Guotong Xie
Scalable Semi-supervised Landmark Localization for X-ray Images using Few-shot Deep Adaptive Graph
Apr 29, 2021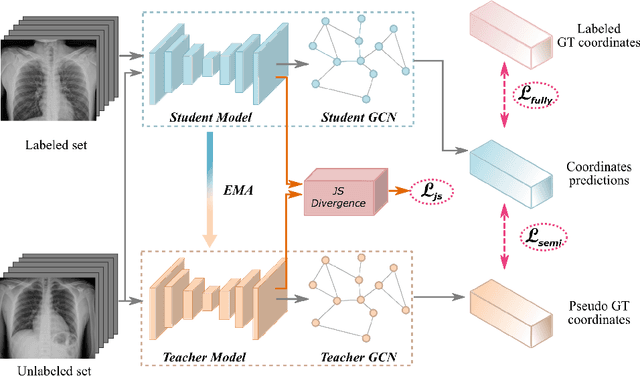
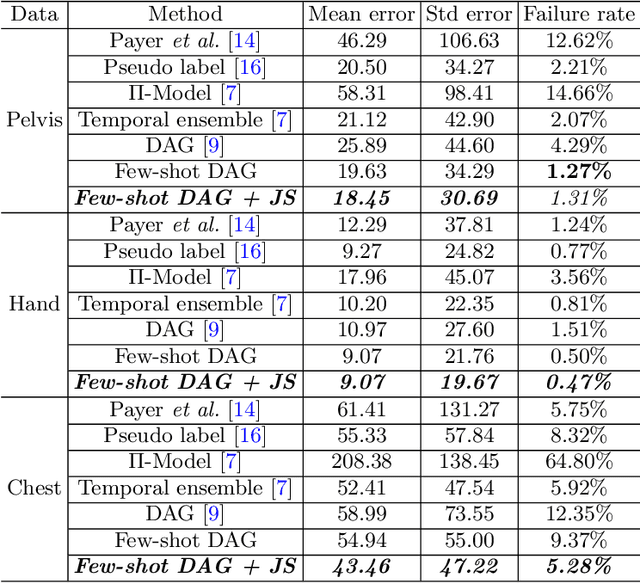
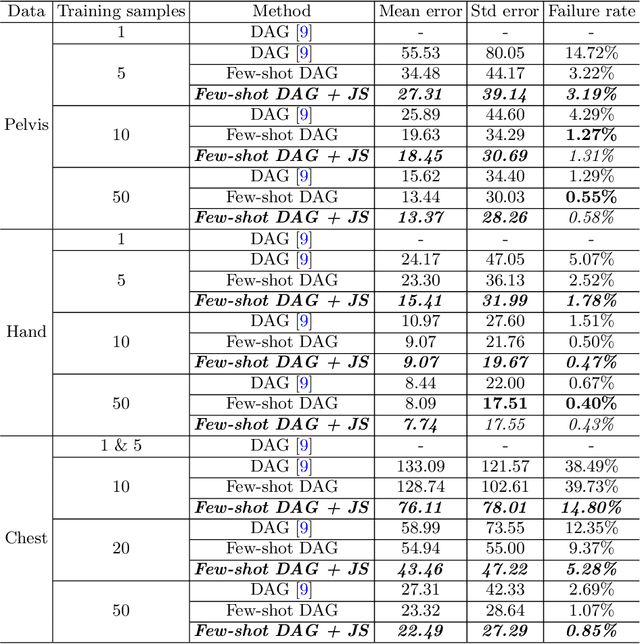
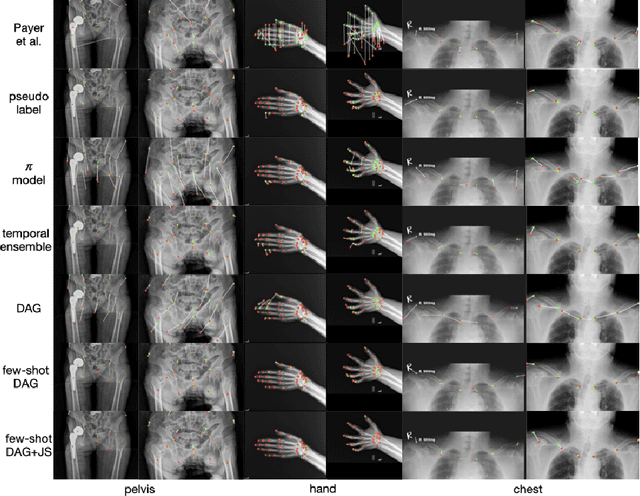
Abstract:Landmark localization plays an important role in medical image analysis. Learning based methods, including CNN and GCN, have demonstrated the state-of-the-art performance. However, most of these methods are fully-supervised and heavily rely on manual labeling of a large training dataset. In this paper, based on a fully-supervised graph-based method, DAG, we proposed a semi-supervised extension of it, termed few-shot DAG, \ie five-shot DAG. It first trains a DAG model on the labeled data and then fine-tunes the pre-trained model on the unlabeled data with a teacher-student SSL mechanism. In addition to the semi-supervised loss, we propose another loss using JS divergence to regulate the consistency of the intermediate feature maps. We extensively evaluated our method on pelvis, hand and chest landmark detection tasks. Our experiment results demonstrate consistent and significant improvements over previous methods.
Learning from Subjective Ratings Using Auto-Decoded Deep Latent Embeddings
Apr 16, 2021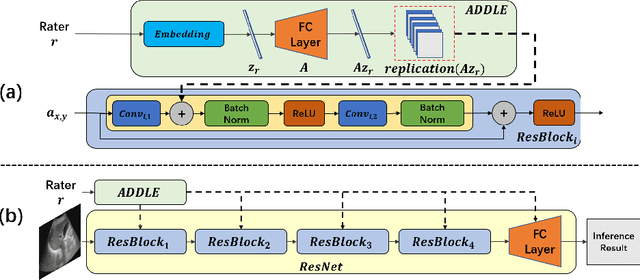
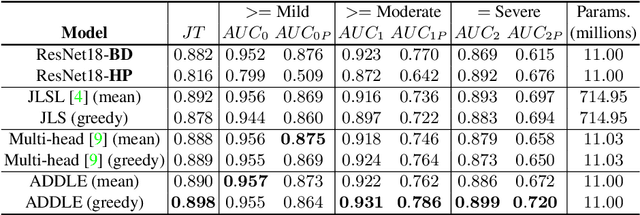
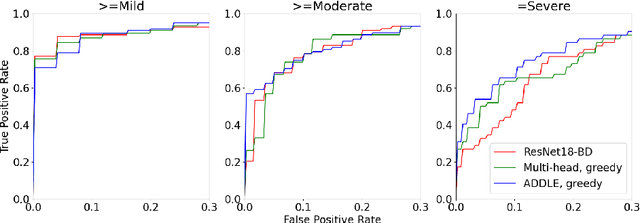
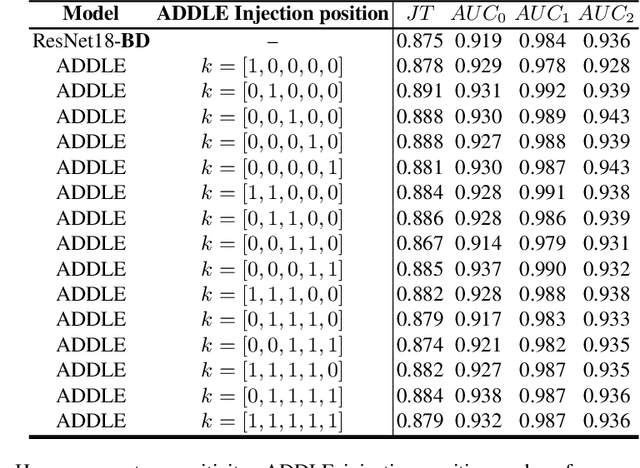
Abstract:Depending on the application, radiological diagnoses can be associated with high inter- and intra-rater variabilities. Most computer-aided diagnosis (CAD) solutions treat such data as incontrovertible, exposing learning algorithms to considerable and possibly contradictory label noise and biases. Thus, managing subjectivity in labels is a fundamental problem in medical imaging analysis. To address this challenge, we introduce auto-decoded deep latent embeddings (ADDLE), which explicitly models the tendencies of each rater using an auto-decoder framework. After a simple linear transformation, the latent variables can be injected into any backbone at any and multiple points, allowing the model to account for rater-specific effects on the diagnosis. Importantly, ADDLE does not expect multiple raters per image in training, meaning it can readily learn from data mined from hospital archives. Moreover, the complexity of training ADDLE does not increase as more raters are added. During inference each rater can be simulated and a 'mean' or 'greedy' virtual rating can be produced. We test ADDLE on the problem of liver steatosis diagnosis from 2D ultrasound (US) by collecting 46 084 studies along with clinical US diagnoses originating from 65 different raters. We evaluated diagnostic performance using a separate dataset with gold-standard biopsy diagnoses. ADDLE can improve the partial areas under the curve (AUCs) for diagnosing severe steatosis by 10.5% over standard classifiers while outperforming other annotator-noise approaches, including those requiring 65 times the parameters.
Semi-Supervised Learning for Bone Mineral Density Estimation in Hip X-ray Images
Mar 24, 2021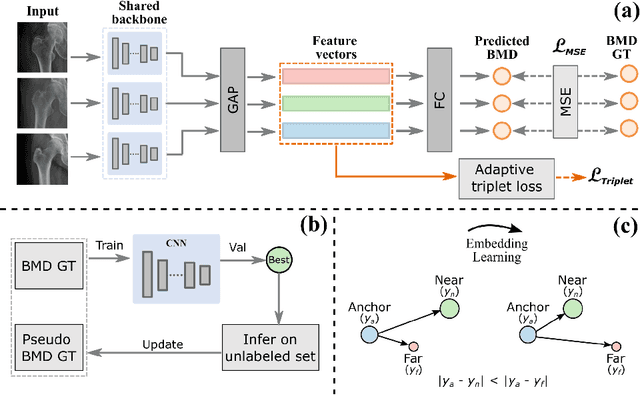

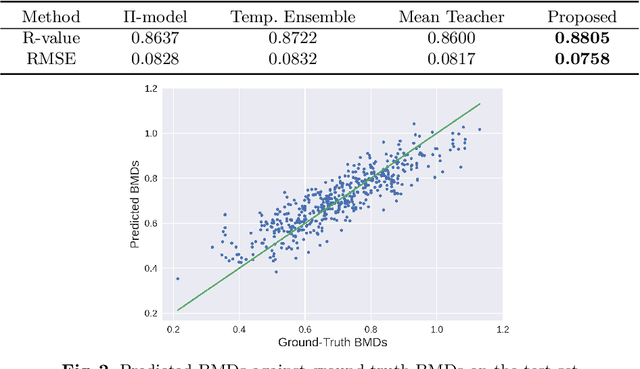
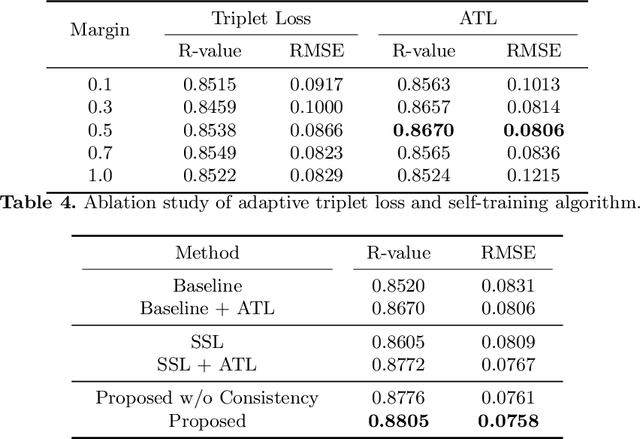
Abstract:Bone mineral density (BMD) is a clinically critical indicator of osteoporosis, usually measured by dual-energy X-ray absorptiometry (DEXA). Due to the limited accessibility of DEXA machines and examinations, osteoporosis is often under-diagnosed and under-treated, leading to increased fragility fracture risks. Thus it is highly desirable to obtain BMDs with alternative cost-effective and more accessible medical imaging examinations such as X-ray plain films. In this work, we formulate the BMD estimation from plain hip X-ray images as a regression problem. Specifically, we propose a new semi-supervised self-training algorithm to train the BMD regression model using images coupled with DEXA measured BMDs and unlabeled images with pseudo BMDs. Pseudo BMDs are generated and refined iteratively for unlabeled images during self-training. We also present a novel adaptive triplet loss to improve the model's regression accuracy. On an in-house dataset of 1,090 images (819 unique patients), our BMD estimation method achieves a high Pearson correlation coefficient of 0.8805 to ground-truth BMDs. It offers good feasibility to use the more accessible and cheaper X-ray imaging for opportunistic osteoporosis screening.
Learn molecular representations from large-scale unlabeled molecules for drug discovery
Dec 21, 2020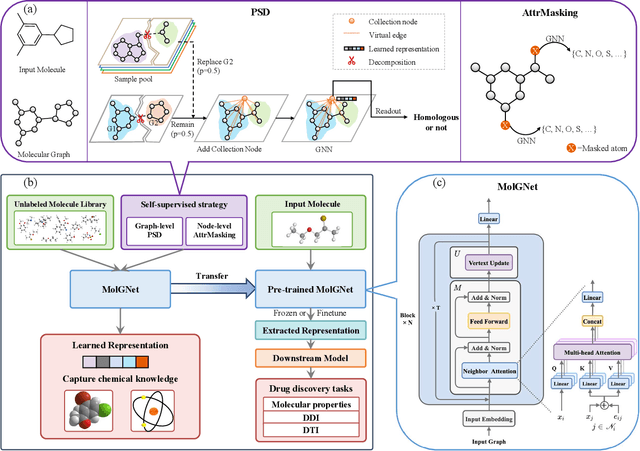
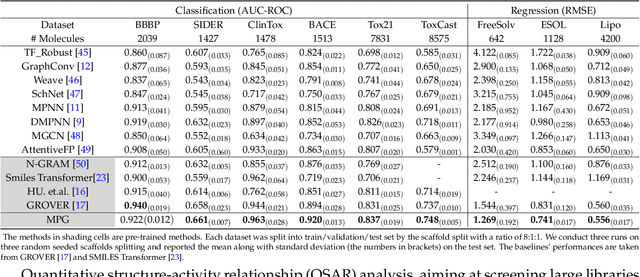
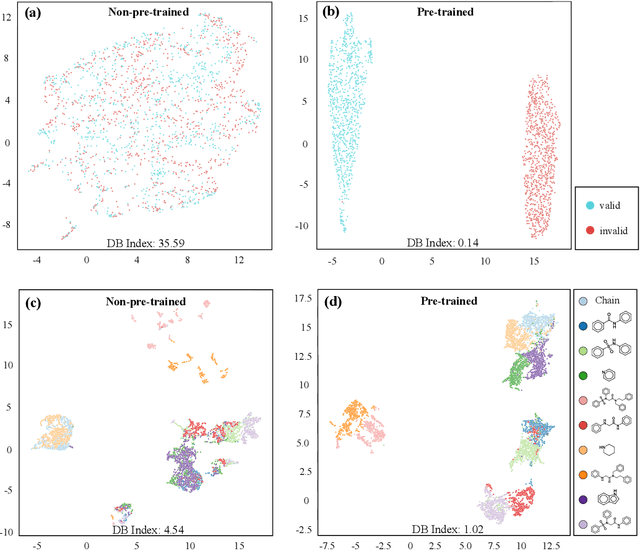
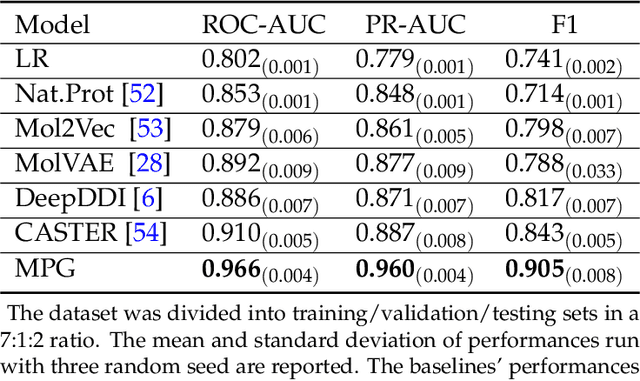
Abstract:How to produce expressive molecular representations is a fundamental challenge in AI-driven drug discovery. Graph neural network (GNN) has emerged as a powerful technique for modeling molecular data. However, previous supervised approaches usually suffer from the scarcity of labeled data and have poor generalization capability. Here, we proposed a novel Molecular Pre-training Graph-based deep learning framework, named MPG, that leans molecular representations from large-scale unlabeled molecules. In MPG, we proposed a powerful MolGNet model and an effective self-supervised strategy for pre-training the model at both the node and graph-level. After pre-training on 11 million unlabeled molecules, we revealed that MolGNet can capture valuable chemistry insights to produce interpretable representation. The pre-trained MolGNet can be fine-tuned with just one additional output layer to create state-of-the-art models for a wide range of drug discovery tasks, including molecular properties prediction, drug-drug interaction, and drug-target interaction, involving 13 benchmark datasets. Our work demonstrates that MPG is promising to become a novel approach in the drug discovery pipeline.
Semi-supervised Active Learning for Instance Segmentation via Scoring Predictions
Dec 09, 2020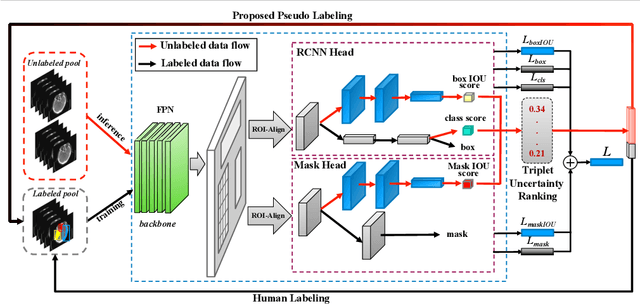
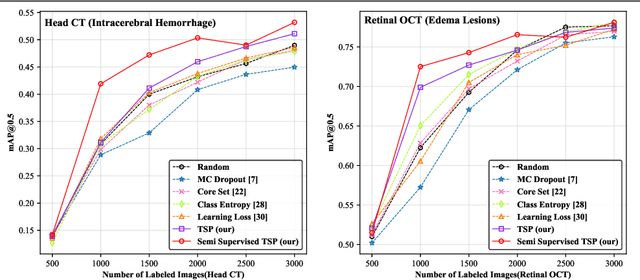
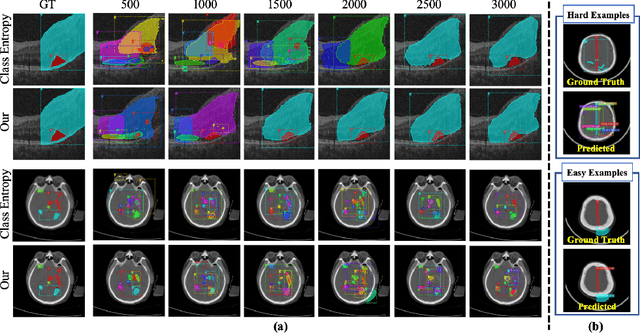
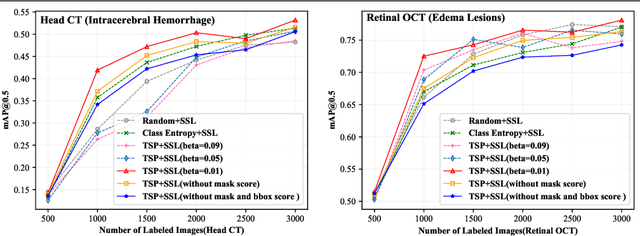
Abstract:Active learning generally involves querying the most representative samples for human labeling, which has been widely studied in many fields such as image classification and object detection. However, its potential has not been explored in the more complex instance segmentation task that usually has relatively higher annotation cost. In this paper, we propose a novel and principled semi-supervised active learning framework for instance segmentation. Specifically, we present an uncertainty sampling strategy named Triplet Scoring Predictions (TSP) to explicitly incorporate samples ranking clues from classes, bounding boxes and masks. Moreover, we devise a progressive pseudo labeling regime using the above TSP in semi-supervised manner, it can leverage both the labeled and unlabeled data to minimize labeling effort while maximize performance of instance segmentation. Results on medical images datasets demonstrate that the proposed method results in the embodiment of knowledge from available data in a meaningful way. The extensive quantitatively and qualitatively experiments show that, our method can yield the best-performing model with notable less annotation costs, compared with state-of-the-arts.
AutoRC: Improving BERT Based Relation Classification Models via Architecture Search
Sep 27, 2020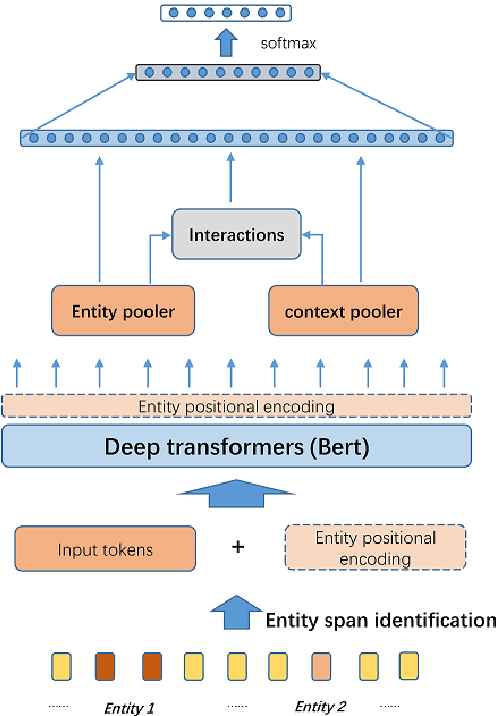
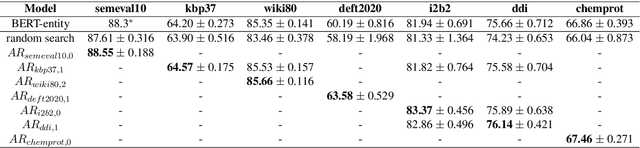
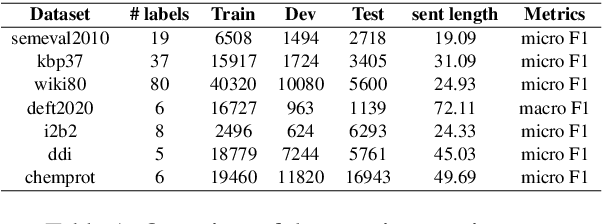
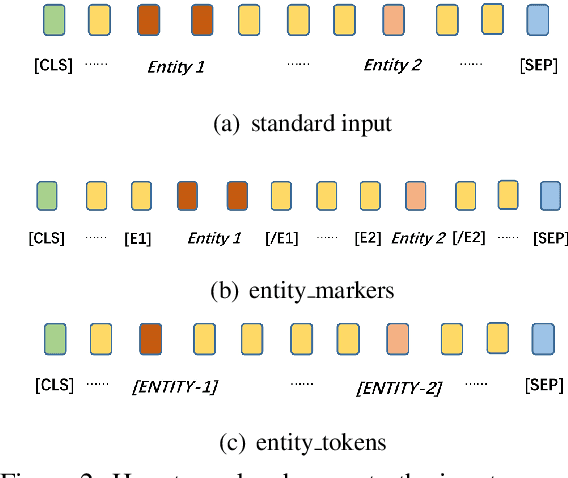

Abstract:Although BERT based relation classification (RC) models have achieved significant improvements over the traditional deep learning models, it seems that no consensus can be reached on what is the optimal architecture. Firstly, there are multiple alternatives for entity span identification. Second, there are a collection of pooling operations to aggregate the representations of entities and contexts into fixed length vectors. Third, it is difficult to manually decide which feature vectors, including their interactions, are beneficial for classifying the relation types. In this work, we design a comprehensive search space for BERT based RC models and employ neural architecture search (NAS) method to automatically discover the design choices mentioned above. Experiments on seven benchmark RC tasks show that our method is efficient and effective in finding better architectures than the baseline BERT based RC model. Ablation study demonstrates the necessity of our search space design and the effectiveness of our search method.
AutoTrans: Automating Transformer Design via Reinforced Architecture Search
Sep 04, 2020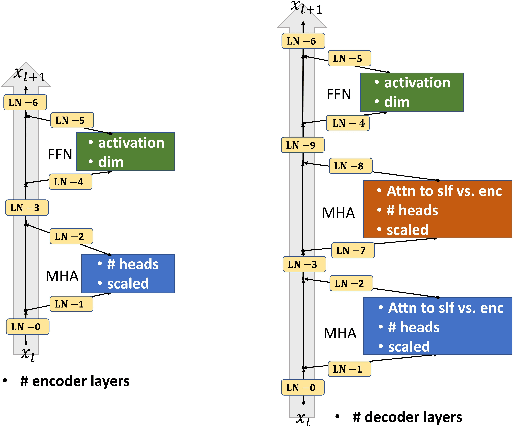
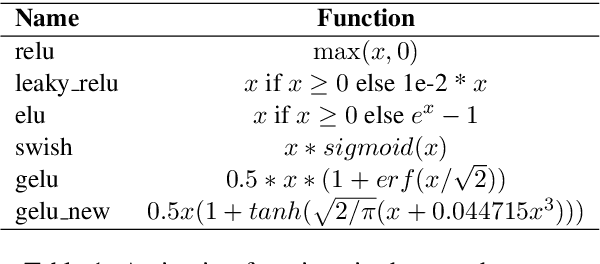

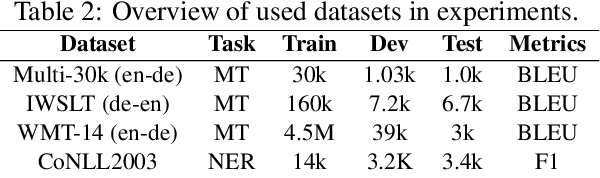
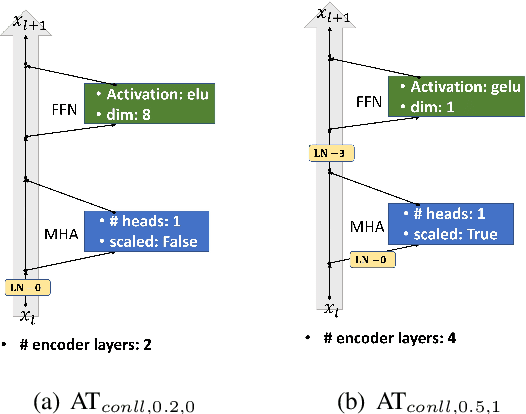
Abstract:Though the transformer architectures have shown dominance in many natural language understanding tasks, there are still unsolved issues for the training of transformer models, especially the need for a principled way of warm-up which has shown importance for stable training of a transformer, as well as whether the task at hand prefer to scale the attention product or not. In this paper, we empirically explore automating the design choices in the transformer model, i.e., how to set layer-norm, whether to scale, number of layers, number of heads, activation function, etc, so that one can obtain a transformer architecture that better suits the tasks at hand. RL is employed to navigate along search space, and special parameter sharing strategies are designed to accelerate the search. It is shown that sampling a proportion of training data per epoch during search help to improve the search quality. Experiments on the CoNLL03, Multi-30k, IWSLT14 and WMT-14 shows that the searched transformer model can outperform the standard transformers. In particular, we show that our learned model can be trained more robustly with large learning rates without warm-up.
Developing Knowledge-enhanced Chronic Disease Risk Prediction Models from Regional EHR Repositories
Jul 31, 2017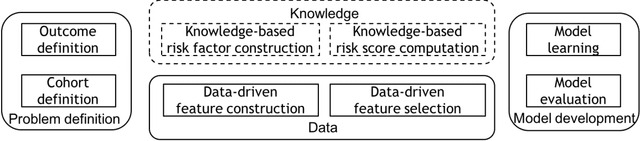
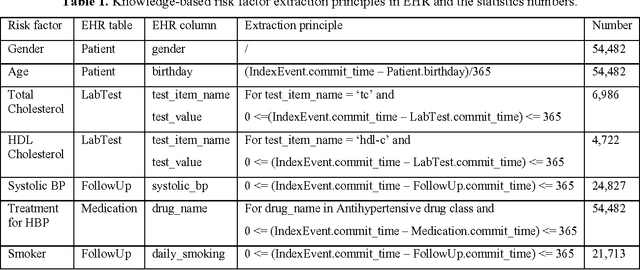

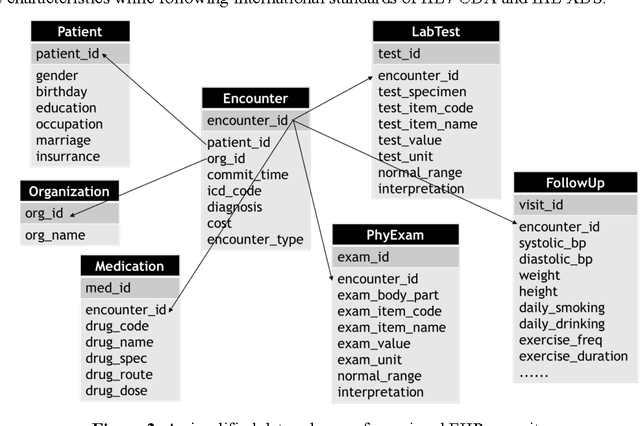
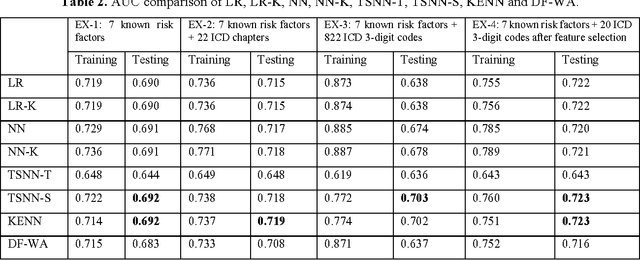
Abstract:Precision medicine requires the precision disease risk prediction models. In literature, there have been a lot well-established (inter-)national risk models, but when applying them into the local population, the prediction performance becomes unsatisfactory. To address the localization issue, this paper exploits the way to develop knowledge-enhanced localized risk models. On the one hand, we tune models by learning from regional Electronic Health Record (EHR) repositories, and on the other hand, we propose knowledge injection into the EHR data learning process. For experiments, we leverage the Pooled Cohort Equations (PCE, as recommended in ACC/AHA guidelines to estimate the risk of ASCVD) to develop a localized ASCVD risk prediction model in diabetes. The experimental results show that, if directly using the PCE algorithm on our cohort, the AUC is only 0.653, while our knowledge-enhanced localized risk model can achieve higher prediction performance with AUC of 0.723 (improved by 10.7%).
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge