Siqi Sun
Design, Actuation, and Functionalization of Untethered Soft Magnetic Robots with Life-Like Motions: A Review
May 28, 2023



Abstract:Soft robots have demonstrated superior flexibility and functionality than conventional rigid robots. These versatile devices can respond to a wide range of external stimuli (including light, magnetic field, heat, electric field, etc.), and can perform sophisticated tasks. Notably, soft magnetic robots exhibit unparalleled advantages among numerous soft robots (such as untethered control, rapid response, and high safety), and have made remarkable progress in small-scale manipulation tasks and biomedical applications. Despite the promising potential, soft magnetic robots are still in their infancy and require significant advancements in terms of fabrication, design principles, and functional development to be viable for real-world applications. Recent progress shows that bionics can serve as an effective tool for developing soft robots. In light of this, the review is presented with two main goals: (i) exploring how innovative bioinspired strategies can revolutionize the design and actuation of soft magnetic robots to realize various life-like motions; (ii) examining how these bionic systems could benefit practical applications in small-scale solid/liquid manipulation and therapeutic/diagnostic-related biomedical fields.
AF2-Mutation: Adversarial Sequence Mutations against AlphaFold2 on Protein Tertiary Structure Prediction
May 15, 2023Abstract:Deep learning-based approaches, such as AlphaFold2 (AF2), have significantly advanced protein tertiary structure prediction, achieving results comparable to real biological experimental methods. While AF2 has shown limitations in predicting the effects of mutations, its robustness against sequence mutations remains to be determined. Starting with the wild-type (WT) sequence, we investigate adversarial sequences generated via an evolutionary approach, which AF2 predicts to be substantially different from WT. Our experiments on CASP14 reveal that by modifying merely three residues in the protein sequence using a combination of replacement, deletion, and insertion strategies, the alteration in AF2's predictions, as measured by the Local Distance Difference Test (lDDT), reaches 46.61. Moreover, when applied to a specific protein, SPNS2, our proposed algorithm successfully identifies biologically meaningful residues critical to protein structure determination and potentially indicates alternative conformations, thus significantly expediting the experimental process.
E2Efold-3D: End-to-End Deep Learning Method for accurate de novo RNA 3D Structure Prediction
Jul 04, 2022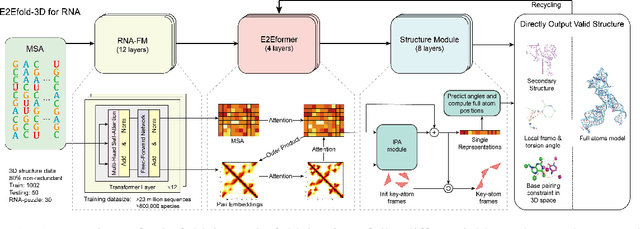
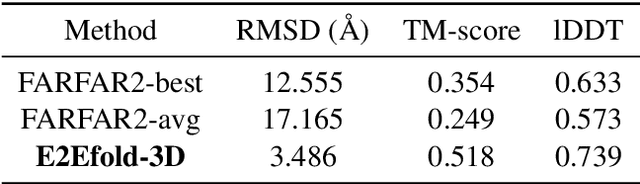
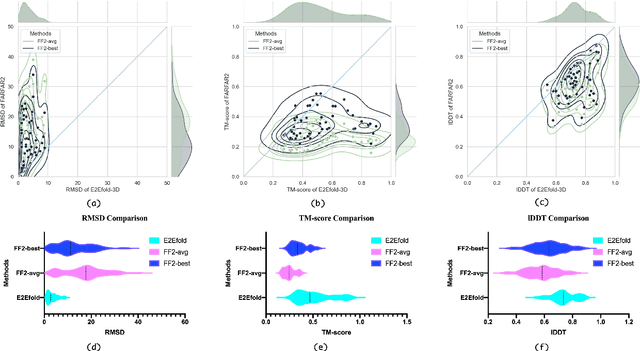
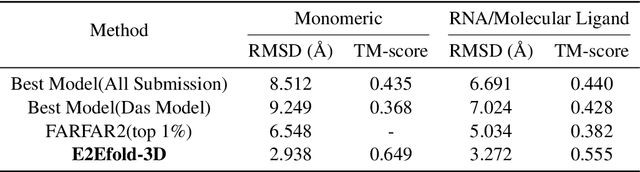
Abstract:RNA structure determination and prediction can promote RNA-targeted drug development and engineerable synthetic elements design. But due to the intrinsic structural flexibility of RNAs, all the three mainstream structure determination methods (X-ray crystallography, NMR, and Cryo-EM) encounter challenges when resolving the RNA structures, which leads to the scarcity of the resolved RNA structures. Computational prediction approaches emerge as complementary to the experimental techniques. However, none of the \textit{de novo} approaches is based on deep learning since too few structures are available. Instead, most of them apply the time-consuming sampling-based strategies, and their performance seems to hit the plateau. In this work, we develop the first end-to-end deep learning approach, E2Efold-3D, to accurately perform the \textit{de novo} RNA structure prediction. Several novel components are proposed to overcome the data scarcity, such as a fully-differentiable end-to-end pipeline, secondary structure-assisted self-distillation, and parameter-efficient backbone formulation. Such designs are validated on the independent, non-overlapping RNA puzzle testing dataset and reach an average sub-4 \AA{} root-mean-square deviation, demonstrating its superior performance compared to state-of-the-art approaches. Interestingly, it also achieves promising results when predicting RNA complex structures, a feat that none of the previous systems could accomplish. When E2Efold-3D is coupled with the experimental techniques, the RNA structure prediction field can be greatly advanced.
Training Data is More Valuable than You Think: A Simple and Effective Method by Retrieving from Training Data
Mar 16, 2022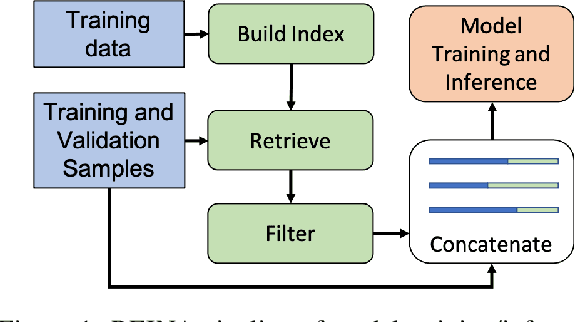
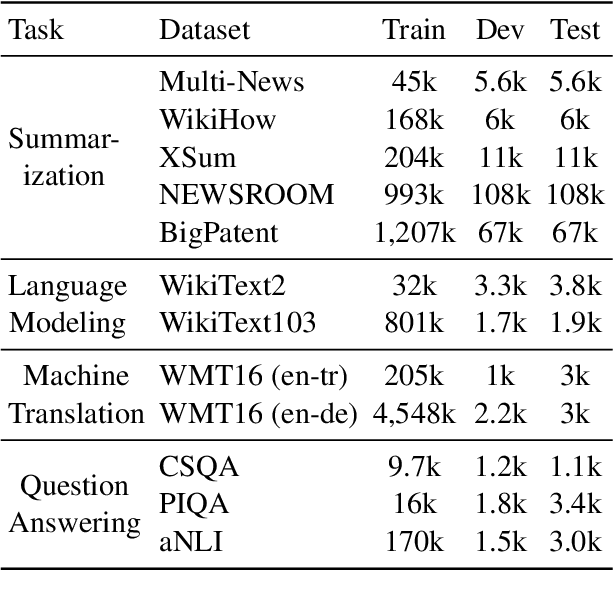
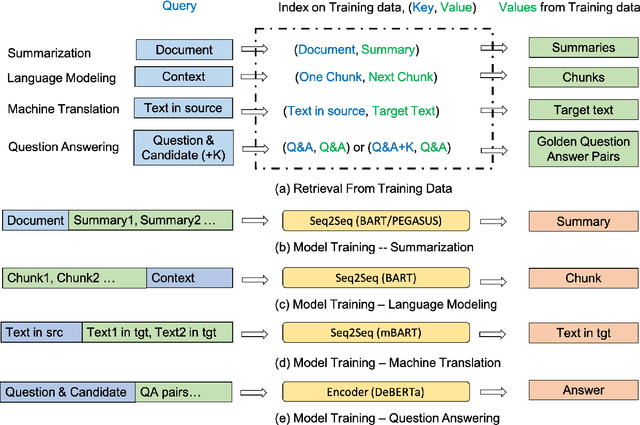
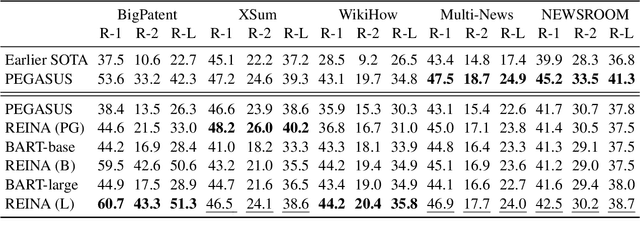
Abstract:Retrieval-based methods have been shown to be effective in NLP tasks via introducing external knowledge. However, the indexing and retrieving of large-scale corpora bring considerable computational cost. Surprisingly, we found that REtrieving from the traINing datA (REINA) only can lead to significant gains on multiple NLG and NLU tasks. We retrieve the labeled training instances most similar to the input text and then concatenate them with the input to feed into the model to generate the output. Experimental results show that this simple method can achieve significantly better performance on a variety of NLU and NLG tasks, including summarization, machine translation, language modeling, and question answering tasks. For instance, our proposed method achieved state-of-the-art results on XSum, BigPatent, and CommonsenseQA. Our code is released, https://github.com/microsoft/REINA .
Human Parity on CommonsenseQA: Augmenting Self-Attention with External Attention
Dec 14, 2021
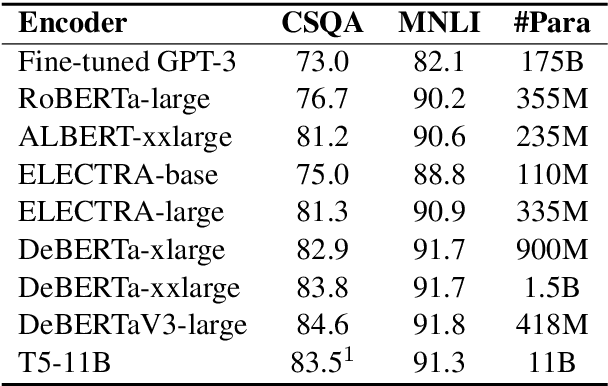
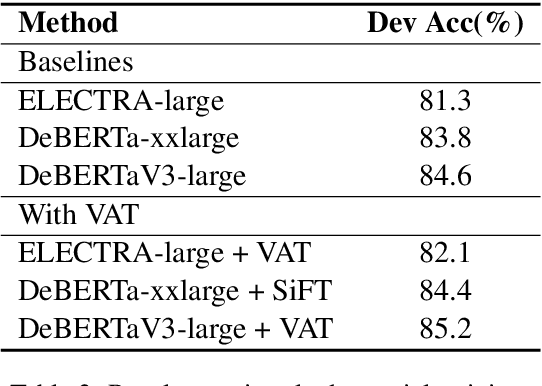

Abstract:Most of today's AI systems focus on using self-attention mechanisms and transformer architectures on large amounts of diverse data to achieve impressive performance gains. In this paper, we propose to augment the transformer architecture with an external attention mechanism to bring external knowledge and context to bear. By integrating external information into the prediction process, we hope to reduce the need for ever-larger models and increase the democratization of AI systems. We find that the proposed external attention mechanism can significantly improve the performance of existing AI systems, allowing practitioners to easily customize foundation AI models to many diverse downstream applications. In particular, we focus on the task of Commonsense Reasoning, demonstrating that the proposed external attention mechanism can augment existing transformer models and significantly improve the model's reasoning capabilities. The proposed system, Knowledgeable External Attention for commonsense Reasoning (KEAR), reaches human parity on the open CommonsenseQA research benchmark with an accuracy of 89.4\% in comparison to the human accuracy of 88.9\%.
Leveraging Knowledge in Multilingual Commonsense Reasoning
Oct 16, 2021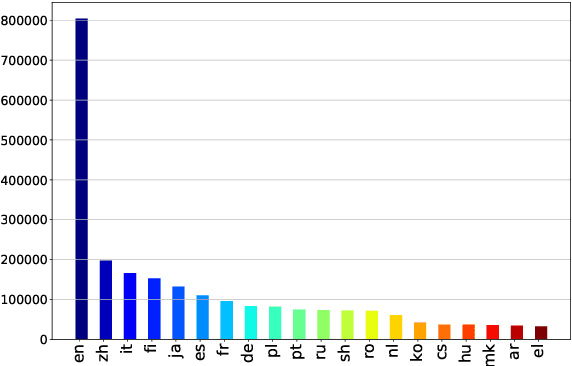

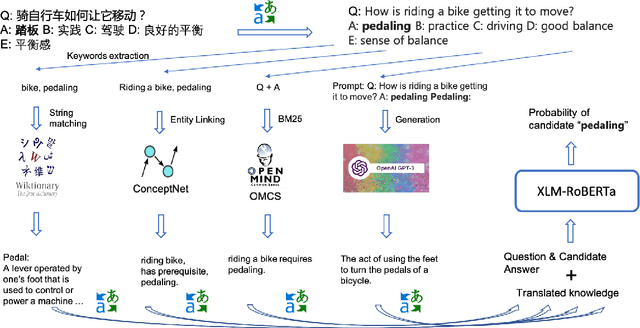
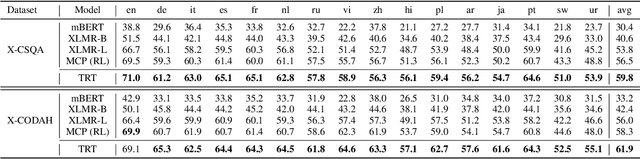
Abstract:Commonsense reasoning (CSR) requires the model to be equipped with general world knowledge. While CSR is a language-agnostic process, most comprehensive knowledge sources are in few popular languages, especially English. Thus, it remains unclear how to effectively conduct multilingual commonsense reasoning (XCSR) for various languages. In this work, we propose to utilize English knowledge sources via a translate-retrieve-translate (TRT) strategy. For multilingual commonsense questions and choices, we collect related knowledge via translation and retrieval from the knowledge sources. The retrieved knowledge is then translated into the target language and integrated into a pre-trained multilingual language model via visible knowledge attention. Then we utilize a diverse of 4 English knowledge sources to provide more comprehensive coverage of knowledge in different formats. Extensive results on the XCSR benchmark demonstrate that TRT with external knowledge can significantly improve multilingual commonsense reasoning in both zero-shot and translate-train settings, outperforming 3.3 and 3.6 points over the previous state-of-the-art on XCSR benchmark datasets (X-CSQA and X-CODAH).
Joint Retrieval and Generation Training for Grounded Text Generation
Jun 03, 2021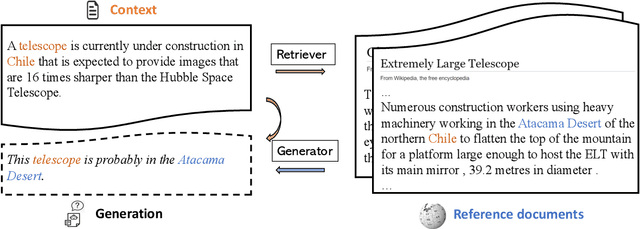
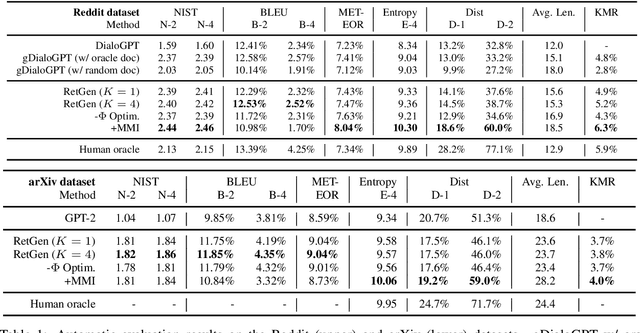
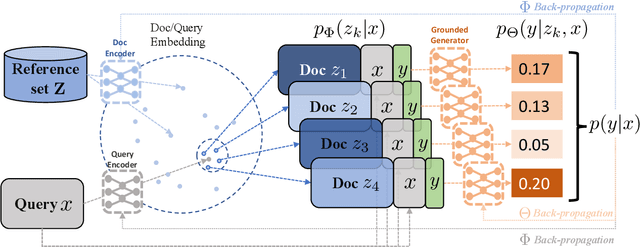

Abstract:Recent advances in large-scale pre-training such as GPT-3 allow seemingly high quality text to be generated from a given prompt. However, such generation systems often suffer from problems of hallucinated facts, and are not inherently designed to incorporate useful external information. Grounded generation models appear to offer remedies, but their training typically relies on rarely-available parallel data where corresponding information-relevant documents are provided for context. We propose a framework that alleviates this data constraint by jointly training a grounded generator and document retriever on the language model signal. The model learns to reward retrieval of the documents with the highest utility in generation, and attentively combines them using a Mixture-of-Experts (MoE) ensemble to generate follow-on text. We demonstrate that both generator and retriever can take advantage of this joint training and work synergistically to produce more informative and relevant text in both prose and dialogue generation.
LightningDOT: Pre-training Visual-Semantic Embeddings for Real-Time Image-Text Retrieval
Apr 11, 2021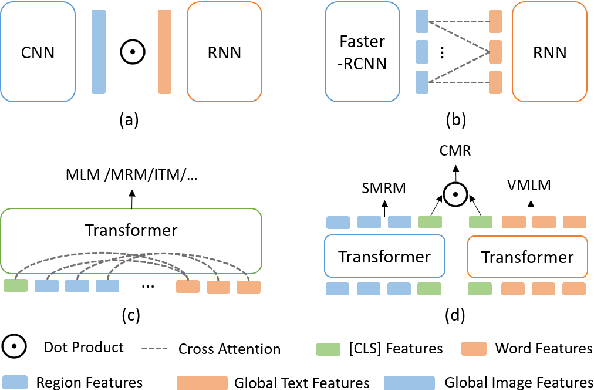
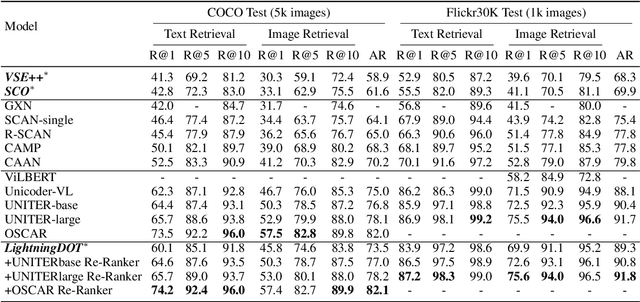
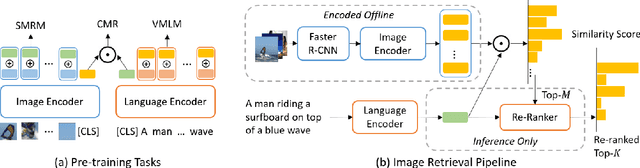
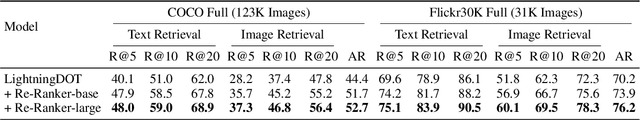
Abstract:Multimodal pre-training has propelled great advancement in vision-and-language research. These large-scale pre-trained models, although successful, fatefully suffer from slow inference speed due to enormous computation cost mainly from cross-modal attention in Transformer architecture. When applied to real-life applications, such latency and computation demand severely deter the practical use of pre-trained models. In this paper, we study Image-text retrieval (ITR), the most mature scenario of V+L application, which has been widely studied even prior to the emergence of recent pre-trained models. We propose a simple yet highly effective approach, LightningDOT that accelerates the inference time of ITR by thousands of times, without sacrificing accuracy. LightningDOT removes the time-consuming cross-modal attention by pre-training on three novel learning objectives, extracting feature indexes offline, and employing instant dot-product matching with further re-ranking, which significantly speeds up retrieval process. In fact, LightningDOT achieves new state of the art across multiple ITR benchmarks such as Flickr30k, COCO and Multi30K, outperforming existing pre-trained models that consume 1000x magnitude of computational hours. Code and pre-training checkpoints are available at https://github.com/intersun/LightningDOT.
Cross-Thought for Sentence Encoder Pre-training
Oct 07, 2020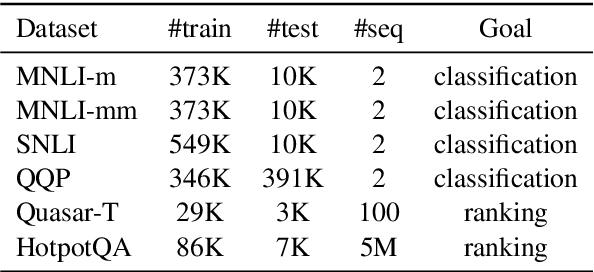
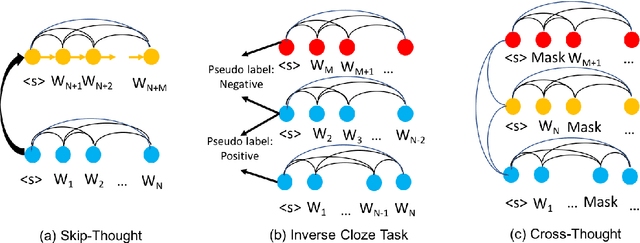
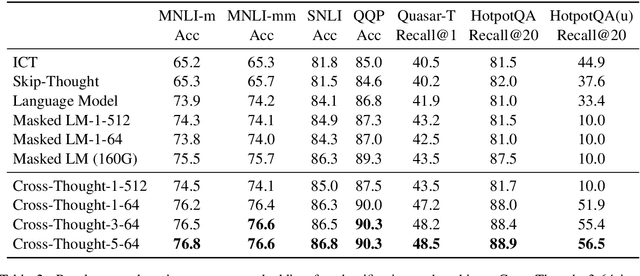
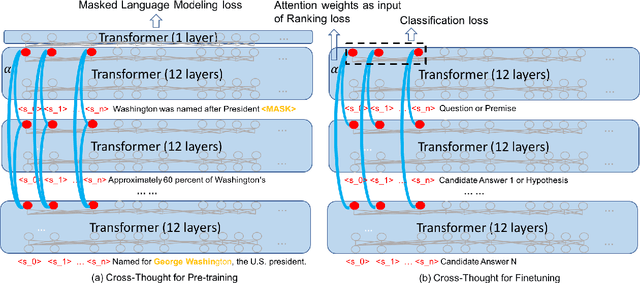
Abstract:In this paper, we propose Cross-Thought, a novel approach to pre-training sequence encoder, which is instrumental in building reusable sequence embeddings for large-scale NLP tasks such as question answering. Instead of using the original signals of full sentences, we train a Transformer-based sequence encoder over a large set of short sequences, which allows the model to automatically select the most useful information for predicting masked words. Experiments on question answering and textual entailment tasks demonstrate that our pre-trained encoder can outperform state-of-the-art encoders trained with continuous sentence signals as well as traditional masked language modeling baselines. Our proposed approach also achieves new state of the art on HotpotQA (full-wiki setting) by improving intermediate information retrieval performance.
Contrastive Distillation on Intermediate Representations for Language Model Compression
Sep 29, 2020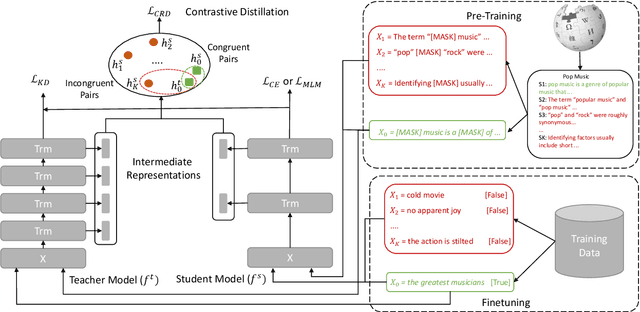
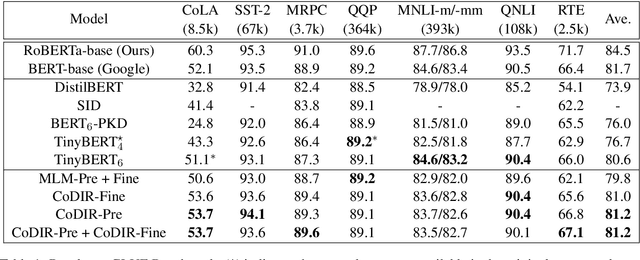
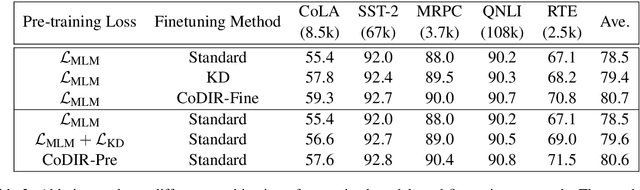
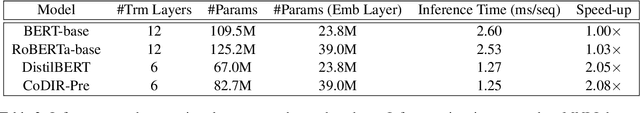
Abstract:Existing language model compression methods mostly use a simple L2 loss to distill knowledge in the intermediate representations of a large BERT model to a smaller one. Although widely used, this objective by design assumes that all the dimensions of hidden representations are independent, failing to capture important structural knowledge in the intermediate layers of the teacher network. To achieve better distillation efficacy, we propose Contrastive Distillation on Intermediate Representations (CoDIR), a principled knowledge distillation framework where the student is trained to distill knowledge through intermediate layers of the teacher via a contrastive objective. By learning to distinguish positive sample from a large set of negative samples, CoDIR facilitates the student's exploitation of rich information in teacher's hidden layers. CoDIR can be readily applied to compress large-scale language models in both pre-training and finetuning stages, and achieves superb performance on the GLUE benchmark, outperforming state-of-the-art compression methods.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge