Jieping Ye
University of Michigan, DiDi Chuxing
On Reward-Free RL with Kernel and Neural Function Approximations: Single-Agent MDP and Markov Game
Oct 19, 2021Abstract:To achieve sample efficiency in reinforcement learning (RL), it necessitates efficiently exploring the underlying environment. Under the offline setting, addressing the exploration challenge lies in collecting an offline dataset with sufficient coverage. Motivated by such a challenge, we study the reward-free RL problem, where an agent aims to thoroughly explore the environment without any pre-specified reward function. Then, given any extrinsic reward, the agent computes the policy via a planning algorithm with offline data collected in the exploration phase. Moreover, we tackle this problem under the context of function approximation, leveraging powerful function approximators. Specifically, we propose to explore via an optimistic variant of the value-iteration algorithm incorporating kernel and neural function approximations, where we adopt the associated exploration bonus as the exploration reward. Moreover, we design exploration and planning algorithms for both single-agent MDPs and zero-sum Markov games and prove that our methods can achieve $\widetilde{\mathcal{O}}(1 /\varepsilon^2)$ sample complexity for generating a $\varepsilon$-suboptimal policy or $\varepsilon$-approximate Nash equilibrium when given an arbitrary extrinsic reward. To the best of our knowledge, we establish the first provably efficient reward-free RL algorithm with kernel and neural function approximators.
A Deep Value-network Based Approach for Multi-Driver Order Dispatching
Jun 08, 2021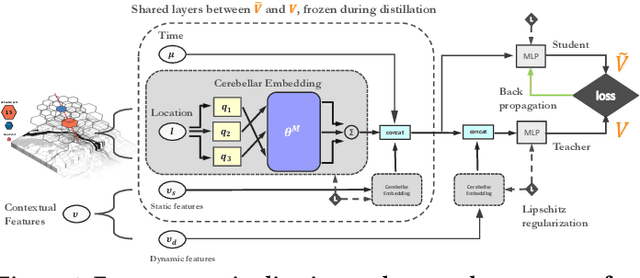
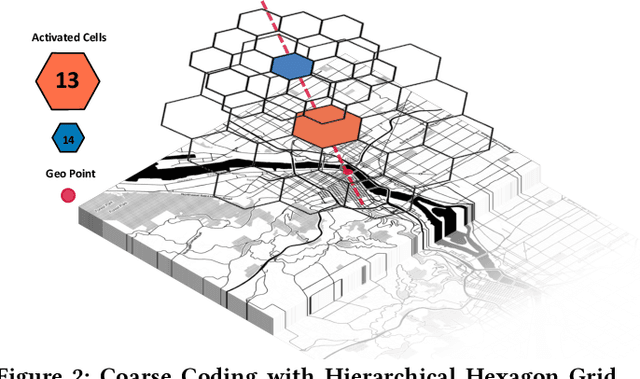
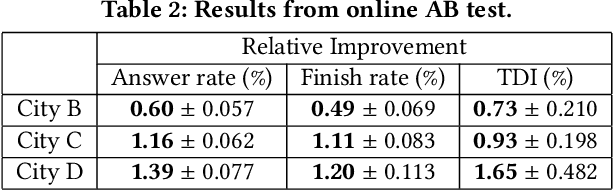
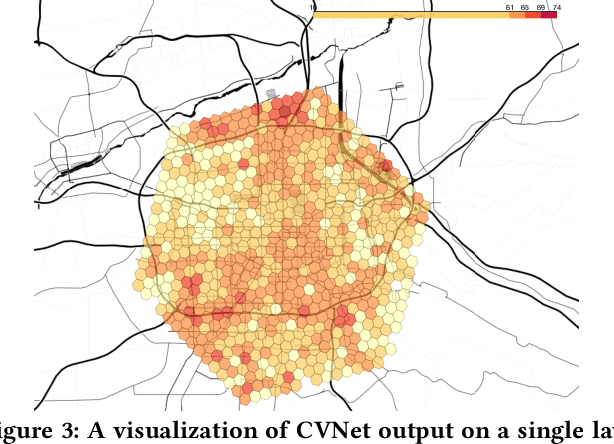
Abstract:Recent works on ride-sharing order dispatching have highlighted the importance of taking into account both the spatial and temporal dynamics in the dispatching process for improving the transportation system efficiency. At the same time, deep reinforcement learning has advanced to the point where it achieves superhuman performance in a number of fields. In this work, we propose a deep reinforcement learning based solution for order dispatching and we conduct large scale online A/B tests on DiDi's ride-dispatching platform to show that the proposed method achieves significant improvement on both total driver income and user experience related metrics. In particular, we model the ride dispatching problem as a Semi Markov Decision Process to account for the temporal aspect of the dispatching actions. To improve the stability of the value iteration with nonlinear function approximators like neural networks, we propose Cerebellar Value Networks (CVNet) with a novel distributed state representation layer. We further derive a regularized policy evaluation scheme for CVNet that penalizes large Lipschitz constant of the value network for additional robustness against adversarial perturbation and noises. Finally, we adapt various transfer learning methods to CVNet for increased learning adaptability and efficiency across multiple cities. We conduct extensive offline simulations based on real dispatching data as well as online AB tests through the DiDi's platform. Results show that CVNet consistently outperforms other recently proposed dispatching methods. We finally show that the performance can be further improved through the efficient use of transfer learning.
Value Function is All You Need: A Unified Learning Framework for Ride Hailing Platforms
Jun 04, 2021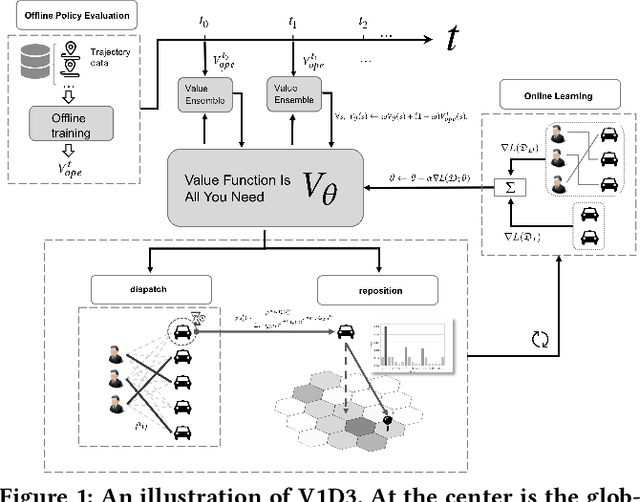
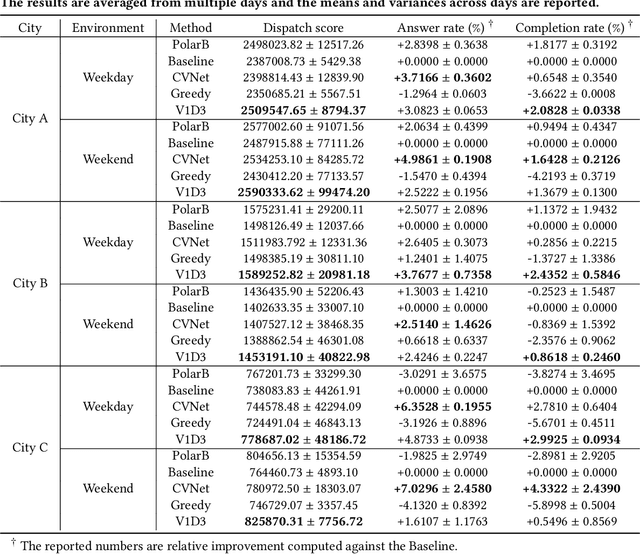
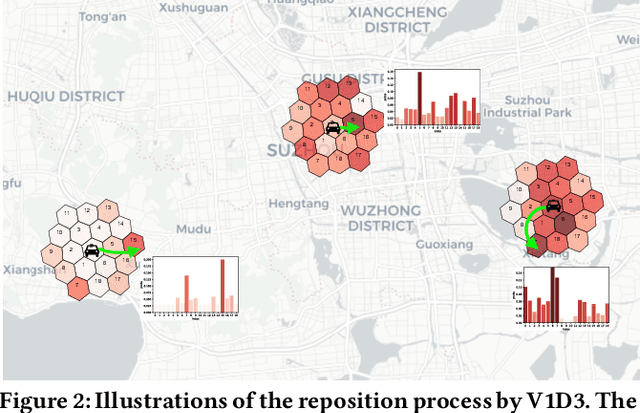
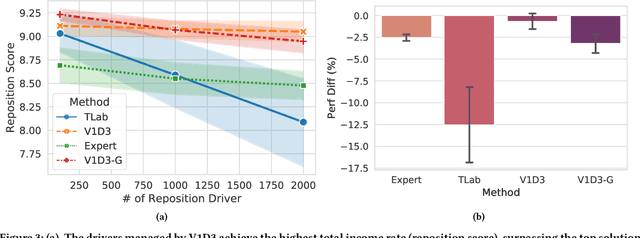
Abstract:Large ride-hailing platforms, such as DiDi, Uber and Lyft, connect tens of thousands of vehicles in a city to millions of ride demands throughout the day, providing great promises for improving transportation efficiency through the tasks of order dispatching and vehicle repositioning. Existing studies, however, usually consider the two tasks in simplified settings that hardly address the complex interactions between the two, the real-time fluctuations between supply and demand, and the necessary coordinations due to the large-scale nature of the problem. In this paper we propose a unified value-based dynamic learning framework (V1D3) for tackling both tasks. At the center of the framework is a globally shared value function that is updated continuously using online experiences generated from real-time platform transactions. To improve the sample-efficiency and the robustness, we further propose a novel periodic ensemble method combining the fast online learning with a large-scale offline training scheme that leverages the abundant historical driver trajectory data. This allows the proposed framework to adapt quickly to the highly dynamic environment, to generalize robustly to recurrent patterns and to drive implicit coordinations among the population of managed vehicles. Extensive experiments based on real-world datasets show considerably improvements over other recently proposed methods on both tasks. Particularly, V1D3 outperforms the first prize winners of both dispatching and repositioning tracks in the KDD Cup 2020 RL competition, achieving state-of-the-art results on improving both total driver income and user experience related metrics.
Reinforcement Learning for Ridesharing: A Survey
May 03, 2021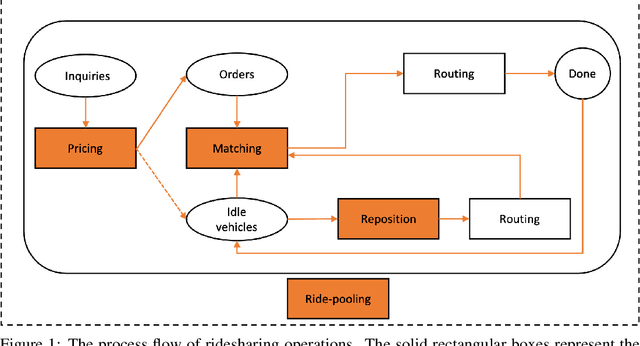
Abstract:In this paper, we present a comprehensive, in-depth survey of the literature on reinforcement learning approaches to ridesharing problems. Papers on the topics of rideshare matching, vehicle repositioning, ride-pooling, and dynamic pricing are covered. Popular data sets and open simulation environments are also introduced. Subsequently, we discuss a number of challenges and opportunities for reinforcement learning research on this important domain.
Real-world Ride-hailing Vehicle Repositioning using Deep Reinforcement Learning
Mar 08, 2021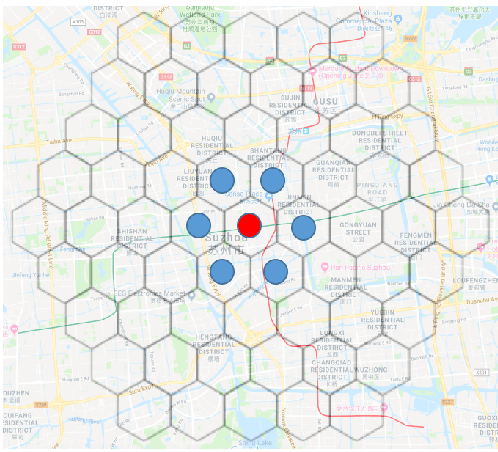
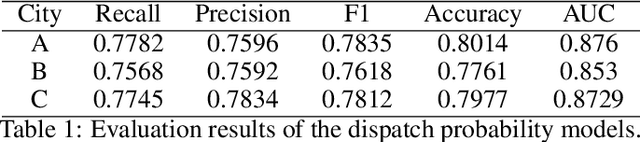
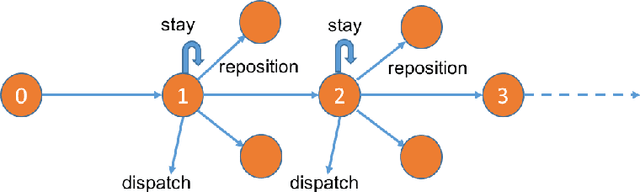

Abstract:We present a new practical framework based on deep reinforcement learning and decision-time planning for real-world vehicle repositioning on ride-hailing (a type of mobility-on-demand, MoD) platforms. Our approach learns the spatiotemporal state-value function using a batch training algorithm with deep value networks. The optimal repositioning action is generated on-demand through value-based policy search, which combines planning and bootstrapping with the value networks. For the large-fleet problems, we develop several algorithmic features that we incorporate into our framework and that we demonstrate to induce coordination among the algorithmically-guided vehicles. We benchmark our algorithm with baselines in a ride-hailing simulation environment to demonstrate its superiority in improving income efficiency meausred by income-per-hour. We have also designed and run a real-world experiment program with regular drivers on a major ride-hailing platform. We have observed significantly positive results on key metrics comparing our method with experienced drivers who performed idle-time repositioning based on their own expertise.
Selective Pseudo-Labeling with Reinforcement Learning for Semi-Supervised Domain Adaptation
Dec 07, 2020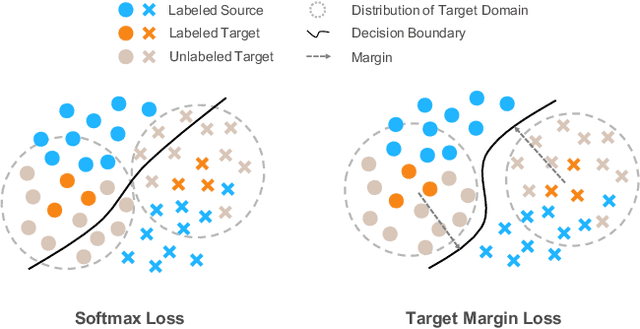
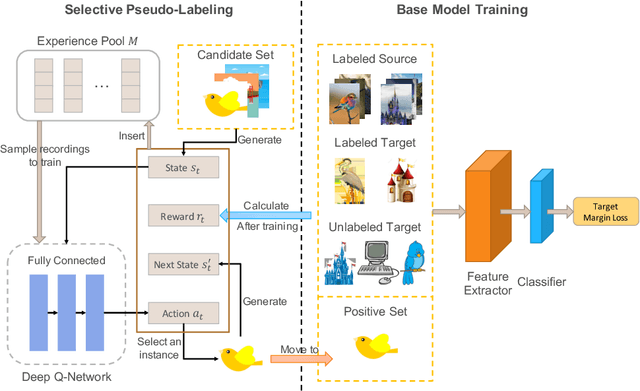
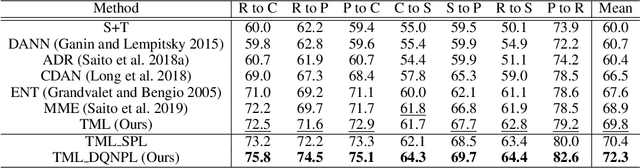
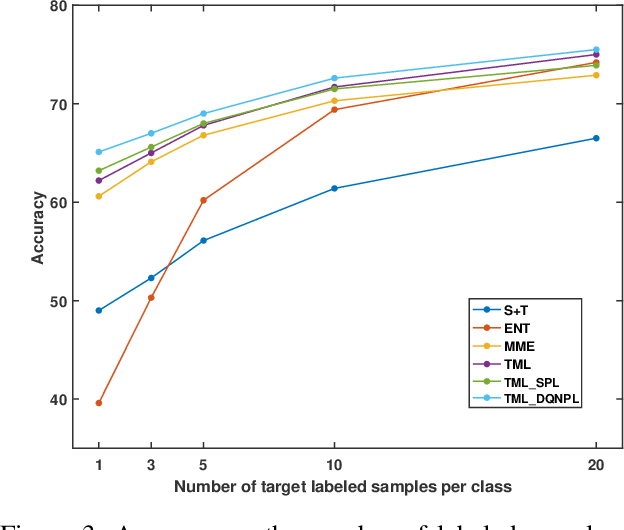
Abstract:Recent domain adaptation methods have demonstrated impressive improvement on unsupervised domain adaptation problems. However, in the semi-supervised domain adaptation (SSDA) setting where the target domain has a few labeled instances available, these methods can fail to improve performance. Inspired by the effectiveness of pseudo-labels in domain adaptation, we propose a reinforcement learning based selective pseudo-labeling method for semi-supervised domain adaptation. It is difficult for conventional pseudo-labeling methods to balance the correctness and representativeness of pseudo-labeled data. To address this limitation, we develop a deep Q-learning model to select both accurate and representative pseudo-labeled instances. Moreover, motivated by large margin loss's capacity on learning discriminative features with little data, we further propose a novel target margin loss for our base model training to improve its discriminability. Our proposed method is evaluated on several benchmark datasets for SSDA, and demonstrates superior performance to all the comparison methods.
Domain Adaptation with Incomplete Target Domains
Dec 03, 2020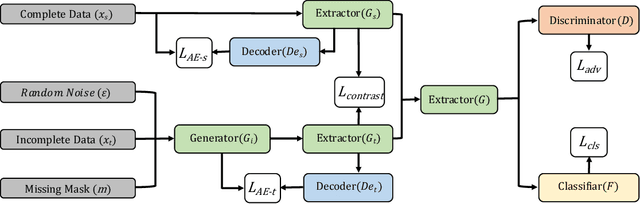
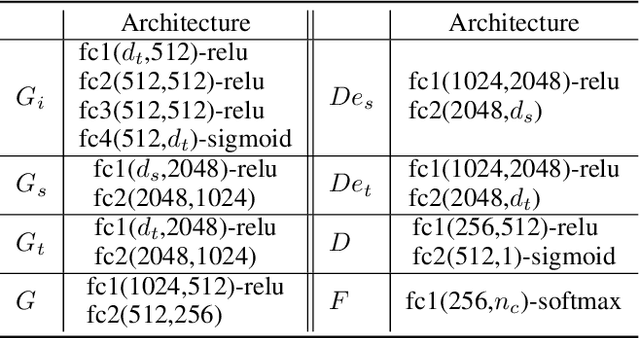
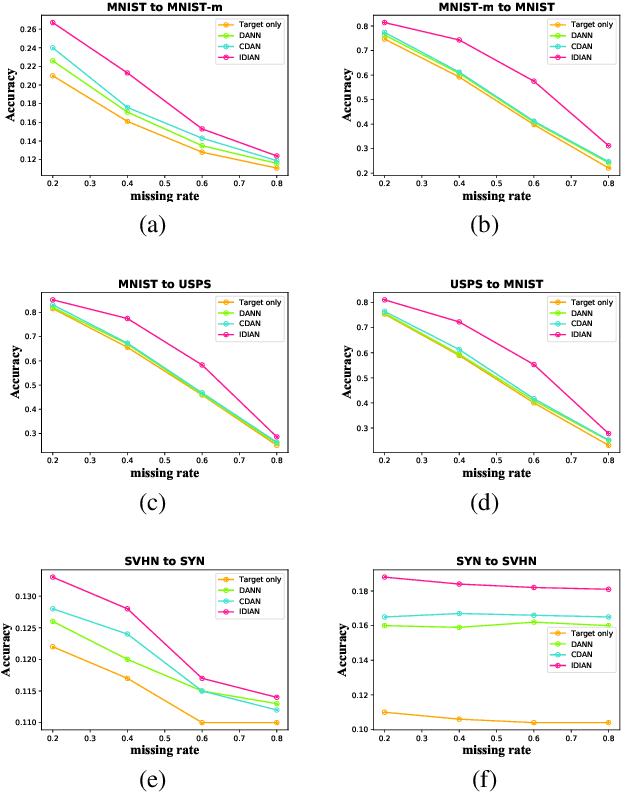
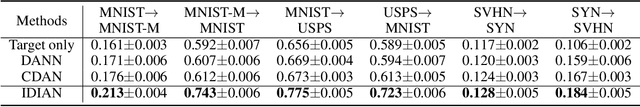
Abstract:Domain adaptation, as a task of reducing the annotation cost in a target domain by exploiting the existing labeled data in an auxiliary source domain, has received a lot of attention in the research community. However, the standard domain adaptation has assumed perfectly observed data in both domains, while in real world applications the existence of missing data can be prevalent. In this paper, we tackle a more challenging domain adaptation scenario where one has an incomplete target domain with partially observed data. We propose an Incomplete Data Imputation based Adversarial Network (IDIAN) model to address this new domain adaptation challenge. In the proposed model, we design a data imputation module to fill the missing feature values based on the partial observations in the target domain, while aligning the two domains via deep adversarial adaption. We conduct experiments on both cross-domain benchmark tasks and a real world adaptation task with imperfect target domains. The experimental results demonstrate the effectiveness of the proposed method.
Bi-Dimensional Feature Alignment for Cross-Domain Object Detection
Nov 14, 2020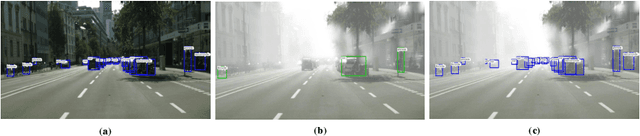
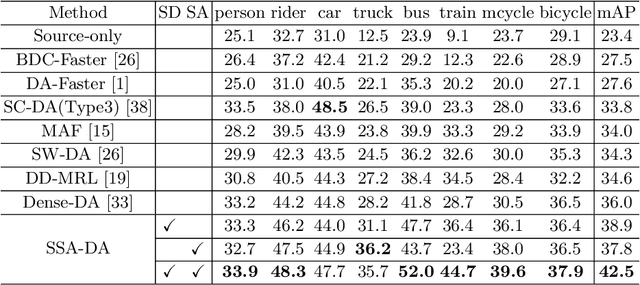
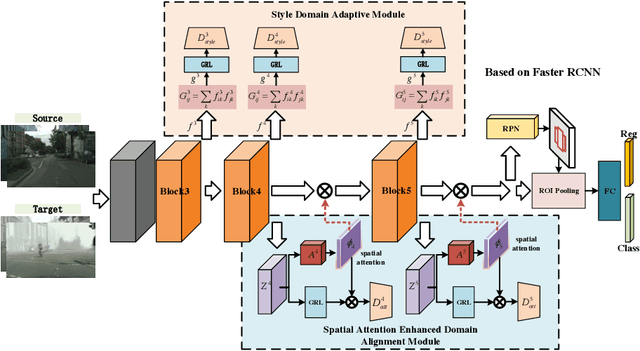
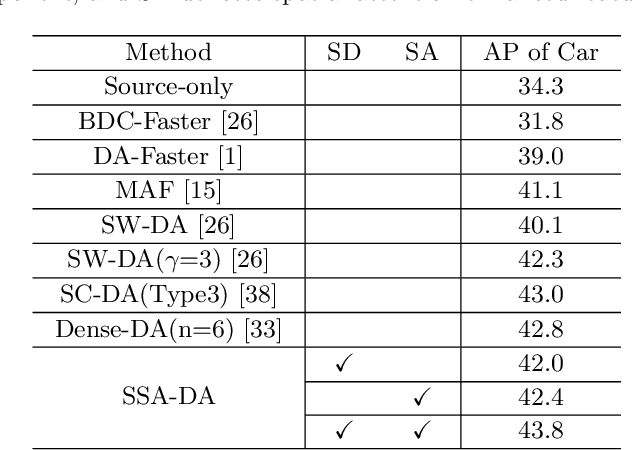
Abstract:Recently the problem of cross-domain object detection has started drawing attention in the computer vision community. In this paper, we propose a novel unsupervised cross-domain detection model that exploits the annotated data in a source domain to train an object detector for a different target domain. The proposed model mitigates the cross-domain representation divergence for object detection by performing cross-domain feature alignment in two dimensions, the depth dimension and the spatial dimension. In the depth dimension of channel layers, it uses inter-channel information to bridge the domain divergence with respect to image style alignment. In the dimension of spatial layers, it deploys spatial attention modules to enhance detection relevant regions and suppress irrelevant regions with respect to cross-domain feature alignment. Experiments are conducted on a number of benchmark cross-domain detection datasets. The empirical results show the proposed method outperforms the state-of-the-art comparison methods.
Joint predictions of multi-modal ride-hailing demands: a deep multi-task multigraph learning-based approach
Nov 11, 2020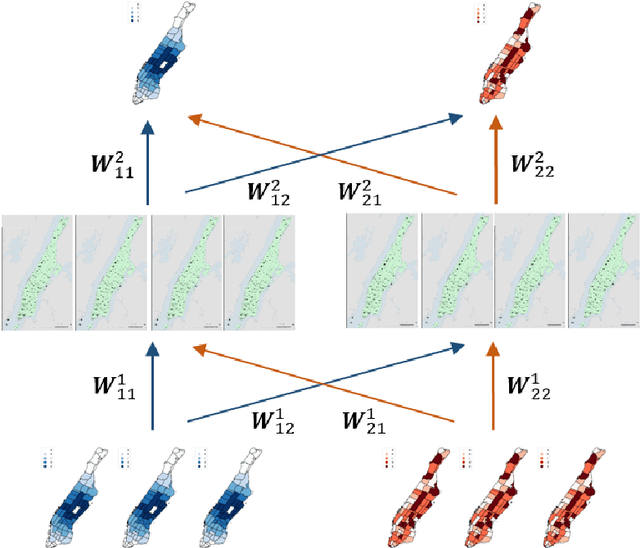
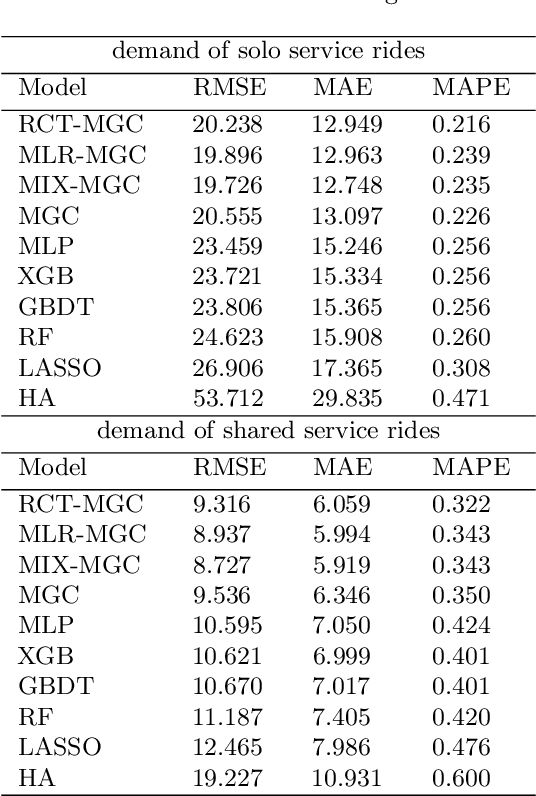
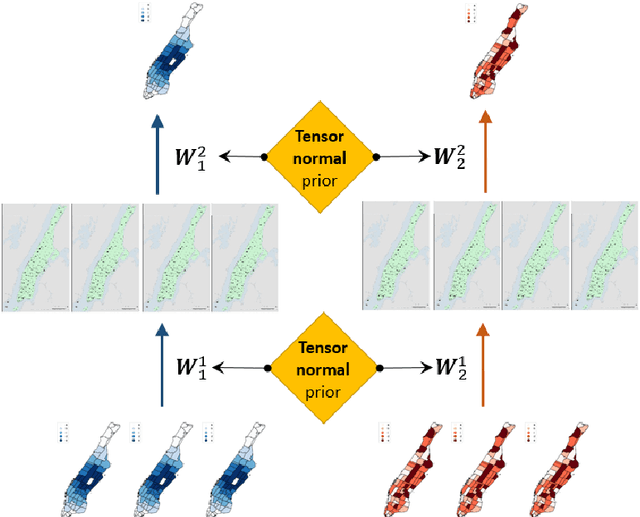
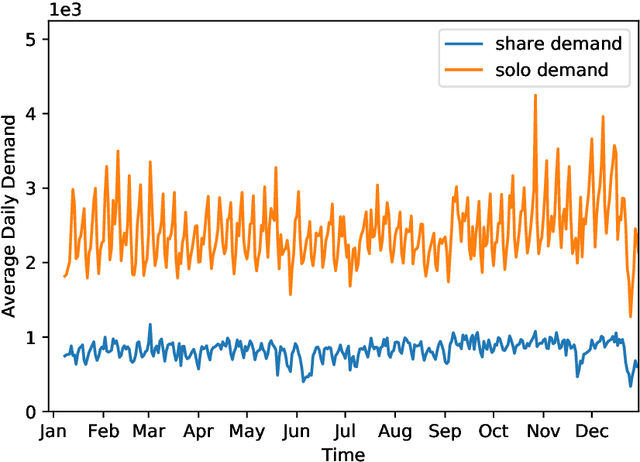
Abstract:Ride-hailing platforms generally provide various service options to customers, such as solo ride services, shared ride services, etc. It is generally expected that demands for different service modes are correlated, and the prediction of demand for one service mode can benefit from historical observations of demands for other service modes. Moreover, an accurate joint prediction of demands for multiple service modes can help the platforms better allocate and dispatch vehicle resources. Although there is a large stream of literature on ride-hailing demand predictions for one specific service mode, little efforts have been paid towards joint predictions of ride-hailing demands for multiple service modes. To address this issue, we propose a deep multi-task multi-graph learning approach, which combines two components: (1) multiple multi-graph convolutional (MGC) networks for predicting demands for different service modes, and (2) multi-task learning modules that enable knowledge sharing across multiple MGC networks. More specifically, two multi-task learning structures are established. The first one is the regularized cross-task learning, which builds cross-task connections among the inputs and outputs of multiple MGC networks. The second one is the multi-linear relationship learning, which imposes a prior tensor normal distribution on the weights of various MGC networks. Although there are no concrete bridges between different MGC networks, the weights of these networks are constrained by each other and subject to a common prior distribution. Evaluated with the for-hire-vehicle datasets in Manhattan, we show that our propose approach outperforms the benchmark algorithms in prediction accuracy for different ride-hailing modes.
Robust Unsupervised Video Anomaly Detection by Multi-Path Frame Prediction
Nov 05, 2020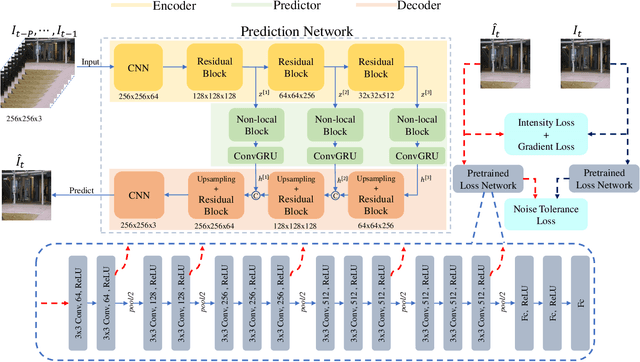
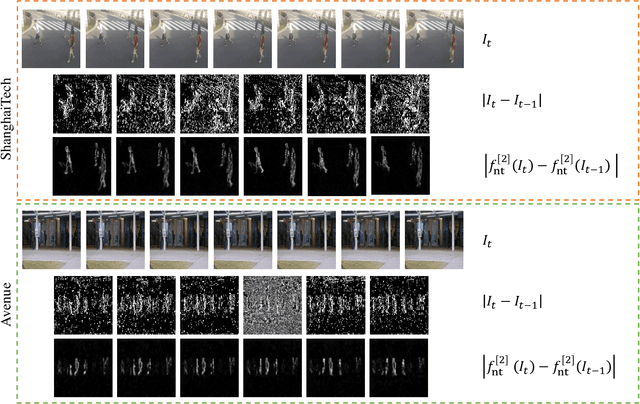

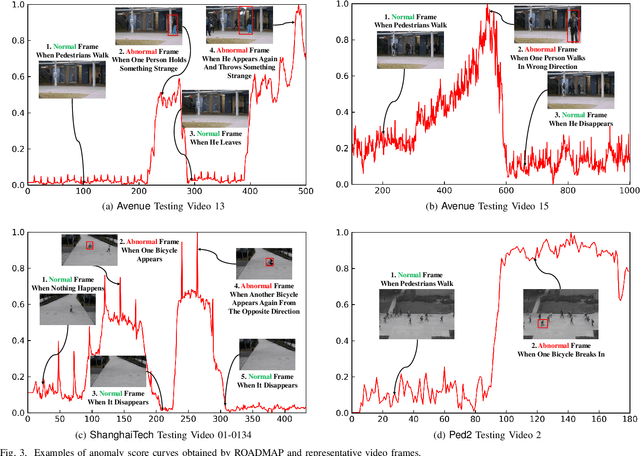
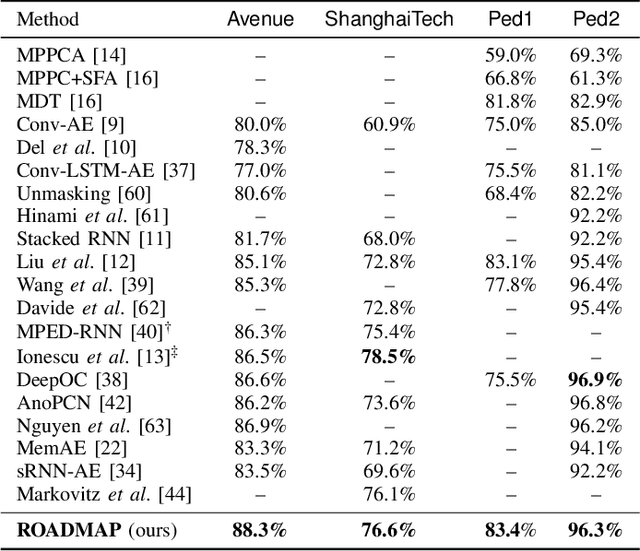
Abstract:Video anomaly detection is commonly used in many applications such as security surveillance and is very challenging. A majority of recent video anomaly detection approaches utilize deep reconstruction models, but their performance is often suboptimal because of insufficient reconstruction error differences between normal and abnormal video frames in practice. Meanwhile, frame prediction-based anomaly detection methods have shown promising performance. In this paper, we propose a novel and robust unsupervised video anomaly detection method by frame prediction with proper design which is more in line with the characteristics of surveillance videos. The proposed method is equipped with a multi-path ConvGRU-based frame prediction network that can better handle semantically informative objects and areas of different scales and capture spatial-temporal dependencies in normal videos. A noise tolerance loss is introduced during training to mitigate the interference caused by background noise. Extensive experiments have been conducted on the CUHK Avenue, ShanghaiTech Campus, and UCSD Pedestrian datasets, and the results show that our proposed method outperforms existing state-of-the-art approaches. Remarkably, our proposed method obtains the frame-level AUC score of 88.3% on the CUHK Avenue dataset.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge