"Image": models, code, and papers
Learning image representations for anomaly detection: application to discovery of histological alterations in drug development
Oct 14, 2022
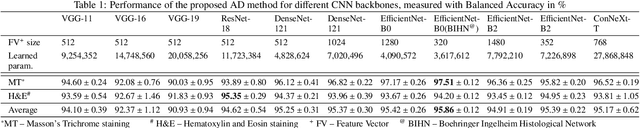
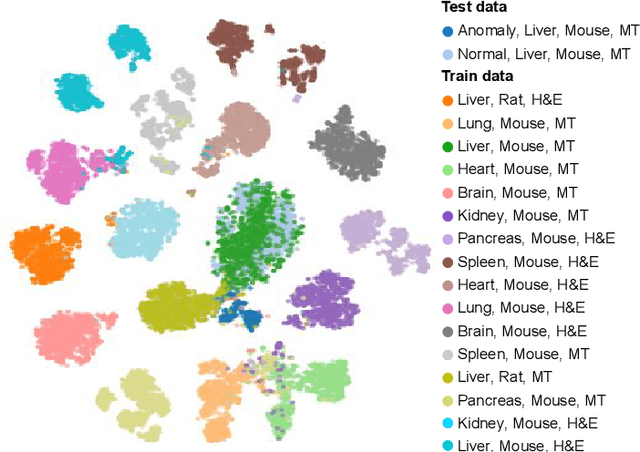
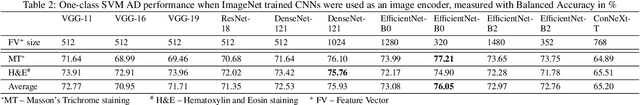
We present a system for anomaly detection in histopathological images. In histology, normal samples are usually abundant, whereas anomalous (pathological) cases are scarce or not available. Under such settings, one-class classifiers trained on healthy data can detect out-of-distribution anomalous samples. Such approaches combined with pre-trained Convolutional Neural Network (CNN) representations of images were previously employed for anomaly detection (AD). However, pre-trained off-the-shelf CNN representations may not be sensitive to abnormal conditions in tissues, while natural variations of healthy tissue may result in distant representations. To adapt representations to relevant details in healthy tissue we propose training a CNN on an auxiliary task that discriminates healthy tissue of different species, organs, and staining reagents. Almost no additional labeling workload is required, since healthy samples come automatically with aforementioned labels. During training we enforce compact image representations with a center-loss term, which further improves representations for AD. The proposed system outperforms established AD methods on a published dataset of liver anomalies. Moreover, it provided comparable results to conventional methods specifically tailored for quantification of liver anomalies. We show that our approach can be used for toxicity assessment of candidate drugs at early development stages and thereby may reduce expensive late-stage drug attrition.
Semi-supervised GAN for Bladder Tissue Classification in Multi-Domain Endoscopic Images
Dec 21, 2022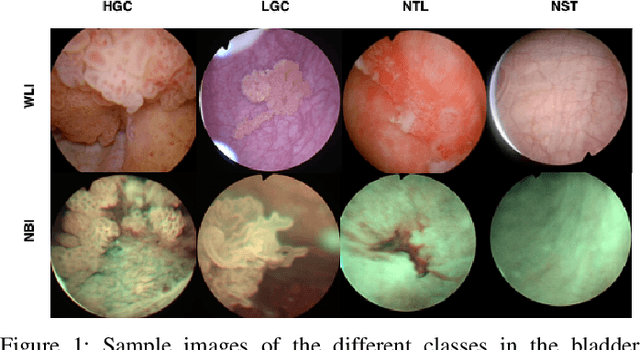
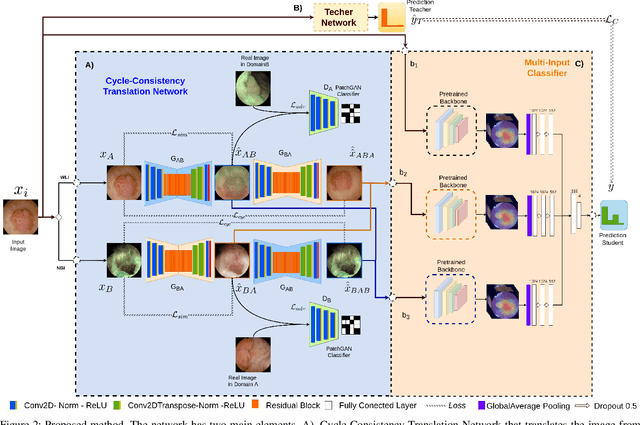
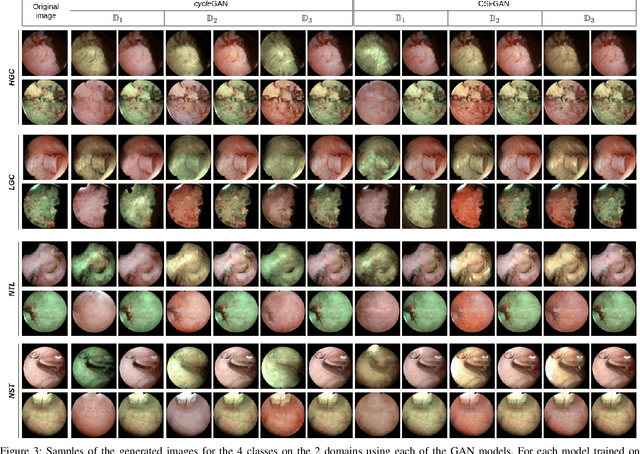
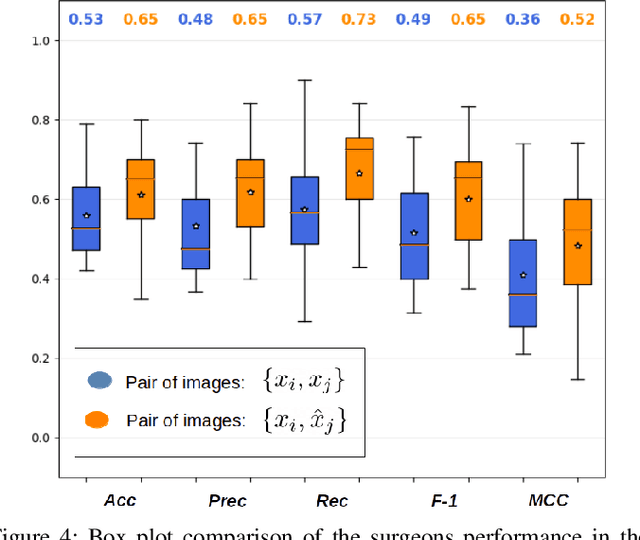
Objective: Accurate visual classification of bladder tissue during Trans-Urethral Resection of Bladder Tumor (TURBT) procedures is essential to improve early cancer diagnosis and treatment. During TURBT interventions, White Light Imaging (WLI) and Narrow Band Imaging (NBI) techniques are used for lesion detection. Each imaging technique provides diverse visual information that allows clinicians to identify and classify cancerous lesions. Computer vision methods that use both imaging techniques could improve endoscopic diagnosis. We address the challenge of tissue classification when annotations are available only in one domain, in our case WLI, and the endoscopic images correspond to an unpaired dataset, i.e. there is no exact equivalent for every image in both NBI and WLI domains. Method: We propose a semi-surprised Generative Adversarial Network (GAN)-based method composed of three main components: a teacher network trained on the labeled WLI data; a cycle-consistency GAN to perform unpaired image-to-image translation, and a multi-input student network. To ensure the quality of the synthetic images generated by the proposed GAN we perform a detailed quantitative, and qualitative analysis with the help of specialists. Conclusion: The overall average classification accuracy, precision, and recall obtained with the proposed method for tissue classification are 0.90, 0.88, and 0.89 respectively, while the same metrics obtained in the unlabeled domain (NBI) are 0.92, 0.64, and 0.94 respectively. The quality of the generated images is reliable enough to deceive specialists. Significance: This study shows the potential of using semi-supervised GAN-based classification to improve bladder tissue classification when annotations are limited in multi-domain data.
Analysis of displacement compensation methods for wavelet lifting of medical 3-D thorax CT volume data
Jan 11, 2023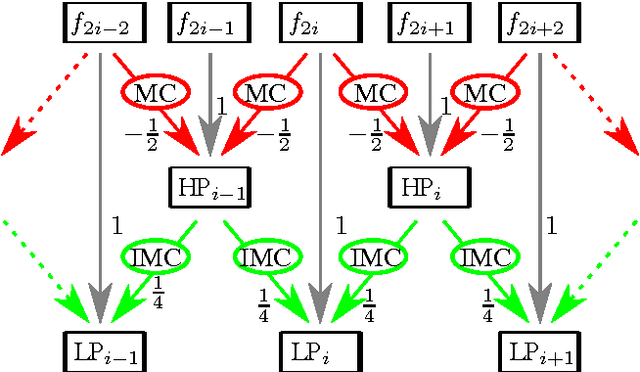
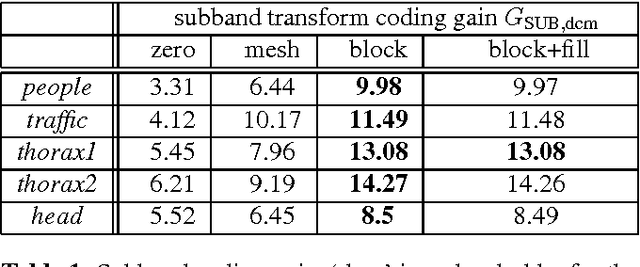
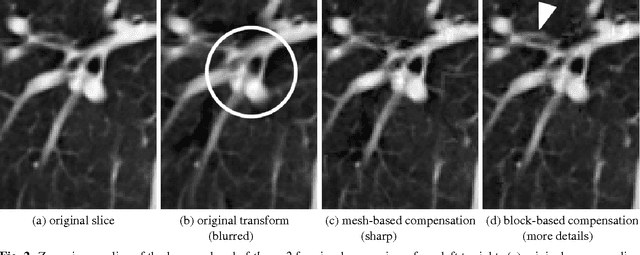
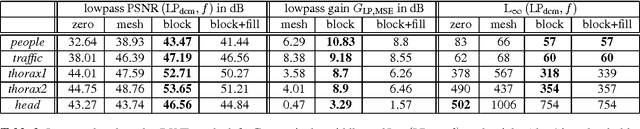
A huge advantage of the wavelet transform in image and video compression is its scalability. Wavelet-based coding of medical computed tomography (CT) data becomes more and more popular. While much effort has been spent on encoding of the wavelet coefficients, the extension of the transform by a compensation method as in video coding has not gained much attention so far. We will analyze two compensation methods for medical CT data and compare the characteristics of the displacement compensated wavelet transform with video data. We will show that for thorax CT data the transform coding gain can be improved by a factor of 2 and the quality of the lowpass band can be improved by 8 dB in terms of PSNR compared to the original transform without compensation.
DELTAR: Depth Estimation from a Light-weight ToF Sensor and RGB Image
Sep 27, 2022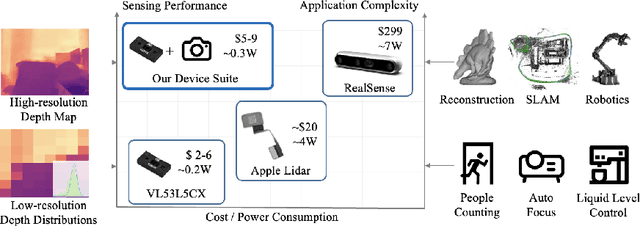
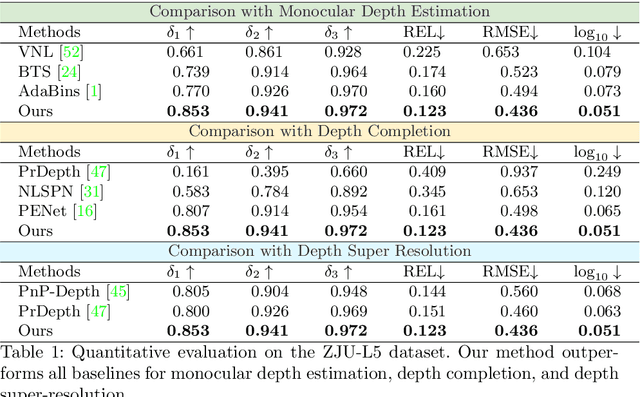
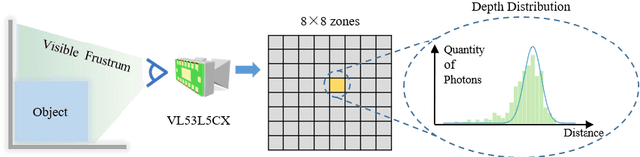
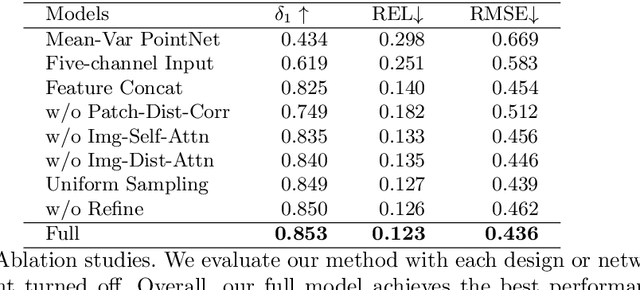
Light-weight time-of-flight (ToF) depth sensors are small, cheap, low-energy and have been massively deployed on mobile devices for the purposes like autofocus, obstacle detection, etc. However, due to their specific measurements (depth distribution in a region instead of the depth value at a certain pixel) and extremely low resolution, they are insufficient for applications requiring high-fidelity depth such as 3D reconstruction. In this paper, we propose DELTAR, a novel method to empower light-weight ToF sensors with the capability of measuring high resolution and accurate depth by cooperating with a color image. As the core of DELTAR, a feature extractor customized for depth distribution and an attention-based neural architecture is proposed to fuse the information from the color and ToF domain efficiently. To evaluate our system in real-world scenarios, we design a data collection device and propose a new approach to calibrate the RGB camera and ToF sensor. Experiments show that our method produces more accurate depth than existing frameworks designed for depth completion and depth super-resolution and achieves on par performance with a commodity-level RGB-D sensor. Code and data are available at https://zju3dv.github.io/deltar/.
Deep Learning for scalp High Frequency Oscillations Identification
Jan 20, 2023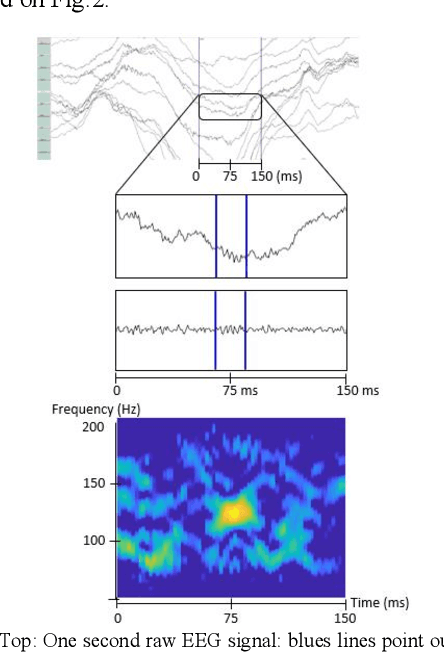
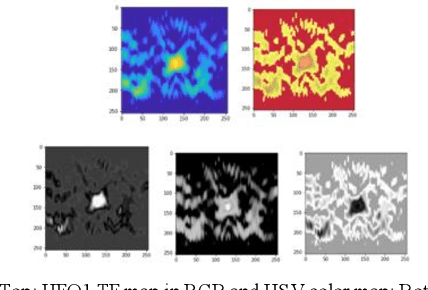
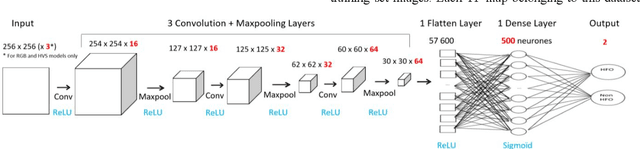
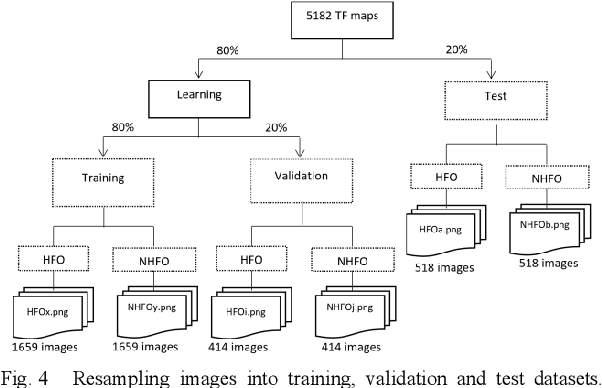
Since last 2 decades, High Frequency Oscillations (HFOs) are studied as a promising biomarker to localize the epileptogenic zone of patients with refractory focal epilepsy. As HFOs visual detection is time consuming and subjective, automatization of HFO detection is required. Most HFO detectors were developed on invasive electroencephalograms (iEEG) whereas scalp electroencephalograms (EEG) are used in clinical routine. In order HFO detection can benefit to more patients, scalp HFO detectors has to be developed. However, HFOs identification in scalp EEG is more challenging than in iEEG since scalp HFOs are of lower rate, lower amplitude and more likely to be corrupted by several sources of artifacts than iEEG HFOs. The main goal of this study is to explore the ability of deep learning architecture to identify scalp HFOs from the remaining EEG signal. Hence, a binary classification Convolutional Neural Network (CNN) is learned to analyze High Density Electroencephalograms (HD-EEG). EEG signals are first mapped into a 2D time-frequency image, several color definitions are then used as an input for the CNN. Experimental results show that deep learning allows simple end-to-end learning of preprocessing, feature extraction and classification modules while reaching competitive performance.
FG-Depth: Flow-Guided Unsupervised Monocular Depth Estimation
Jan 20, 2023
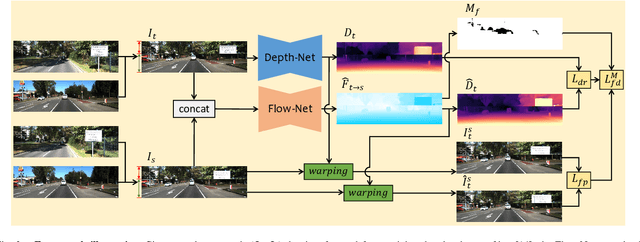
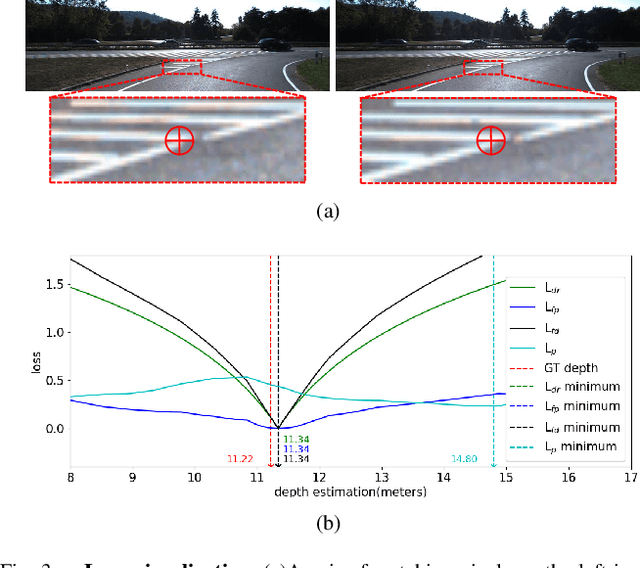

The great potential of unsupervised monocular depth estimation has been demonstrated by many works due to low annotation cost and impressive accuracy comparable to supervised methods. To further improve the performance, recent works mainly focus on designing more complex network structures and exploiting extra supervised information, e.g., semantic segmentation. These methods optimize the models by exploiting the reconstructed relationship between the target and reference images in varying degrees. However, previous methods prove that this image reconstruction optimization is prone to get trapped in local minima. In this paper, our core idea is to guide the optimization with prior knowledge from pretrained Flow-Net. And we show that the bottleneck of unsupervised monocular depth estimation can be broken with our simple but effective framework named FG-Depth. In particular, we propose (i) a flow distillation loss to replace the typical photometric loss that limits the capacity of the model and (ii) a prior flow based mask to remove invalid pixels that bring the noise in training loss. Extensive experiments demonstrate the effectiveness of each component, and our approach achieves state-of-the-art results on both KITTI and NYU-Depth-v2 datasets.
Deep Bayesian Active-Learning-to-Rank for Endoscopic Image Data
Aug 05, 2022Automatic image-based disease severity estimation generally uses discrete (i.e., quantized) severity labels. Annotating discrete labels is often difficult due to the images with ambiguous severity. An easier alternative is to use relative annotation, which compares the severity level between image pairs. By using a learning-to-rank framework with relative annotation, we can train a neural network that estimates rank scores that are relative to severity levels. However, the relative annotation for all possible pairs is prohibitive, and therefore, appropriate sample pair selection is mandatory. This paper proposes a deep Bayesian active-learning-to-rank, which trains a Bayesian convolutional neural network while automatically selecting appropriate pairs for relative annotation. We confirmed the efficiency of the proposed method through experiments on endoscopic images of ulcerative colitis. In addition, we confirmed that our method is useful even with the severe class imbalance because of its ability to select samples from minor classes automatically.
Automatic Classification of Single Tree Decay Stages from Combined ALS Data and Aerial Imagery using Machine Learning
Jan 04, 2023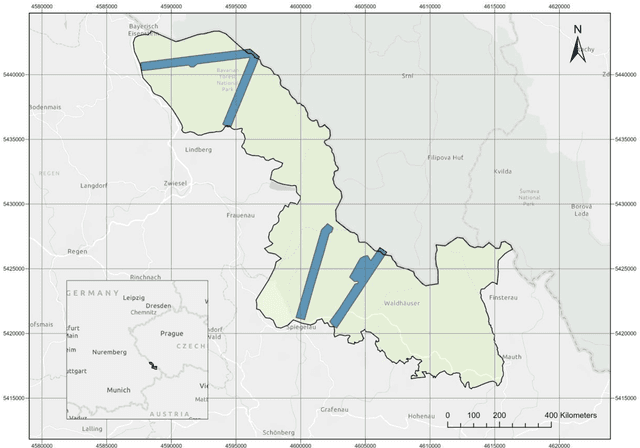
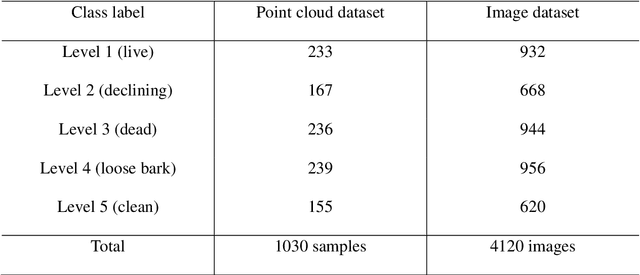

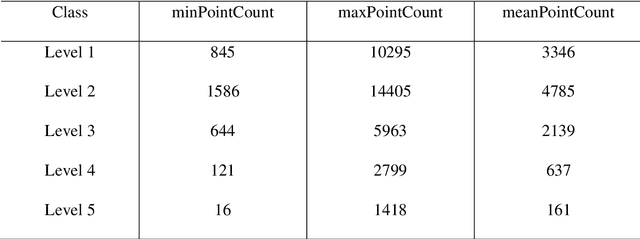
Understanding forest health is of great importance for the conservation of the integrity of forest ecosystems. The monitoring of forest health is, therefore, indispensable for the long-term conservation of forests and their sustainable management. In this regard, evaluating the amount and quality of dead wood is of utmost interest as they are favorable indicators of biodiversity. Apparently, remote sensing-based machine learning techniques have proven to be more efficient and sustainable with unprecedented accuracy in forest inventory. However, the application of these techniques is still in its infancy with respect to dead wood mapping. This study investigates for the first time the automatic classification of individual coniferous trees into five decay stages (live, declining, dead, loose bark, and clean) from combined airborne laser scanning (ALS) point clouds and CIR images using three Machine Learning methods - 3D point cloud-based deep learning (PointNet), Convolutional Neural Network (CNN), and Random Forest (RF). All models achieved promising results, reaching overall accuracy (OA) up to 90.9%, 90.6%, and 80.6% for CNN, RF, and PointNet, respectively. The experimental results reveal that the image-based approach notably outperformed the 3D point cloud-based one, while spectral image texture is of the highest relevance to the success of categorizing tree decay. Our models could therefore be used for automatic determination of single tree decay stages and landscape-wide assessment of dead wood amount and quality using modern airborne remote sensing techniques with machine/deep learning. The proposed method can contribute as an important and rigorous tool for monitoring biodiversity in forest ecosystems.
Adaptive adversarial training method for improving multi-scale GAN based on generalization bound theory
Nov 30, 2022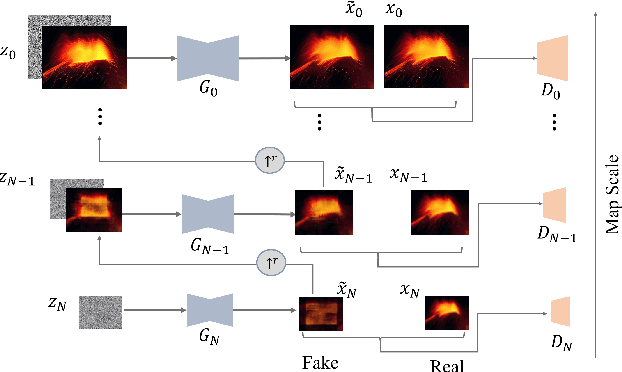
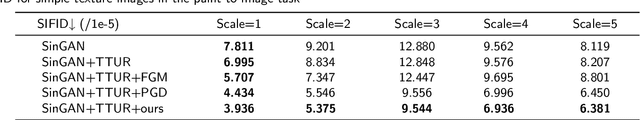
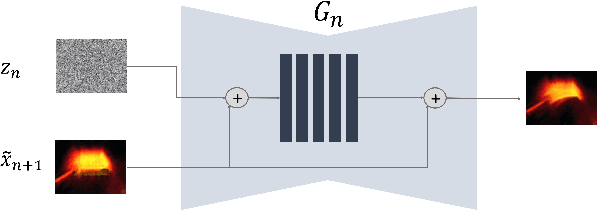
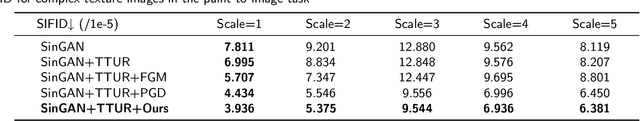
In recent years, multi-scale generative adversarial networks (GANs) have been proposed to build generalized image processing models based on single sample. Constraining on the sample size, multi-scale GANs have much difficulty converging to the global optimum, which ultimately leads to limitations in their capabilities. In this paper, we pioneered the introduction of PAC-Bayes generalized bound theory into the training analysis of specific models under different adversarial training methods, which can obtain a non-vacuous upper bound on the generalization error for the specified multi-scale GAN structure. Based on the drastic changes we found of the generalization error bound under different adversarial attacks and different training states, we proposed an adaptive training method which can greatly improve the image manipulation ability of multi-scale GANs. The final experimental results show that our adaptive training method in this paper has greatly contributed to the improvement of the quality of the images generated by multi-scale GANs on several image manipulation tasks. In particular, for the image super-resolution restoration task, the multi-scale GAN model trained by the proposed method achieves a 100% reduction in natural image quality evaluator (NIQE) and a 60% reduction in root mean squared error (RMSE), which is better than many models trained on large-scale datasets.
SGDraw: Scene Graph Drawing Interface Using Object-Oriented Representation
Nov 30, 2022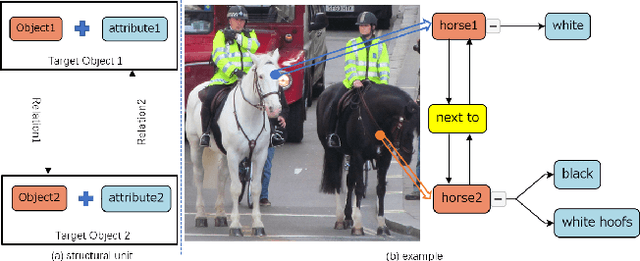
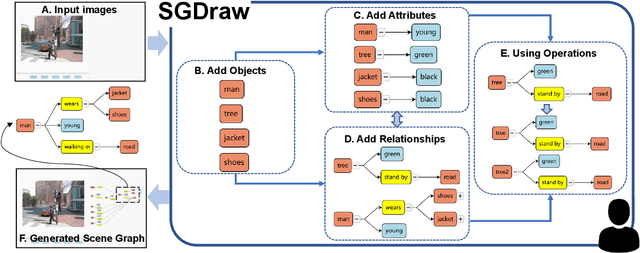
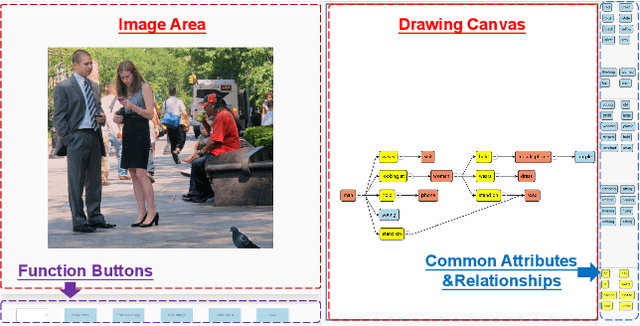
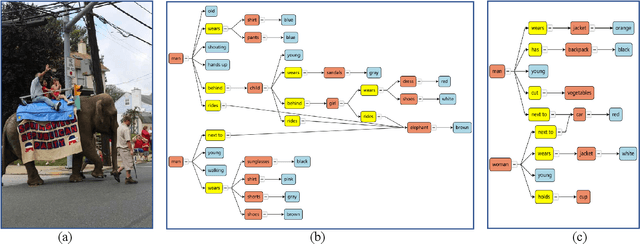
Scene understanding is an essential and challenging task in computer vision. To provide the visually fundamental graphical structure of an image, the scene graph has received increased attention due to its powerful semantic representation. However, it is difficult to draw a proper scene graph for image retrieval, image generation, and multi-modal applications. The conventional scene graph annotation interface is not easy to use in image annotations, and the automatic scene graph generation approaches using deep neural networks are prone to generate redundant content while disregarding details. In this work, we propose SGDraw, a scene graph drawing interface using object-oriented scene graph representation to help users draw and edit scene graphs interactively. For the proposed object-oriented representation, we consider the objects, attributes, and relationships of objects as a structural unit. SGDraw provides a web-based scene graph annotation and generation tool for scene understanding applications. To verify the effectiveness of the proposed interface, we conducted a comparison study with the conventional tool and the user experience study. The results show that SGDraw can help generate scene graphs with richer details and describe the images more accurately than traditional bounding box annotations. We believe the proposed SGDraw can be useful in various vision tasks, such as image retrieval and generation.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge