Jun Zhang
National Innovation Institute of Defense Technology, Chinese Academy of Military Science
Bringing RGB and IR Together: Hierarchical Multi-Modal Enhancement for Robust Transmission Line Detection
Jan 25, 2025Abstract:Ensuring a stable power supply in rural areas relies heavily on effective inspection of power equipment, particularly transmission lines (TLs). However, detecting TLs from aerial imagery can be challenging when dealing with misalignments between visible light (RGB) and infrared (IR) images, as well as mismatched high- and low-level features in convolutional networks. To address these limitations, we propose a novel Hierarchical Multi-Modal Enhancement Network (HMMEN) that integrates RGB and IR data for robust and accurate TL detection. Our method introduces two key components: (1) a Mutual Multi-Modal Enhanced Block (MMEB), which fuses and enhances hierarchical RGB and IR feature maps in a coarse-to-fine manner, and (2) a Feature Alignment Block (FAB) that corrects misalignments between decoder outputs and IR feature maps by leveraging deformable convolutions. We employ MobileNet-based encoders for both RGB and IR inputs to accommodate edge-computing constraints and reduce computational overhead. Experimental results on diverse weather and lighting conditionsfog, night, snow, and daytimedemonstrate the superiority and robustness of our approach compared to state-of-the-art methods, resulting in fewer false positives, enhanced boundary delineation, and better overall detection performance. This framework thus shows promise for practical large-scale power line inspections with unmanned aerial vehicles.
Joint Automatic Speech Recognition And Structure Learning For Better Speech Understanding
Jan 13, 2025



Abstract:Spoken language understanding (SLU) is a structure prediction task in the field of speech. Recently, many works on SLU that treat it as a sequence-to-sequence task have achieved great success. However, This method is not suitable for simultaneous speech recognition and understanding. In this paper, we propose a joint speech recognition and structure learning framework (JSRSL), an end-to-end SLU model based on span, which can accurately transcribe speech and extract structured content simultaneously. We conduct experiments on name entity recognition and intent classification using the Chinese dataset AISHELL-NER and the English dataset SLURP. The results show that our proposed method not only outperforms the traditional sequence-to-sequence method in both transcription and extraction capabilities but also achieves state-of-the-art performance on the two datasets.
Dual Mutual Learning Network with Global-local Awareness for RGB-D Salient Object Detection
Jan 03, 2025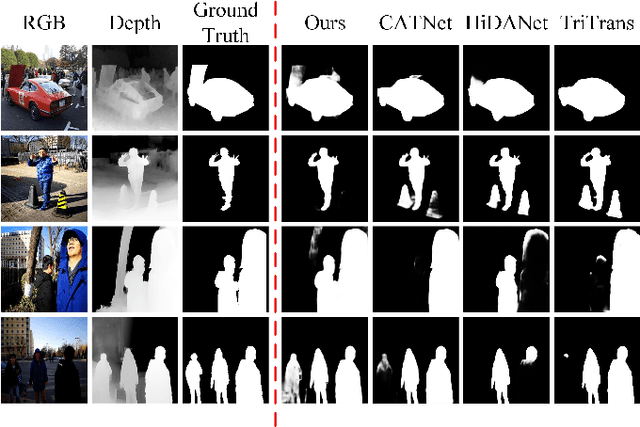
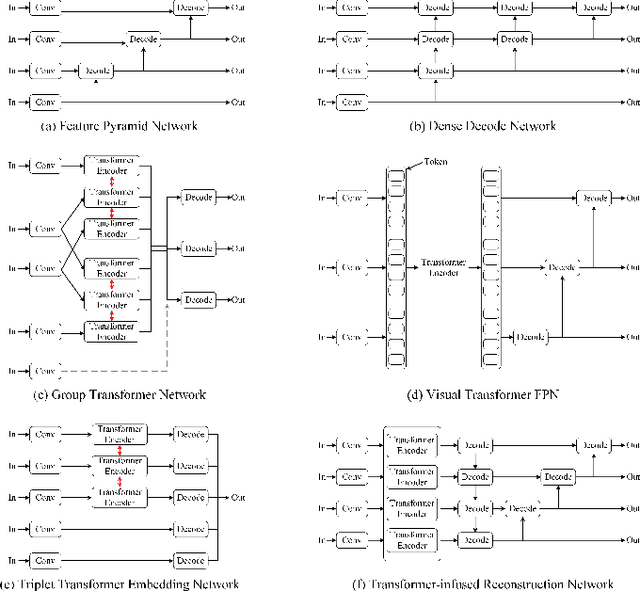
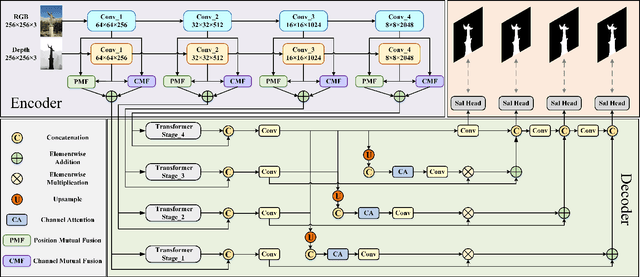
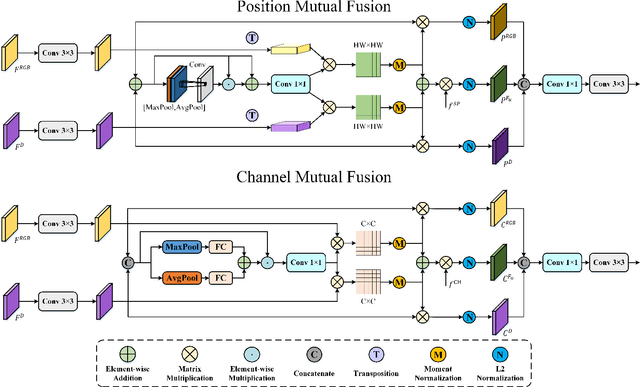
Abstract:RGB-D salient object detection (SOD), aiming to highlight prominent regions of a given scene by jointly modeling RGB and depth information, is one of the challenging pixel-level prediction tasks. Recently, the dual-attention mechanism has been devoted to this area due to its ability to strengthen the detection process. However, most existing methods directly fuse attentional cross-modality features under a manual-mandatory fusion paradigm without considering the inherent discrepancy between the RGB and depth, which may lead to a reduction in performance. Moreover, the long-range dependencies derived from global and local information make it difficult to leverage a unified efficient fusion strategy. Hence, in this paper, we propose the GL-DMNet, a novel dual mutual learning network with global-local awareness. Specifically, we present a position mutual fusion module and a channel mutual fusion module to exploit the interdependencies among different modalities in spatial and channel dimensions. Besides, we adopt an efficient decoder based on cascade transformer-infused reconstruction to integrate multi-level fusion features jointly. Extensive experiments on six benchmark datasets demonstrate that our proposed GL-DMNet performs better than 24 RGB-D SOD methods, achieving an average improvement of ~3% across four evaluation metrics compared to the second-best model (S3Net). Codes and results are available at https://github.com/kingkung2016/GL-DMNet.
Deep Learning Models for Colloidal Nanocrystal Synthesis
Dec 14, 2024



Abstract:Colloidal synthesis of nanocrystals usually includes complex chemical reactions and multi-step crystallization processes. Despite the great success in the past 30 years, it remains challenging to clarify the correlations between synthetic parameters of chemical reaction and physical properties of nanocrystals. Here, we developed a deep learning-based nanocrystal synthesis model that correlates synthetic parameters with the final size and shape of target nanocrystals, using a dataset of 3500 recipes covering 348 distinct nanocrystal compositions. The size and shape labels were obtained from transmission electron microscope images using a segmentation model trained with a semi-supervised algorithm on a dataset comprising 1.2 million nanocrystals. By applying the reaction intermediate-based data augmentation method and elaborated descriptors, the synthesis model was able to predict nanocrystal's size with a mean absolute error of 1.39 nm, while reaching an 89% average accuracy for shape classification. The synthesis model shows knowledge transfer capabilities across different nanocrystals with inputs of new recipes. With that, the influence of chemicals on the final size of nanocrystals was further evaluated, revealing the importance order of nanocrystal composition, precursor or ligand, and solvent. Overall, the deep learning-based nanocrystal synthesis model offers a powerful tool to expedite the development of high-quality nanocrystals.
AI and Deep Learning for THz Ultra-Massive MIMO: From Model-Driven Approaches to Foundation Models
Dec 13, 2024Abstract:In this paper, we explore the potential of artificial intelligence (AI) to address the challenges posed by terahertz ultra-massive multiple-input multiple-output (THz UM-MIMO) systems. We begin by outlining the characteristics of THz UM-MIMO systems, and identify three primary challenges for the transceiver design: 'hard to compute', 'hard to model', and 'hard to measure'. We argue that AI can provide a promising solution to these challenges. We then propose two systematic research roadmaps for developing AI algorithms tailored for THz UM-MIMO systems. The first roadmap, called model-driven deep learning (DL), emphasizes the importance to leverage available domain knowledge and advocates for adopting AI only to enhance the bottleneck modules within an established signal processing or optimization framework. We discuss four essential steps to make it work, including algorithmic frameworks, basis algorithms, loss function design, and neural architecture design. Afterwards, we present a forward-looking vision through the second roadmap, i.e., physical layer foundation models. This approach seeks to unify the design of different transceiver modules by focusing on their common foundation, i.e., the wireless channel. We propose to train a single, compact foundation model to estimate the score function of wireless channels, which can serve as a versatile prior for designing a wide variety of transceiver modules. We will also guide the readers through four essential steps, including general frameworks, conditioning, site-specific adaptation, and the joint design of foundation models and model-driven DL.
Learn How to Query from Unlabeled Data Streams in Federated Learning
Dec 12, 2024



Abstract:Federated learning (FL) enables collaborative learning among decentralized clients while safeguarding the privacy of their local data. Existing studies on FL typically assume offline labeled data available at each client when the training starts. Nevertheless, the training data in practice often arrive at clients in a streaming fashion without ground-truth labels. Given the expensive annotation cost, it is critical to identify a subset of informative samples for labeling on clients. However, selecting samples locally while accommodating the global training objective presents a challenge unique to FL. In this work, we tackle this conundrum by framing the data querying process in FL as a collaborative decentralized decision-making problem and proposing an effective solution named LeaDQ, which leverages multi-agent reinforcement learning algorithms. In particular, under the implicit guidance from global information, LeaDQ effectively learns the local policies for distributed clients and steers them towards selecting samples that can enhance the global model's accuracy. Extensive simulations on image and text tasks show that LeaDQ advances the model performance in various FL scenarios, outperforming the benchmarking algorithms.
From Principles to Practice: A Deep Dive into AI Ethics and Regulations
Dec 06, 2024



Abstract:In the rapidly evolving domain of Artificial Intelligence (AI), the complex interaction between innovation and regulation has become an emerging focus of our society. Despite tremendous advancements in AI's capabilities to excel in specific tasks and contribute to diverse sectors, establishing a high degree of trust in AI-generated outputs and decisions necessitates meticulous caution and continuous oversight. A broad spectrum of stakeholders, including governmental bodies, private sector corporations, academic institutions, and individuals, have launched significant initiatives. These efforts include developing ethical guidelines for AI and engaging in vibrant discussions on AI ethics, both among AI practitioners and within the broader society. This article thoroughly analyzes the ground-breaking AI regulatory framework proposed by the European Union. It delves into the fundamental ethical principles of safety, transparency, non-discrimination, traceability, and environmental sustainability for AI developments and deployments. Considering the technical efforts and strategies undertaken by academics and industry to uphold these principles, we explore the synergies and conflicts among the five ethical principles. Through this lens, work presents a forward-looking perspective on the future of AI regulations, advocating for a harmonized approach that safeguards societal values while encouraging technological advancement.
WRF-GS: Wireless Radiation Field Reconstruction with 3D Gaussian Splatting
Dec 06, 2024Abstract:Wireless channel modeling plays a pivotal role in designing, analyzing, and optimizing wireless communication systems. Nevertheless, developing an effective channel modeling approach has been a longstanding challenge. This issue has been escalated due to the denser network deployment, larger antenna arrays, and wider bandwidth in 5G and beyond networks. To address this challenge, we put forth WRF-GS, a novel framework for channel modeling based on wireless radiation field (WRF) reconstruction using 3D Gaussian splatting. WRF-GS employs 3D Gaussian primitives and neural networks to capture the interactions between the environment and radio signals, enabling efficient WRF reconstruction and visualization of the propagation characteristics. The reconstructed WRF can then be used to synthesize the spatial spectrum for comprehensive wireless channel characterization. Notably, with a small number of measurements, WRF-GS can synthesize new spatial spectra within milliseconds for a given scene, thereby enabling latency-sensitive applications. Experimental results demonstrate that WRF-GS outperforms existing methods for spatial spectrum synthesis, such as ray tracing and other deep-learning approaches. Moreover, WRF-GS achieves superior performance in the channel state information prediction task, surpassing existing methods by a significant margin of more than 2.43 dB.
p-MoD: Building Mixture-of-Depths MLLMs via Progressive Ratio Decay
Dec 05, 2024



Abstract:Despite the remarkable performance of multimodal large language models (MLLMs) across diverse tasks, the substantial training and inference costs impede their advancement. The majority of computation stems from the overwhelming volume of vision tokens processed by the transformer decoder. In this paper, we propose to build efficient MLLMs by leveraging the Mixture-of-Depths (MoD) mechanism, where each transformer decoder layer selects essential vision tokens to process while skipping redundant ones. However, integrating MoD into MLLMs is non-trivial. To address the challenges of training and inference stability as well as limited training data, we adapt the MoD module with two novel designs: tanh-gated weight normalization (TanhNorm) and symmetric token reweighting (STRing). Moreover, we observe that vision tokens exhibit higher redundancy in deeper layer and thus design a progressive ratio decay (PRD) strategy, which gradually reduces the token retention ratio layer by layer, employing a shifted cosine schedule. This crucial design fully unleashes the potential of MoD, significantly boosting the efficiency and performance of our models. To validate the effectiveness of our approach, we conduct extensive experiments with two baseline models across 14 benchmarks. Our model, p-MoD, matches or even surpasses the performance of the baseline models, with only 55.6% TFLOPs and 53.8% KV cache storage during inference, and 77.7% GPU hours during training.
Siamese Machine Unlearning with Knowledge Vaporization and Concentration
Dec 02, 2024



Abstract:In response to the practical demands of the ``right to be forgotten" and the removal of undesired data, machine unlearning emerges as an essential technique to remove the learned knowledge of a fraction of data points from trained models. However, existing methods suffer from limitations such as insufficient methodological support, high computational complexity, and significant memory demands. In this work, we propose the concepts of knowledge vaporization and concentration to selectively erase learned knowledge from specific data points while maintaining representations for the remaining data. Utilizing the Siamese networks, we exemplify the proposed concepts and develop an efficient method for machine unlearning. Our proposed Siamese unlearning method does not require additional memory overhead and full access to the remaining dataset. Extensive experiments conducted across multiple unlearning scenarios showcase the superiority of Siamese unlearning over baseline methods, illustrating its ability to effectively remove knowledge from forgetting data, enhance model utility on remaining data, and reduce susceptibility to membership inference attacks.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge