Ling Zhang
Missouri S&T EMC Laboratory, Rolla, MO, USA
Towards a Single Unified Model for Effective Detection, Segmentation, and Diagnosis of Eight Major Cancers Using a Large Collection of CT Scans
Jan 28, 2023



Abstract:Human readers or radiologists routinely perform full-body multi-organ multi-disease detection and diagnosis in clinical practice, while most medical AI systems are built to focus on single organs with a narrow list of a few diseases. This might severely limit AI's clinical adoption. A certain number of AI models need to be assembled non-trivially to match the diagnostic process of a human reading a CT scan. In this paper, we construct a Unified Tumor Transformer (UniT) model to detect (tumor existence and location) and diagnose (tumor characteristics) eight major cancer-prevalent organs in CT scans. UniT is a query-based Mask Transformer model with the output of multi-organ and multi-tumor semantic segmentation. We decouple the object queries into organ queries, detection queries and diagnosis queries, and further establish hierarchical relationships among the three groups. This clinically-inspired architecture effectively assists inter- and intra-organ representation learning of tumors and facilitates the resolution of these complex, anatomically related multi-organ cancer image reading tasks. UniT is trained end-to-end using a curated large-scale CT images of 10,042 patients including eight major types of cancers and occurring non-cancer tumors (all are pathology-confirmed with 3D tumor masks annotated by radiologists). On the test set of 631 patients, UniT has demonstrated strong performance under a set of clinically relevant evaluation metrics, substantially outperforming both multi-organ segmentation methods and an assembly of eight single-organ expert models in tumor detection, segmentation, and diagnosis. Such a unified multi-cancer image reading model (UniT) can significantly reduce the number of false positives produced by combined multi-system models. This moves one step closer towards a universal high-performance cancer screening tool.
An Electromagnetic-Information-Theory Based Model for Efficient Characterization of MIMO Systems in Complex Space
Jan 13, 2023



Abstract:It is the pursuit of a multiple-input-multiple-output (MIMO) system to approach and even break the limit of channel capacity. However, it is always a big challenge to efficiently characterize the MIMO systems in complex space and get better propagation performance than the conventional MIMO systems considering only free space, which is important for guiding the power and phase allocation of antenna units. In this manuscript, an Electromagnetic-Information-Theory (EMIT) based model is developed for efficient characterization of MIMO systems in complex space. The group-T-matrix-based multiple scattering fast algorithm, the mode-decomposition-based characterization method, and their joint theoretical framework in complex space are discussed. Firstly, key informatics parameters in free electromagnetic space based on a dyadic Green's function are derived. Next, a novel group-T-matrix-based multiple scattering fast algorithm is developed to describe a representative inhomogeneous electromagnetic space. All the analytical results are validated by simulations. In addition, the complete form of the EMIT-based model is proposed to derive the informatics parameters frequently used in electromagnetic propagation, through integrating the mode analysis method with the dyadic Green's function matrix. Finally, as a proof-or-concept, microwave anechoic chamber measurements of a cylindrical array is performed, demonstrating the effectiveness of the EMIT-based model. Meanwhile, a case of image transmission with limited power is presented to illustrate how to use this EMIT-based model to guide the power and phase allocation of antenna units for real MIMO applications.
* 13 pages, 14 figures
A deep local attention network for pre-operative lymph node metastasis prediction in pancreatic cancer via multiphase CT imaging
Jan 04, 2023



Abstract:Lymph node (LN) metastasis status is one of the most critical prognostic and cancer staging factors for patients with resectable pancreatic ductal adenocarcinoma (PDAC), or in general, for any types of solid malignant tumors. Preoperative prediction of LN metastasis from non-invasive CT imaging is highly desired, as it might be straightforwardly used to guide the following neoadjuvant treatment decision and surgical planning. Most studies only capture the tumor characteristics in CT imaging to implicitly infer LN metastasis and very few work exploit direct LN's CT imaging information. To the best of our knowledge, this is the first work to propose a fully-automated LN segmentation and identification network to directly facilitate the LN metastasis status prediction task. Nevertheless LN segmentation/detection is very challenging since LN can be easily confused with other hard negative anatomic structures (e.g., vessels) from radiological images. We explore the anatomical spatial context priors of pancreatic LN locations by generating a guiding attention map from related organs and vessels to assist segmentation and infer LN status. As such, LN segmentation is impelled to focus on regions that are anatomically adjacent or plausible with respect to the specific organs and vessels. The metastasized LN identification network is trained to classify the segmented LN instances into positives or negatives by reusing the segmentation network as a pre-trained backbone and padding a new classification head. More importantly, we develop a LN metastasis status prediction network that combines the patient-wise aggregation results of LN segmentation/identification and deep imaging features extracted from the tumor region. Extensive quantitative nested five-fold cross-validation is conducted on a discovery dataset of 749 patients with PDAC.
FedTADBench: Federated Time-Series Anomaly Detection Benchmark
Dec 19, 2022



Abstract:Time series anomaly detection strives to uncover potential abnormal behaviors and patterns from temporal data, and has fundamental significance in diverse application scenarios. Constructing an effective detection model usually requires adequate training data stored in a centralized manner, however, this requirement sometimes could not be satisfied in realistic scenarios. As a prevailing approach to address the above problem, federated learning has demonstrated its power to cooperate with the distributed data available while protecting the privacy of data providers. However, it is still unclear that how existing time series anomaly detection algorithms perform with decentralized data storage and privacy protection through federated learning. To study this, we conduct a federated time series anomaly detection benchmark, named FedTADBench, which involves five representative time series anomaly detection algorithms and four popular federated learning methods. We would like to answer the following questions: (1)How is the performance of time series anomaly detection algorithms when meeting federated learning? (2) Which federated learning method is the most appropriate one for time series anomaly detection? (3) How do federated time series anomaly detection approaches perform on different partitions of data in clients? Numbers of results as well as corresponding analysis are provided from extensive experiments with various settings. The source code of our benchmark is publicly available at https://github.com/fanxingliu2020/FedTADBench.
SnapshotNet: Self-supervised Feature Learning for Point Cloud Data Segmentation Using Minimal Labeled Data
Jan 13, 2022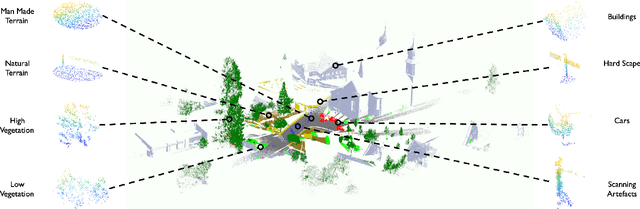
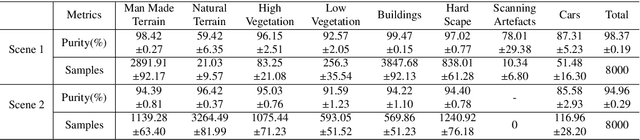
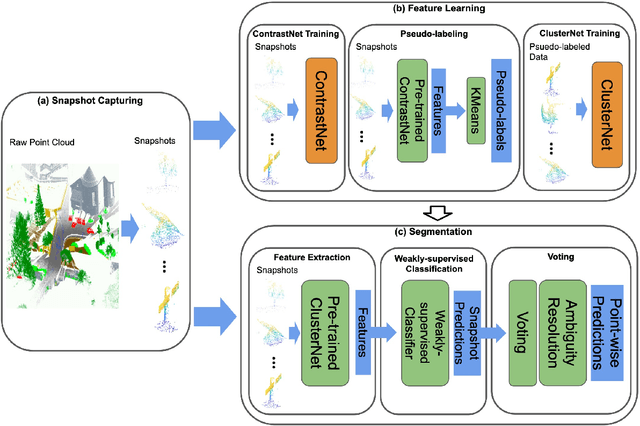
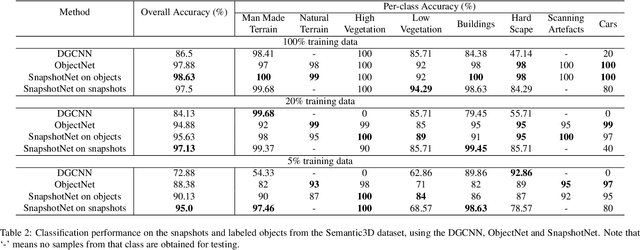
Abstract:Manually annotating complex scene point cloud datasets is both costly and error-prone. To reduce the reliance on labeled data, a new model called SnapshotNet is proposed as a self-supervised feature learning approach, which directly works on the unlabeled point cloud data of a complex 3D scene. The SnapshotNet pipeline includes three stages. In the snapshot capturing stage, snapshots, which are defined as local collections of points, are sampled from the point cloud scene. A snapshot could be a view of a local 3D scan directly captured from the real scene, or a virtual view of such from a large 3D point cloud dataset. Snapshots could also be sampled at different sampling rates or fields of view (FOVs), thus multi-FOV snapshots, to capture scale information from the scene. In the feature learning stage, a new pre-text task called multi-FOV contrasting is proposed to recognize whether two snapshots are from the same object or not, within the same FOV or across different FOVs. Snapshots go through two self-supervised learning steps: the contrastive learning step with both part and scale contrasting, followed by a snapshot clustering step to extract higher level semantic features. Then a weakly-supervised segmentation stage is implemented by first training a standard SVM classifier on the learned features with a small fraction of labeled snapshots. The trained SVM is used to predict labels for input snapshots and predicted labels are converted into point-wise label assignments for semantic segmentation of the entire scene using a voting procedure. The experiments are conducted on the Semantic3D dataset and the results have shown that the proposed method is capable of learning effective features from snapshots of complex scene data without any labels. Moreover, the proposed method has shown advantages when comparing to the SOA method on weakly-supervised point cloud semantic segmentation.
A free lunch from ViT:Adaptive Attention Multi-scale Fusion Transformer for Fine-grained Visual Recognition
Oct 11, 2021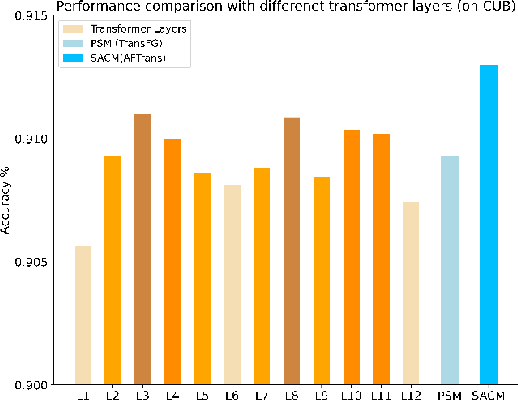
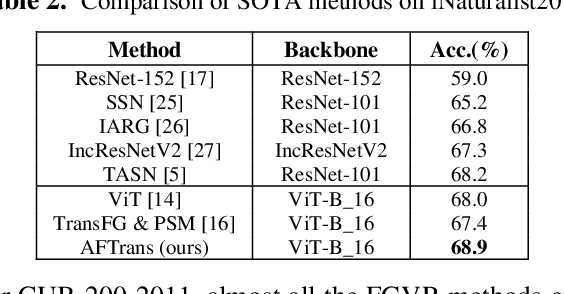
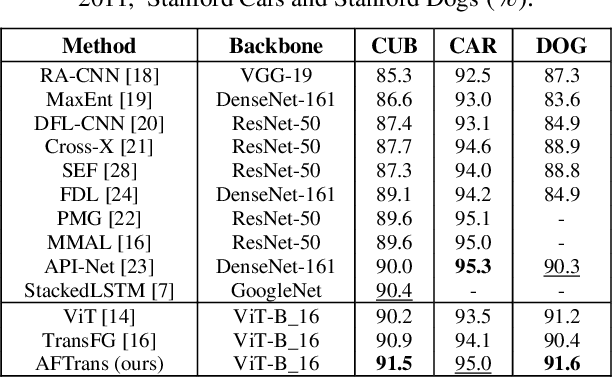
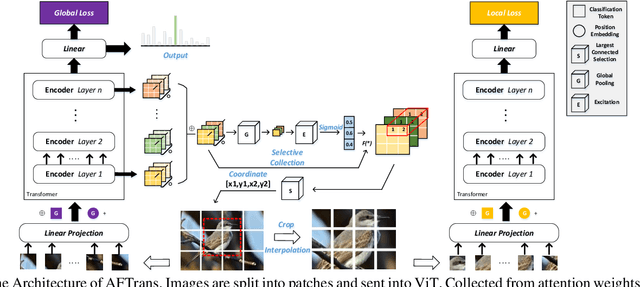

Abstract:Learning subtle representation about object parts plays a vital role in fine-grained visual recognition (FGVR) field. The vision transformer (ViT) achieves promising results on computer vision due to its attention mechanism. Nonetheless, with the fixed size of patches in ViT, the class token in deep layer focuses on the global receptive field and cannot generate multi-granularity features for FGVR. To capture region attention without box annotations and compensate for ViT shortcomings in FGVR, we propose a novel method named Adaptive attention multi-scale Fusion Transformer (AFTrans). The Selective Attention Collection Module (SACM) in our approach leverages attention weights in ViT and filters them adaptively to correspond with the relative importance of input patches. The multiple scales (global and local) pipeline is supervised by our weights sharing encoder and can be easily trained end-to-end. Comprehensive experiments demonstrate that AFTrans can achieve SOTA performance on three published fine-grained benchmarks: CUB-200-2011, Stanford Dogs and iNat2017.
Learning to Solve the AC Optimal Power Flow via a Lagrangian Approach
Oct 04, 2021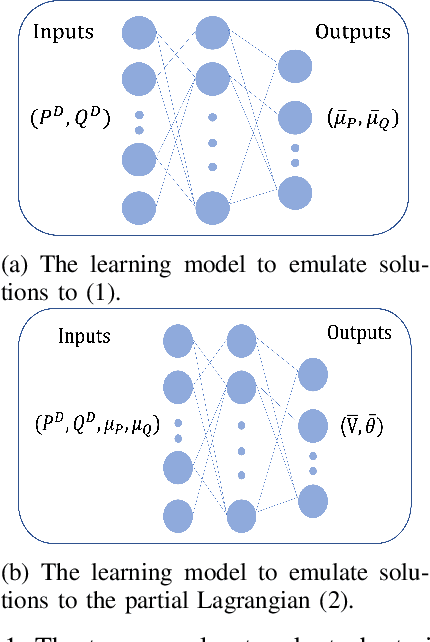
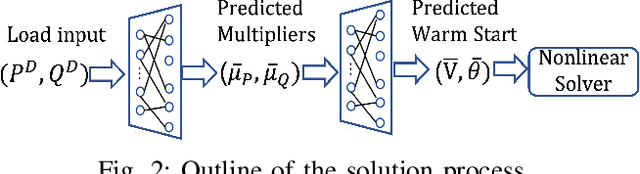
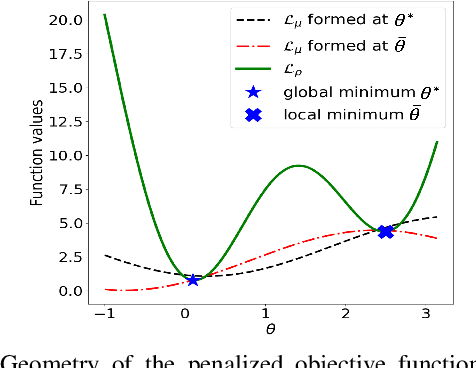
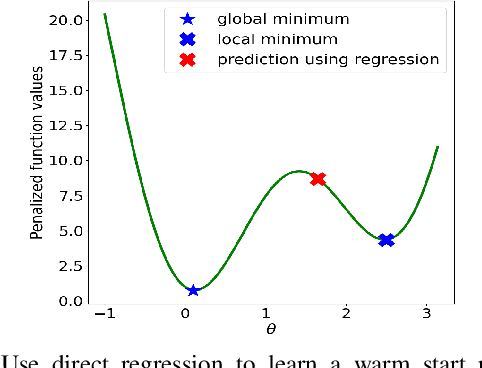
Abstract:Using deep neural networks to predict the solutions of AC optimal power flow (ACOPF) problems has been an active direction of research. However, because the ACOPF is nonconvex, it is difficult to construct a good data set that contains mostly globally optimal solutions. To overcome the challenge that the training data may contain suboptimal solutions, we propose a Lagrangian-based approach. First, we use a neural network to learn the dual variables of the ACOPF problem. Then we use a second neural network to predict solutions of the partial Lagrangian from the predicted dual variables. Since the partial Lagrangian has a much better optimization landscape, we use the predicted solutions from the neural network as a warm start for the ACOPF problem. Using standard and modified IEEE 22-bus, 39-bus, and 118-bus networks, we show that our approach is able to obtain the globally optimal cost even when the training data is mostly comprised of suboptimal solutions.
CANet: A Context-Aware Network for Shadow Removal
Aug 23, 2021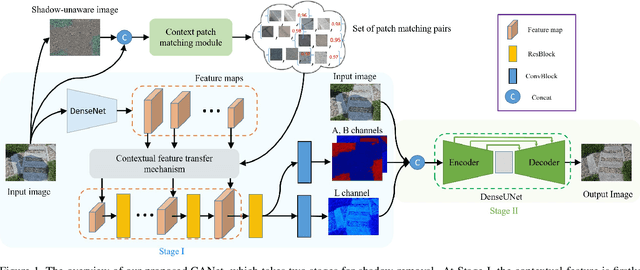
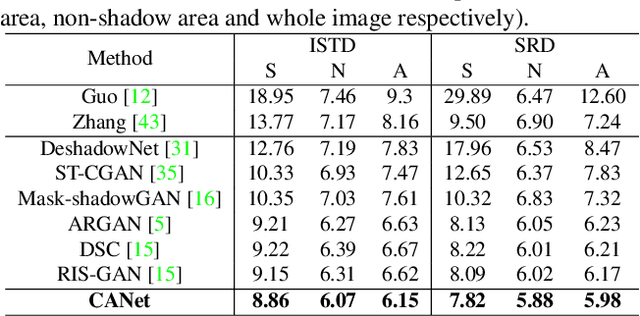
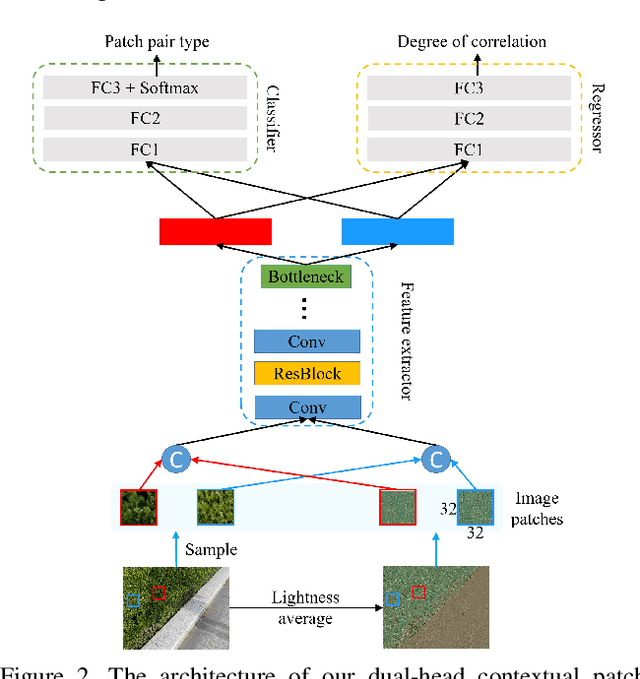
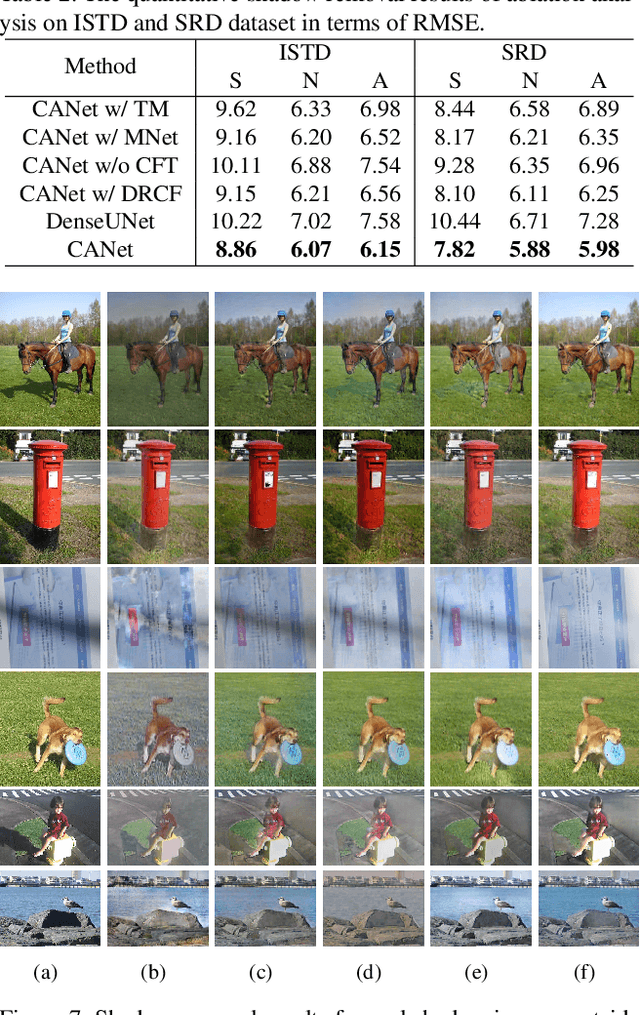
Abstract:In this paper, we propose a novel two-stage context-aware network named CANet for shadow removal, in which the contextual information from non-shadow regions is transferred to shadow regions at the embedded feature spaces. At Stage-I, we propose a contextual patch matching (CPM) module to generate a set of potential matching pairs of shadow and non-shadow patches. Combined with the potential contextual relationships between shadow and non-shadow regions, our well-designed contextual feature transfer (CFT) mechanism can transfer contextual information from non-shadow to shadow regions at different scales. With the reconstructed feature maps, we remove shadows at L and A/B channels separately. At Stage-II, we use an encoder-decoder to refine current results and generate the final shadow removal results. We evaluate our proposed CANet on two benchmark datasets and some real-world shadow images with complex scenes. Extensive experimental results strongly demonstrate the efficacy of our proposed CANet and exhibit superior performance to state-of-the-arts.
Distilling Neuron Spike with High Temperature in Reinforcement Learning Agents
Aug 05, 2021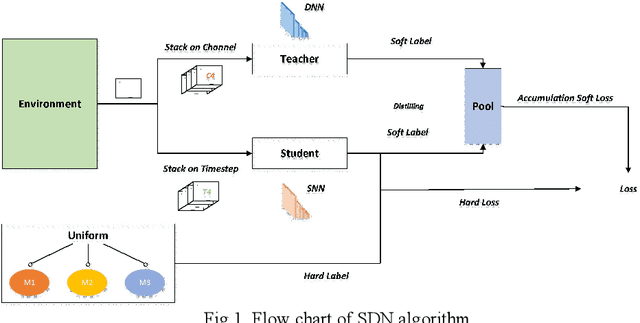
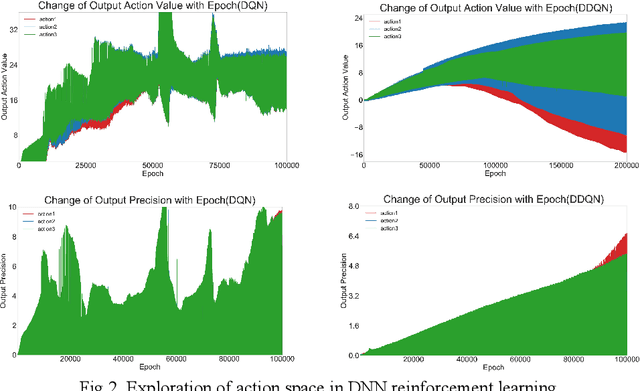
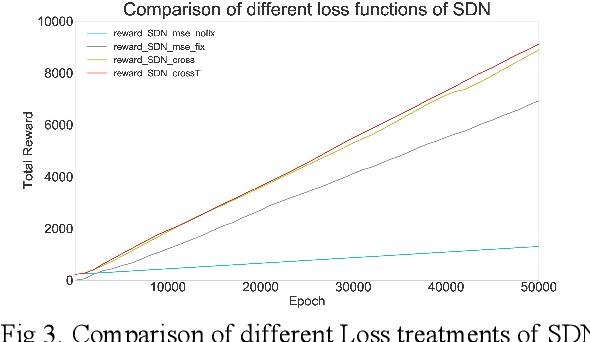
Abstract:Spiking neural network (SNN), compared with depth neural network (DNN), has faster processing speed, lower energy consumption and more biological interpretability, which is expected to approach Strong AI. Reinforcement learning is similar to learning in biology. It is of great significance to study the combination of SNN and RL. We propose the reinforcement learning method of spike distillation network (SDN) with STBP. This method uses distillation to effectively avoid the weakness of STBP, which can achieve SOTA performance in classification, and can obtain a smaller, faster convergence and lower power consumption SNN reinforcement learning model. Experiments show that our method can converge faster than traditional SNN reinforcement learning and DNN reinforcement learning methods, about 1000 epochs faster, and obtain SNN 200 times smaller than DNN. We also deploy SDN to the PKU nc64c chip, which proves that SDN has lower power consumption than DNN, and the power consumption of SDN is more than 600 times lower than DNN on large-scale devices. SDN provides a new way of SNN reinforcement learning, and can achieve SOTA performance, which proves the possibility of further development of SNN reinforcement learning.
Fast PDN Impedance Prediction Using Deep Learning
Jun 20, 2021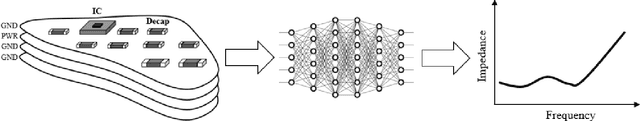
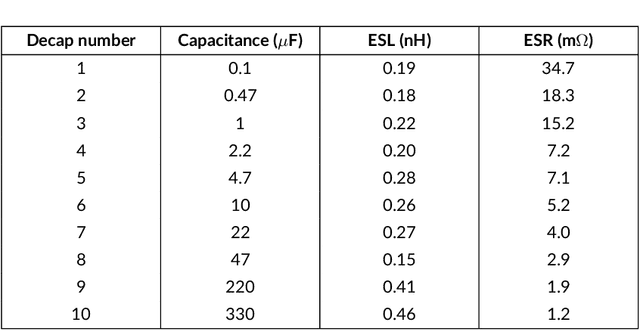
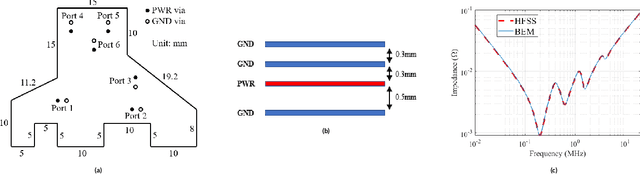

Abstract:Modeling and simulating a power distribution network (PDN) for printed circuit boards (PCBs) with irregular board shapes and multi-layer stackup is computationally inefficient using full-wave simulations. This paper presents a new concept of using deep learning for PDN impedance prediction. A boundary element method (BEM) is applied to efficiently calculate the impedance for arbitrary board shape and stackup. Then over one million boards with different shapes, stackup, IC location, and decap placement are randomly generated to train a deep neural network (DNN). The trained DNN can predict the impedance accurately for new board configurations that have not been used for training. The consumed time using the trained DNN is only 0.1 seconds, which is over 100 times faster than the BEM method and 5000 times faster than full-wave simulations.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge