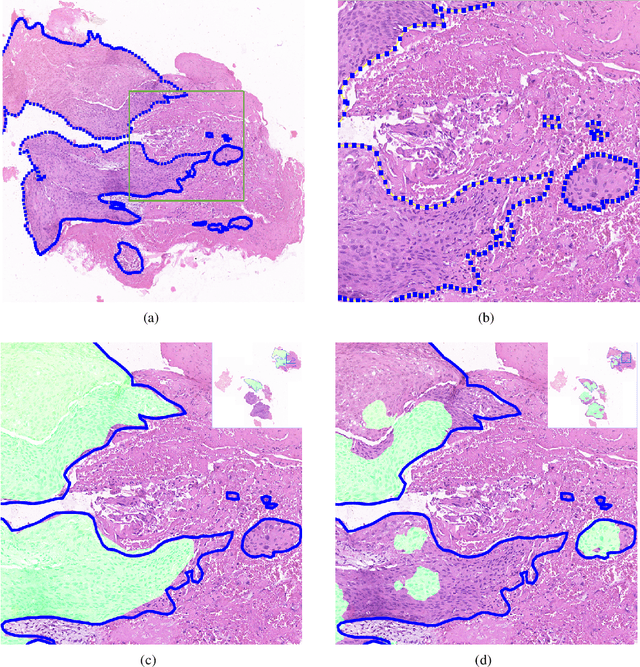
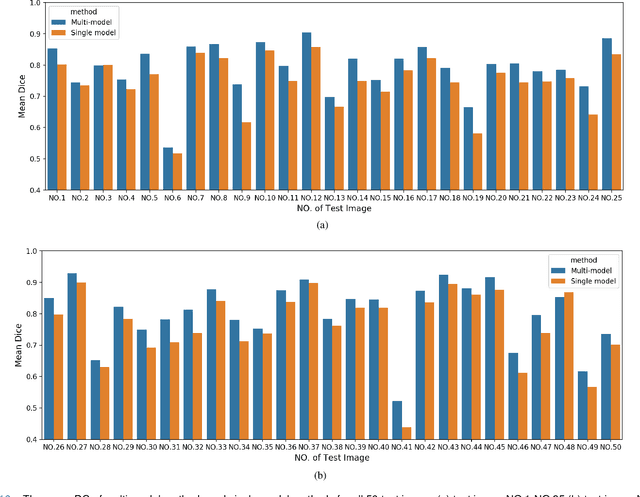
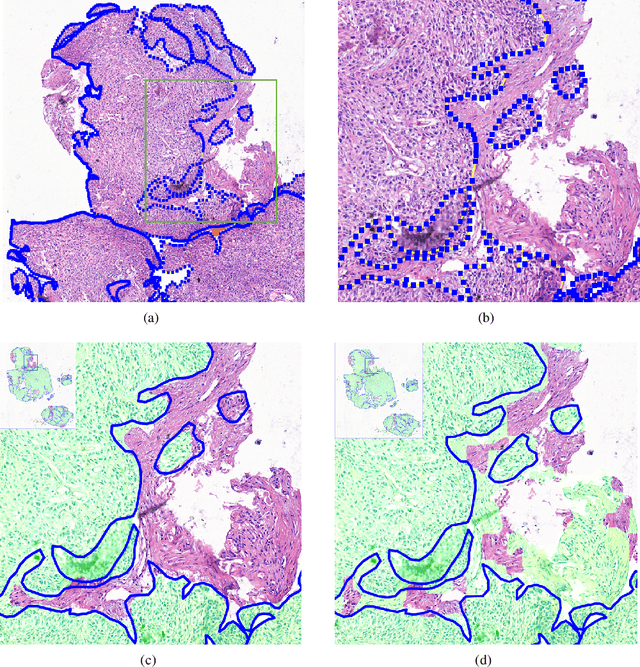

Abstract:Accurate segmentation of lung cancer in pathology slides is a critical step in improving patient care. We proposed the ACDC@LungHP (Automatic Cancer Detection and Classification in Whole-slide Lung Histopathology) challenge for evaluating different computer-aided diagnosis (CADs) methods on the automatic diagnosis of lung cancer. The ACDC@LungHP 2019 focused on segmentation (pixel-wise detection) of cancer tissue in whole slide imaging (WSI), using an annotated dataset of 150 training images and 50 test images from 200 patients. This paper reviews this challenge and summarizes the top 10 submitted methods for lung cancer segmentation. All methods were evaluated using the false positive rate, false negative rate, and DICE coefficient (DC). The DC ranged from 0.7354$\pm$0.1149 to 0.8372$\pm$0.0858. The DC of the best method was close to the inter-observer agreement (0.8398$\pm$0.0890). All methods were based on deep learning and categorized into two groups: multi-model method and single model method. In general, multi-model methods were significantly better ($\textit{p}$<$0.01$) than single model methods, with mean DC of 0.7966 and 0.7544, respectively. Deep learning based methods could potentially help pathologists find suspicious regions for further analysis of lung cancer in WSI.
Deep Learning Based Brain Tumor Segmentation: A Survey
Jul 21, 2020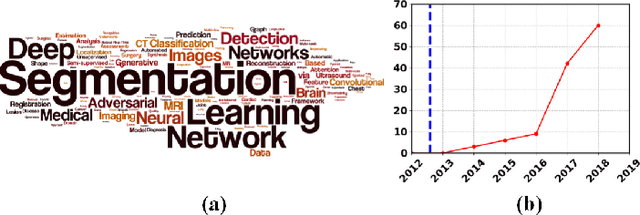
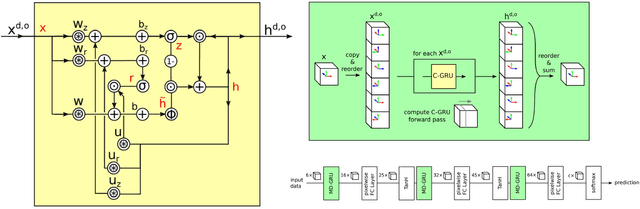
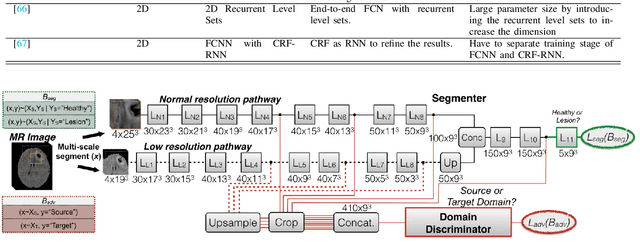
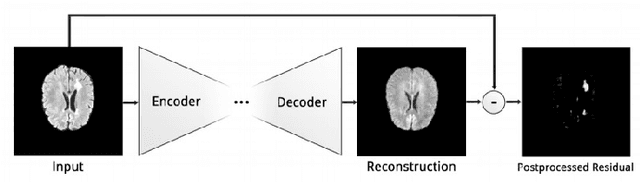
Abstract:Brain tumor segmentation is a challenging problem in medical image analysis. The goal of brain tumor segmentation is to generate accurate delineation of brain tumor regions with correctly located masks. In recent years, deep learning methods have shown very promising performance in solving various computer vision problems, such as image classification, object detection and semantic segmentation. A number of deep learning based methods have been applied to brain tumor segmentation and achieved impressive system performance. Considering state-of-the-art technologies and their performance, the purpose of this paper is to provide a comprehensive survey of recently developed deep learning based brain tumor segmentation techniques. The established works included in this survey extensively cover technical aspects such as the strengths and weaknesses of different approaches, pre- and post-processing frameworks, datasets and evaluation metrics. Finally, we conclude this survey by discussing the potential development in future research work.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge