Tao Shen
PCL: Peer-Contrastive Learning with Diverse Augmentations for Unsupervised Sentence Embeddings
Jan 28, 2022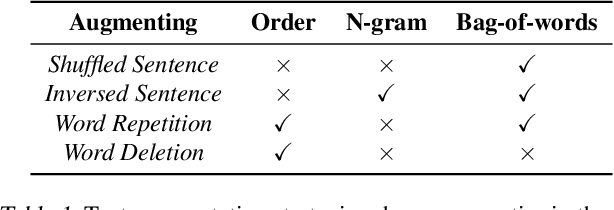
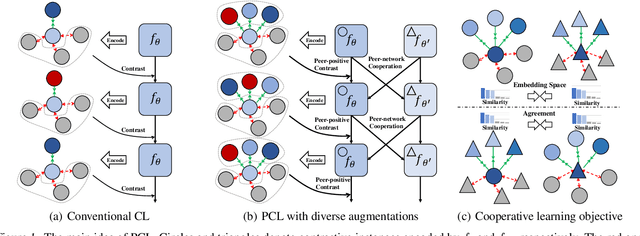
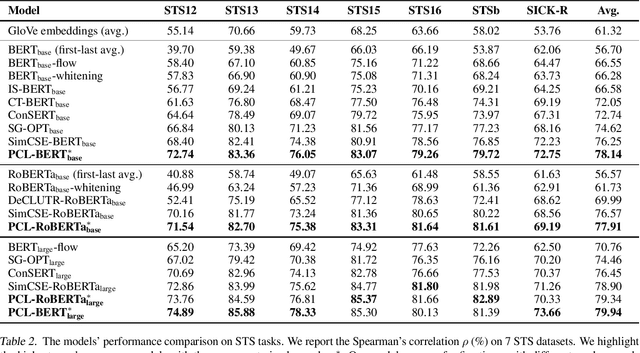
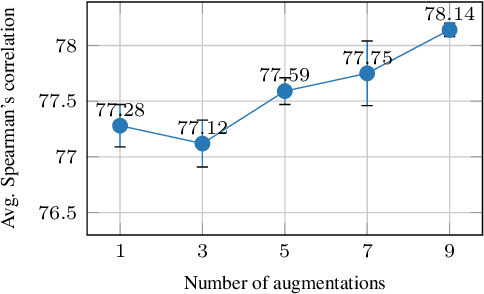
Abstract:Learning sentence embeddings in an unsupervised manner is fundamental in natural language processing. Recent common practice is to couple pre-trained language models with unsupervised contrastive learning, whose success relies on augmenting a sentence with a semantically-close positive instance to construct contrastive pairs. Nonetheless, existing approaches usually depend on a mono-augmenting strategy, which causes learning shortcuts towards the augmenting biases and thus corrupts the quality of sentence embeddings. A straightforward solution is resorting to more diverse positives from a multi-augmenting strategy, while an open question remains about how to unsupervisedly learn from the diverse positives but with uneven augmenting qualities in the text field. As one answer, we propose a novel Peer-Contrastive Learning (PCL) with diverse augmentations. PCL constructs diverse contrastive positives and negatives at the group level for unsupervised sentence embeddings. PCL can perform peer-positive contrast as well as peer-network cooperation, which offers an inherent anti-bias ability and an effective way to learn from diverse augmentations. Experiments on STS benchmarks verify the effectiveness of our PCL against its competitors in unsupervised sentence embeddings.
Edge-Cloud Polarization and Collaboration: A Comprehensive Survey
Nov 12, 2021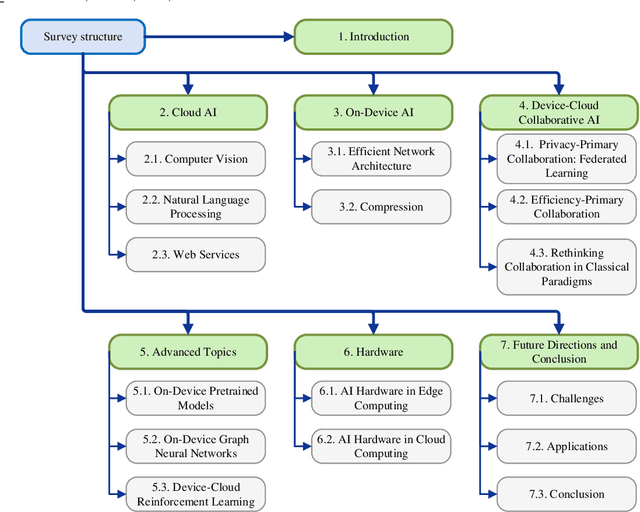
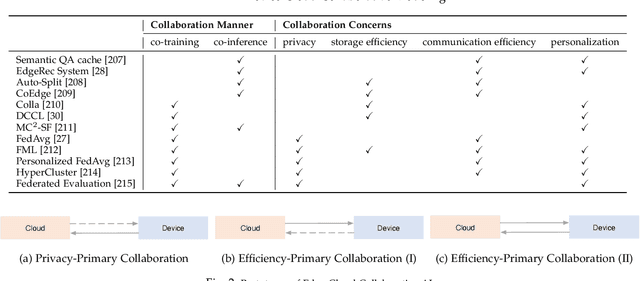
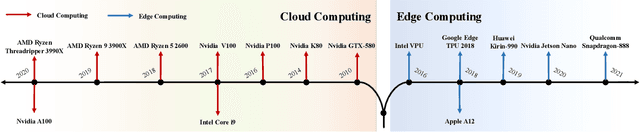
Abstract:Influenced by the great success of deep learning via cloud computing and the rapid development of edge chips, research in artificial intelligence (AI) has shifted to both of the computing paradigms, i.e., cloud computing and edge computing. In recent years, we have witnessed significant progress in developing more advanced AI models on cloud servers that surpass traditional deep learning models owing to model innovations (e.g., Transformers, Pretrained families), explosion of training data and soaring computing capabilities. However, edge computing, especially edge and cloud collaborative computing, are still in its infancy to announce their success due to the resource-constrained IoT scenarios with very limited algorithms deployed. In this survey, we conduct a systematic review for both cloud and edge AI. Specifically, we are the first to set up the collaborative learning mechanism for cloud and edge modeling with a thorough review of the architectures that enable such mechanism. We also discuss potentials and practical experiences of some on-going advanced edge AI topics including pretraining models, graph neural networks and reinforcement learning. Finally, we discuss the promising directions and challenges in this field.
EventBERT: A Pre-Trained Model for Event Correlation Reasoning
Oct 13, 2021
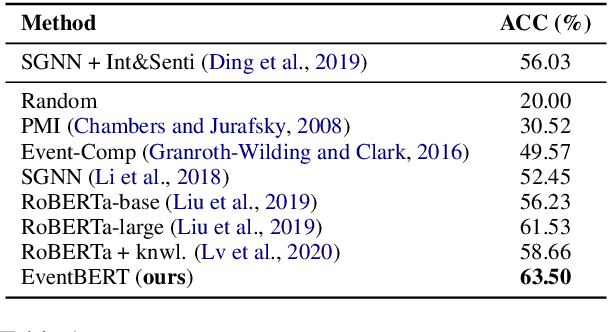
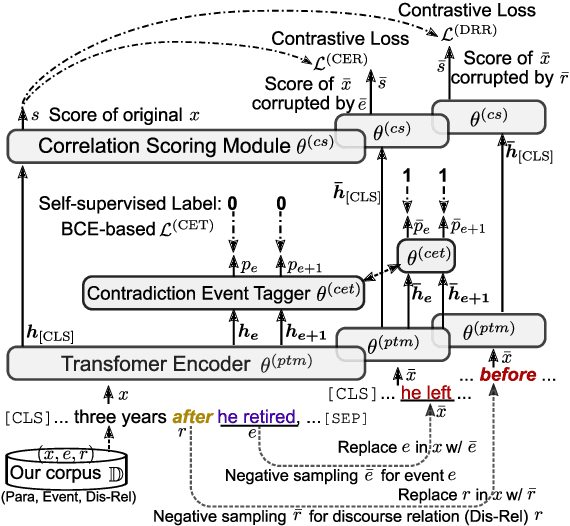
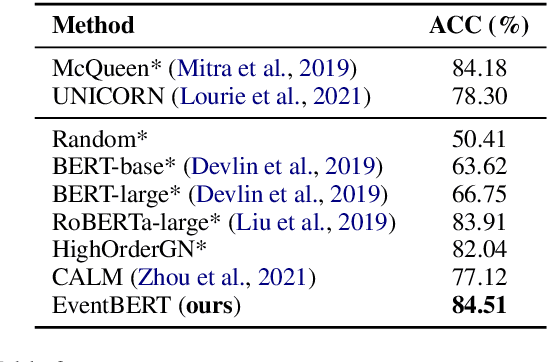
Abstract:Event correlation reasoning infers whether a natural language paragraph containing multiple events conforms to human common sense. For example, "Andrew was very drowsy, so he took a long nap, and now he is very alert" is sound and reasonable. In contrast, "Andrew was very drowsy, so he stayed up a long time, now he is very alert" does not comply with human common sense. Such reasoning capability is essential for many downstream tasks, such as script reasoning, abductive reasoning, narrative incoherence, story cloze test, etc. However, conducting event correlation reasoning is challenging due to a lack of large amounts of diverse event-based knowledge and difficulty in capturing correlation among multiple events. In this paper, we propose EventBERT, a pre-trained model to encapsulate eventuality knowledge from unlabeled text. Specifically, we collect a large volume of training examples by identifying natural language paragraphs that describe multiple correlated events and further extracting event spans in an unsupervised manner. We then propose three novel event- and correlation-based learning objectives to pre-train an event correlation model on our created training corpus. Empirical results show EventBERT outperforms strong baselines on four downstream tasks, and achieves SoTA results on most of them. Besides, it outperforms existing pre-trained models by a large margin, e.g., 6.5~23%, in zero-shot learning of these tasks.
Hierarchical Relation-Guided Type-Sentence Alignment for Long-Tail Relation Extraction with Distant Supervision
Sep 19, 2021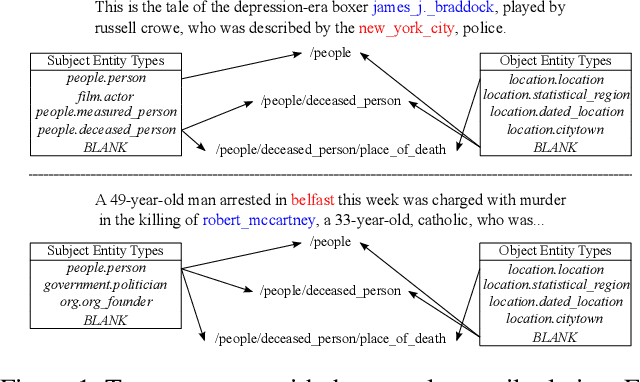
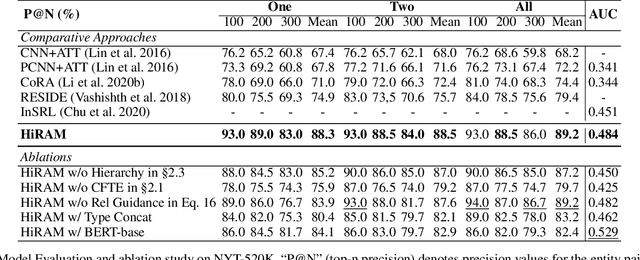
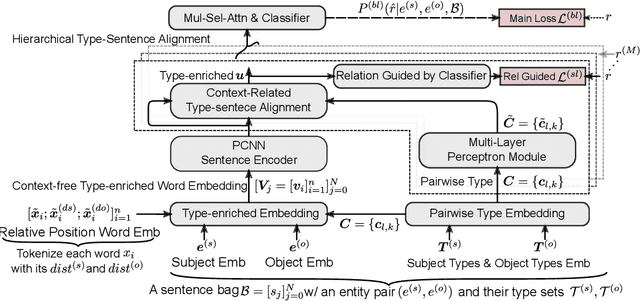
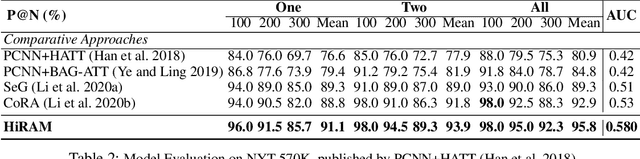
Abstract:Distant supervision uses triple facts in knowledge graphs to label a corpus for relation extraction, leading to wrong labeling and long-tail problems. Some works use the hierarchy of relations for knowledge transfer to long-tail relations. However, a coarse-grained relation often implies only an attribute (e.g., domain or topic) of the distant fact, making it hard to discriminate relations based solely on sentence semantics. One solution is resorting to entity types, but open questions remain about how to fully leverage the information of entity types and how to align multi-granular entity types with sentences. In this work, we propose a novel model to enrich distantly-supervised sentences with entity types. It consists of (1) a pairwise type-enriched sentence encoding module injecting both context-free and -related backgrounds to alleviate sentence-level wrong labeling, and (2) a hierarchical type-sentence alignment module enriching a sentence with the triple fact's basic attributes to support long-tail relations. Our model achieves new state-of-the-art results in overall and long-tail performance on benchmarks.
Sequential Diagnosis Prediction with Transformer and Ontological Representation
Sep 07, 2021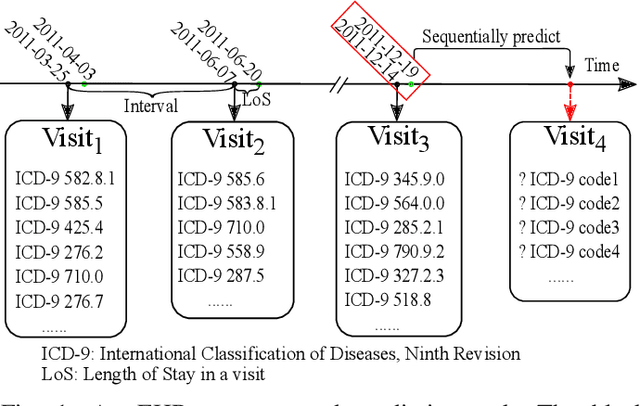
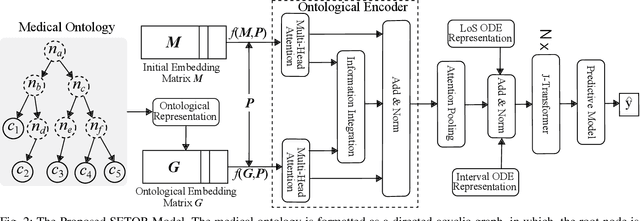
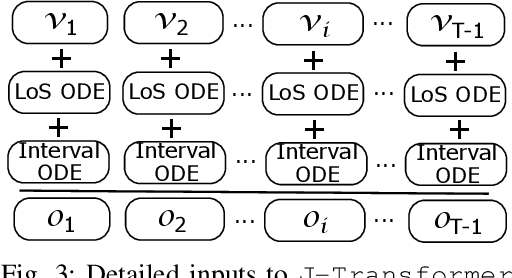
Abstract:Sequential diagnosis prediction on the Electronic Health Record (EHR) has been proven crucial for predictive analytics in the medical domain. EHR data, sequential records of a patient's interactions with healthcare systems, has numerous inherent characteristics of temporality, irregularity and data insufficiency. Some recent works train healthcare predictive models by making use of sequential information in EHR data, but they are vulnerable to irregular, temporal EHR data with the states of admission/discharge from hospital, and insufficient data. To mitigate this, we propose an end-to-end robust transformer-based model called SETOR, which exploits neural ordinary differential equation to handle both irregular intervals between a patient's visits with admitted timestamps and length of stay in each visit, to alleviate the limitation of insufficient data by integrating medical ontology, and to capture the dependencies between the patient's visits by employing multi-layer transformer blocks. Experiments conducted on two real-world healthcare datasets show that, our sequential diagnoses prediction model SETOR not only achieves better predictive results than previous state-of-the-art approaches, irrespective of sufficient or insufficient training data, but also derives more interpretable embeddings of medical codes. The experimental codes are available at the GitHub repository (https://github.com/Xueping/SETOR).
Eliminating Sentiment Bias for Aspect-Level Sentiment Classification with Unsupervised Opinion Extraction
Sep 07, 2021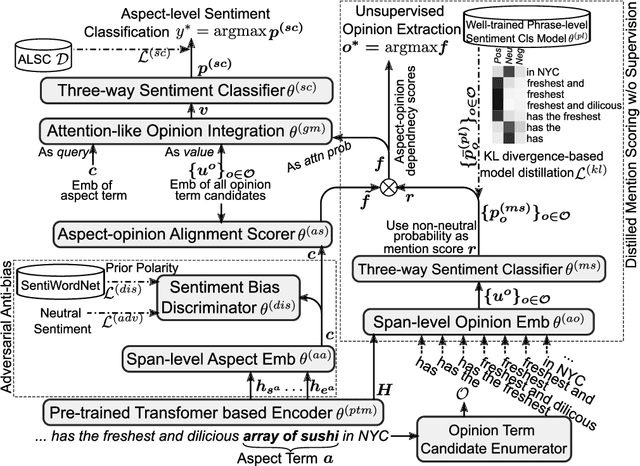
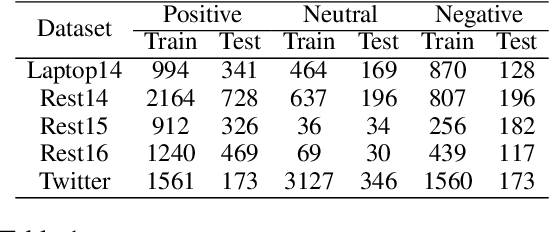

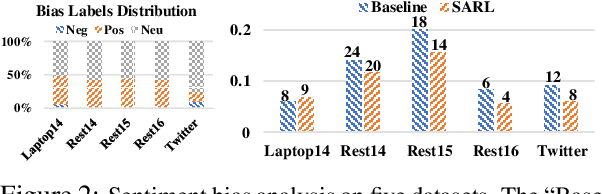
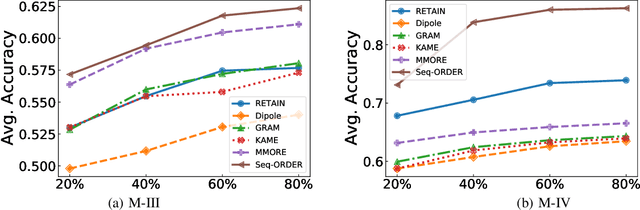
Abstract:Aspect-level sentiment classification (ALSC) aims at identifying the sentiment polarity of a specified aspect in a sentence. ALSC is a practical setting in aspect-based sentiment analysis due to no opinion term labeling needed, but it fails to interpret why a sentiment polarity is derived for the aspect. To address this problem, recent works fine-tune pre-trained Transformer encoders for ALSC to extract an aspect-centric dependency tree that can locate the opinion words. However, the induced opinion words only provide an intuitive cue far below human-level interpretability. Besides, the pre-trained encoder tends to internalize an aspect's intrinsic sentiment, causing sentiment bias and thus affecting model performance. In this paper, we propose a span-based anti-bias aspect representation learning framework. It first eliminates the sentiment bias in the aspect embedding by adversarial learning against aspects' prior sentiment. Then, it aligns the distilled opinion candidates with the aspect by span-based dependency modeling to highlight the interpretable opinion terms. Our method achieves new state-of-the-art performance on five benchmarks, with the capability of unsupervised opinion extraction.
Federated Learning for Privacy-Preserving Open Innovation Future on Digital Health
Aug 24, 2021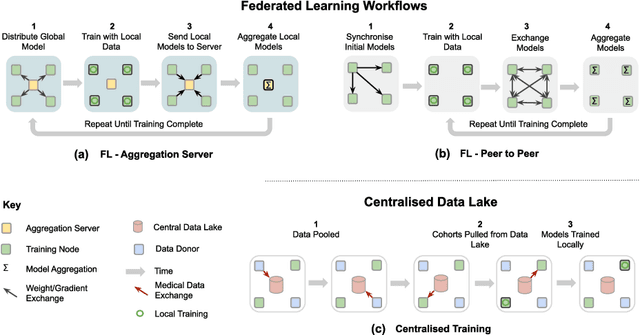


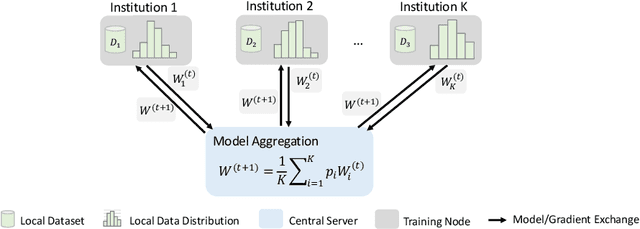
Abstract:Privacy protection is an ethical issue with broad concern in Artificial Intelligence (AI). Federated learning is a new machine learning paradigm to learn a shared model across users or organisations without direct access to the data. It has great potential to be the next-general AI model training framework that offers privacy protection and therefore has broad implications for the future of digital health and healthcare informatics. Implementing an open innovation framework in the healthcare industry, namely open health, is to enhance innovation and creative capability of health-related organisations by building a next-generation collaborative framework with partner organisations and the research community. In particular, this game-changing collaborative framework offers knowledge sharing from diverse data with a privacy-preserving. This chapter will discuss how federated learning can enable the development of an open health ecosystem with the support of AI. Existing challenges and solutions for federated learning will be discussed.
Multi-Center Federated Learning
Aug 21, 2021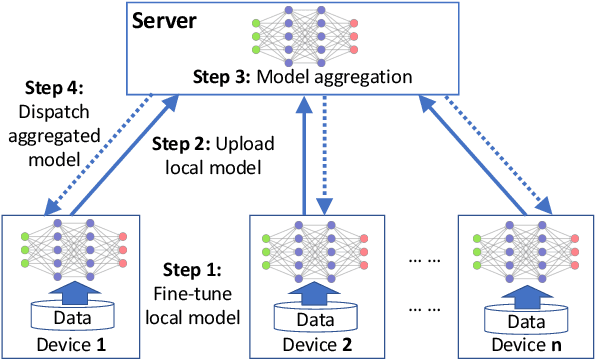
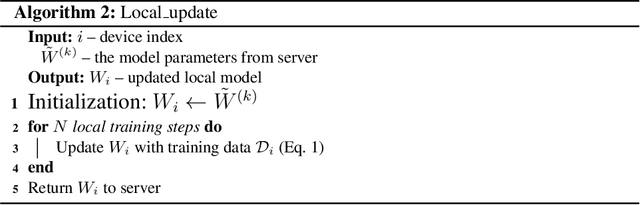

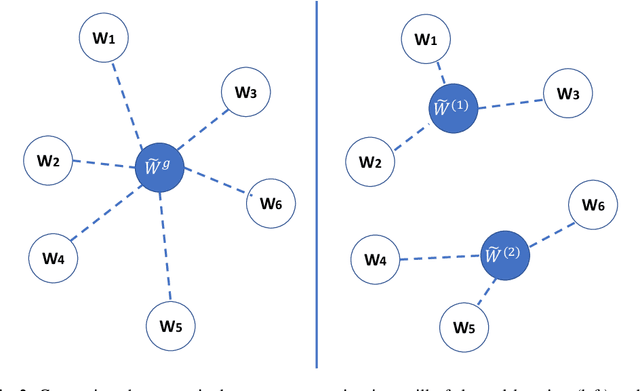
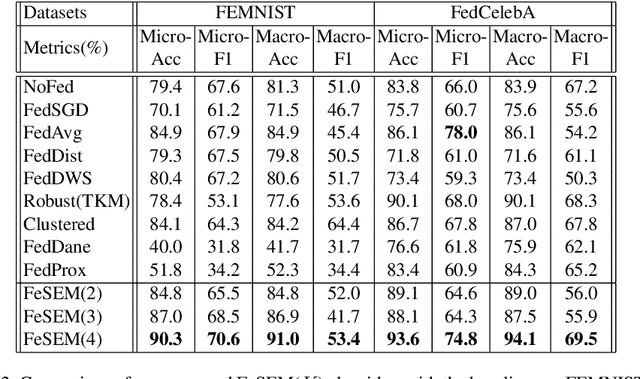
Abstract:Federated learning (FL) can protect data privacy in distributed learning since it merely collects local gradients from users without access to their data. However, FL is fragile in the presence of heterogeneity that is commonly encountered in practical settings, e.g., non-IID data over different users. Existing FL approaches usually update a single global model to capture the shared knowledge of all users by aggregating their gradients, regardless of the discrepancy between their data distributions. By comparison, a mixture of multiple global models could capture the heterogeneity across various users if assigning the users to different global models (i.e., centers) in FL. To this end, we propose a novel multi-center aggregation mechanism . It learns multiple global models from data, and simultaneously derives the optimal matching between users and centers. We then formulate it as a bi-level optimization problem that can be efficiently solved by a stochastic expectation maximization (EM) algorithm. Experiments on multiple benchmark datasets of FL show that our method outperforms several popular FL competitors. The source code are open source on Github.
Dynamic Prediction Model for NOx Emission of SCR System Based on Hybrid Data-driven Algorithms
Aug 03, 2021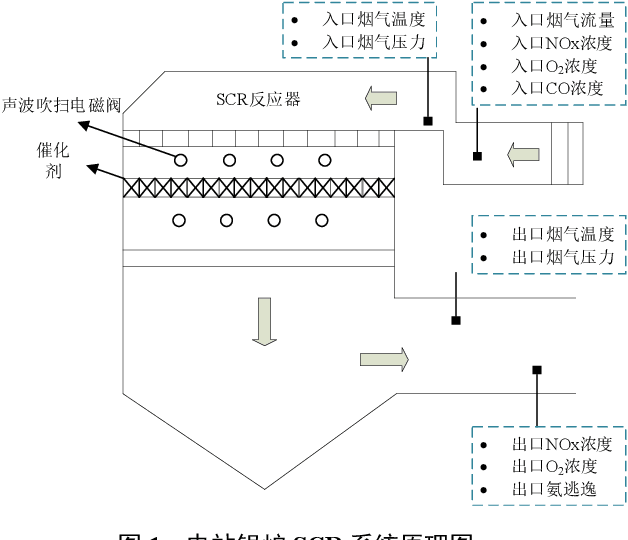
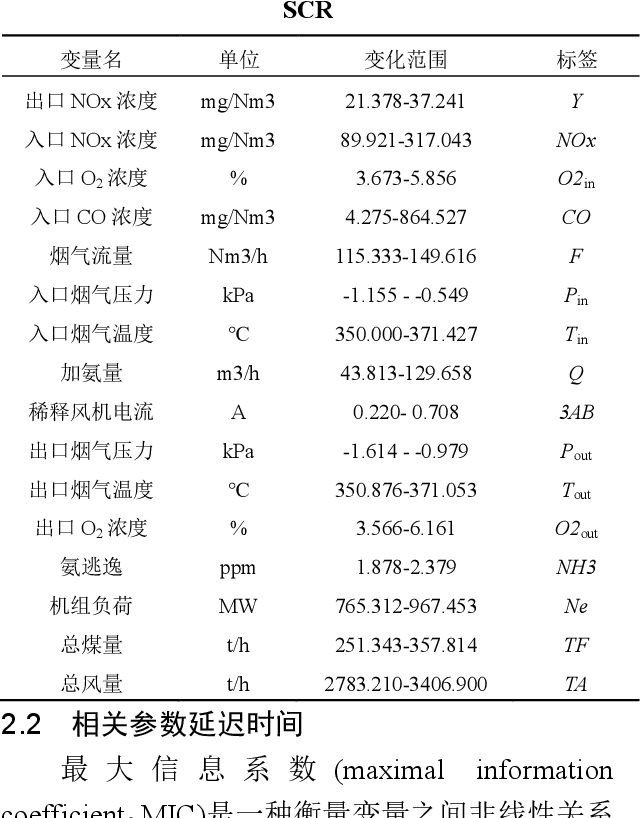
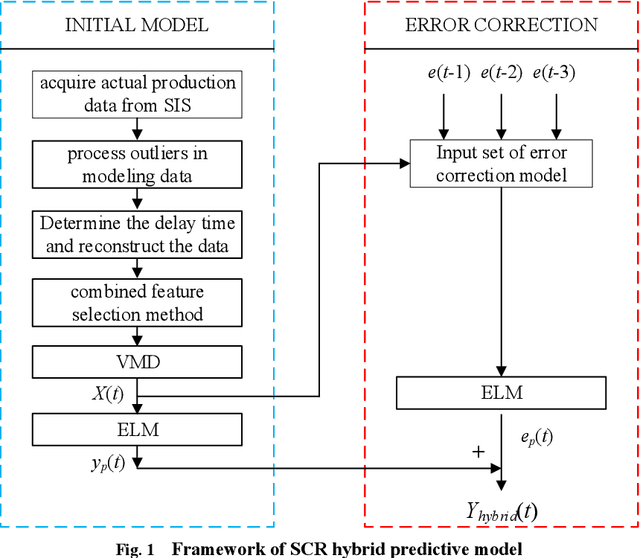
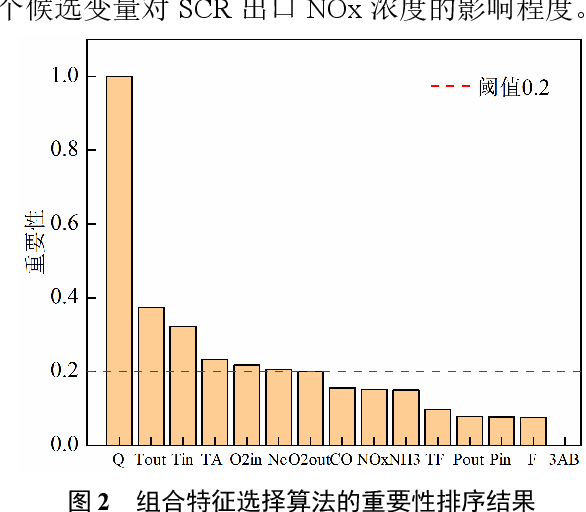
Abstract:Aiming at the problem that delay time is difficult to determine and prediction accuracy is low in building prediction model of SCR system, a dynamic modeling scheme based on a hybrid of multiple data-driven algorithms was proposed. First, processed abnormal values and normalized the data. To improve the relevance of the input data, used MIC to estimate delay time and reconstructed production data. Then used combined feature selection method to determine input variables. To further mine data information, VMD was used to decompose input time series. Finally, established NOx emission prediction model combining ELM and EC model. Experimental results based on actual historical operating data show that the MAPE of predicted results is 2.61%. Model sensitivity analysis shows that besides the amount of ammonia injection, the inlet oxygen concentration and the flue gas temperature have a significant impact on NOx emission, which should be considered in SCR process control and optimization.
MIMO: Mutual Integration of Patient Journey and Medical Ontology for Healthcare Representation Learning
Jul 23, 2021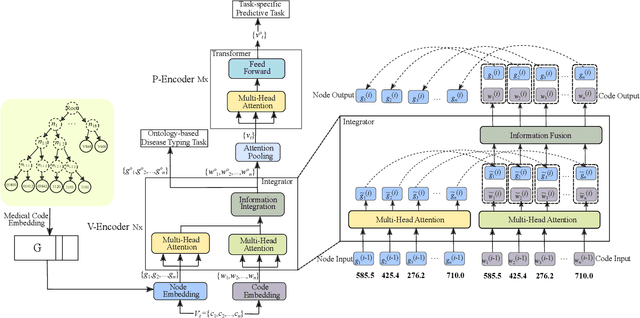
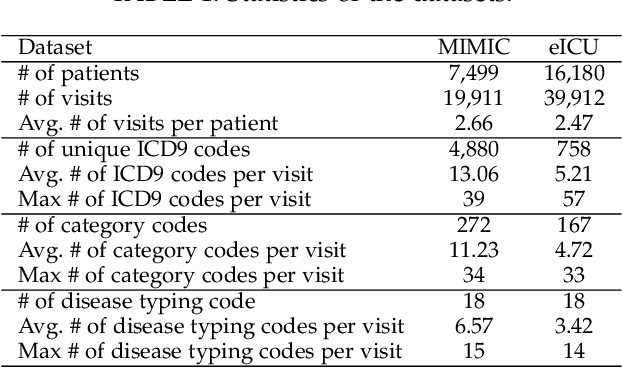
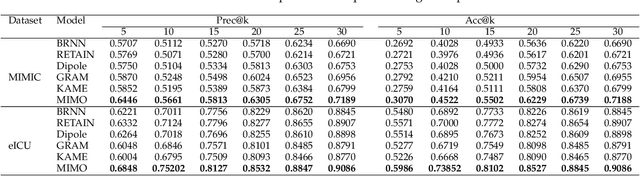
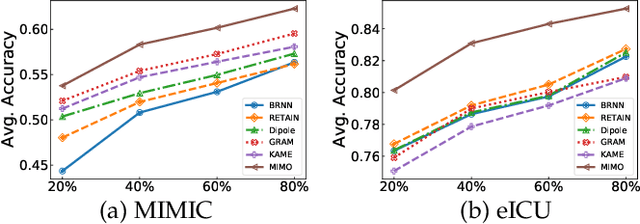
Abstract:Healthcare representation learning on the Electronic Health Record (EHR) is seen as crucial for predictive analytics in the medical field. Many natural language processing techniques, such as word2vec, RNN and self-attention, have been adapted for use in hierarchical and time stamped EHR data, but fail when they lack either general or task-specific data. Hence, some recent works train healthcare representations by incorporating medical ontology (a.k.a. knowledge graph), by self-supervised tasks like diagnosis prediction, but (1) the small-scale, monotonous ontology is insufficient for robust learning, and (2) critical contexts or dependencies underlying patient journeys are never exploited to enhance ontology learning. To address this, we propose an end-to-end robust Transformer-based solution, Mutual Integration of patient journey and Medical Ontology (MIMO) for healthcare representation learning and predictive analytics. Specifically, it consists of task-specific representation learning and graph-embedding modules to learn both patient journey and medical ontology interactively. Consequently, this creates a mutual integration to benefit both healthcare representation learning and medical ontology embedding. Moreover, such integration is achieved by a joint training of both task-specific predictive and ontology-based disease typing tasks based on fused embeddings of the two modules. Experiments conducted on two real-world diagnosis prediction datasets show that, our healthcare representation model MIMO not only achieves better predictive results than previous state-of-the-art approaches regardless of sufficient or insufficient training data, but also derives more interpretable embeddings of diagnoses.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge