Yihang Zhou
Rooftop Wind Field Reconstruction Using Sparse Sensors: From Deterministic to Generative Learning Methods
Mar 13, 2026Abstract:Real-time rooftop wind-speed distribution is important for the safe operation of drones and urban air mobility systems, wind control systems, and rooftop utilization. However, rooftop flows show strong nonlinearity, separation, and cross-direction variability, which make flow field reconstruction from sparse sensors difficult. This study develops a learning-from-observation framework using wind-tunnel experimental data obtained by Particle Image Velocimetry (PIV) and compares Kriging interpolation with three deep learning models: UNet, Vision Transformer Autoencoder (ViTAE), and Conditional Wasserstein GAN (CWGAN). We evaluate two training strategies, single wind-direction training (SDT) and mixed wind-direction training (MDT), across sensor densities from 5 to 30, test robustness under sensor position perturbations of plus or minus 1 grid, and optimize sensor placement via Proper Orthogonal Decomposition with QR decomposition. Results show that deep learning methods can reconstruct rooftop wind fields from sparse sensor data effectively. Compared with Kriging interpolation, the deep learning models improved SSIM by up to 32.7%, FAC2 by 24.2%, and NMSE by 27.8%. Mixed wind-direction training further improved performance, with gains of up to 173.7% in SSIM, 16.7% in FAC2, and 98.3% in MG compared with single-direction training. The results also show that sensor configuration, optimization, and training strategy should be considered jointly for reliable deployment. QR-based optimization improved robustness by up to 27.8% under sensor perturbations, although with metric-dependent trade-offs. Training on experimental rather than simulated data also provides practical guidance for method selection and sensor placement in different scenarios.
Contact-Anchored Policies: Contact Conditioning Creates Strong Robot Utility Models
Feb 09, 2026Abstract:The prevalent paradigm in robot learning attempts to generalize across environments, embodiments, and tasks with language prompts at runtime. A fundamental tension limits this approach: language is often too abstract to guide the concrete physical understanding required for robust manipulation. In this work, we introduce Contact-Anchored Policies (CAP), which replace language conditioning with points of physical contact in space. Simultaneously, we structure CAP as a library of modular utility models rather than a monolithic generalist policy. This factorization allows us to implement a real-to-sim iteration cycle: we build EgoGym, a lightweight simulation benchmark, to rapidly identify failure modes and refine our models and datasets prior to real-world deployment. We show that by conditioning on contact and iterating via simulation, CAP generalizes to novel environments and embodiments out of the box on three fundamental manipulation skills while using only 23 hours of demonstration data, and outperforms large, state-of-the-art VLAs in zero-shot evaluations by 56%. All model checkpoints, codebase, hardware, simulation, and datasets will be open-sourced. Project page: https://cap-policy.github.io/
Extreme Value Policy Optimization for Safe Reinforcement Learning
Jan 17, 2026Abstract:Ensuring safety is a critical challenge in applying Reinforcement Learning (RL) to real-world scenarios. Constrained Reinforcement Learning (CRL) addresses this by maximizing returns under predefined constraints, typically formulated as the expected cumulative cost. However, expectation-based constraints overlook rare but high-impact extreme value events in the tail distribution, such as black swan incidents, which can lead to severe constraint violations. To address this issue, we propose the Extreme Value policy Optimization (EVO) algorithm, leveraging Extreme Value Theory (EVT) to model and exploit extreme reward and cost samples, reducing constraint violations. EVO introduces an extreme quantile optimization objective to explicitly capture extreme samples in the cost tail distribution. Additionally, we propose an extreme prioritization mechanism during replay, amplifying the learning signal from rare but high-impact extreme samples. Theoretically, we establish upper bounds on expected constraint violations during policy updates, guaranteeing strict constraint satisfaction at a zero-violation quantile level. Further, we demonstrate that EVO achieves a lower probability of constraint violations than expectation-based methods and exhibits lower variance than quantile regression methods. Extensive experiments show that EVO significantly reduces constraint violations during training while maintaining competitive policy performance compared to baselines.
NeuroSwift: A Lightweight Cross-Subject Framework for fMRI Visual Reconstruction of Complex Scenes
Oct 02, 2025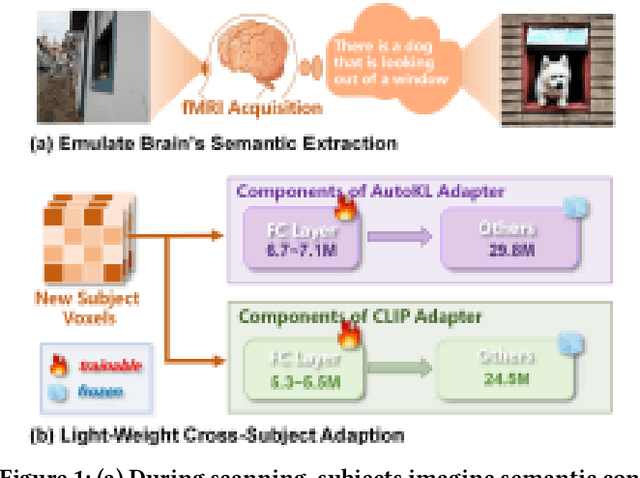
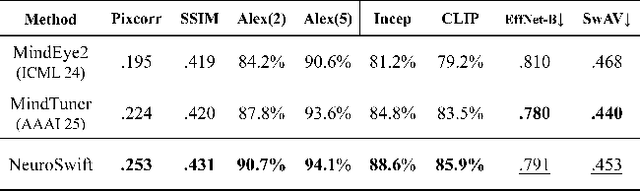
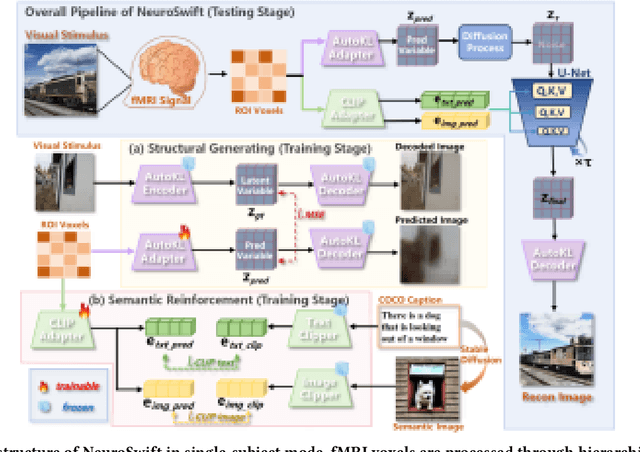
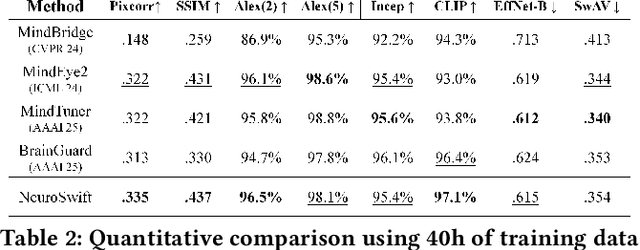
Abstract:Reconstructing visual information from brain activity via computer vision technology provides an intuitive understanding of visual neural mechanisms. Despite progress in decoding fMRI data with generative models, achieving accurate cross-subject reconstruction of visual stimuli remains challenging and computationally demanding. This difficulty arises from inter-subject variability in neural representations and the brain's abstract encoding of core semantic features in complex visual inputs. To address these challenges, we propose NeuroSwift, which integrates complementary adapters via diffusion: AutoKL for low-level features and CLIP for semantics. NeuroSwift's CLIP Adapter is trained on Stable Diffusion generated images paired with COCO captions to emulate higher visual cortex encoding. For cross-subject generalization, we pretrain on one subject and then fine-tune only 17 percent of parameters (fully connected layers) for new subjects, while freezing other components. This enables state-of-the-art performance with only one hour of training per subject on lightweight GPUs (three RTX 4090), and it outperforms existing methods.
HAVIR: HierArchical Vision to Image Reconstruction using CLIP-Guided Versatile Diffusion
Jun 06, 2025Abstract:Reconstructing visual information from brain activity bridges the gap between neuroscience and computer vision. Even though progress has been made in decoding images from fMRI using generative models, a challenge remains in accurately recovering highly complex visual stimuli. This difficulty stems from their elemental density and diversity, sophisticated spatial structures, and multifaceted semantic information. To address these challenges, we propose HAVIR that contains two adapters: (1) The AutoKL Adapter transforms fMRI voxels into a latent diffusion prior, capturing topological structures; (2) The CLIP Adapter converts the voxels to CLIP text and image embeddings, containing semantic information. These complementary representations are fused by Versatile Diffusion to generate the final reconstructed image. To extract the most essential semantic information from complex scenarios, the CLIP Adapter is trained with text captions describing the visual stimuli and their corresponding semantic images synthesized from these captions. The experimental results demonstrate that HAVIR effectively reconstructs both structural features and semantic information of visual stimuli even in complex scenarios, outperforming existing models.
Comparative and Interpretative Analysis of CNN and Transformer Models in Predicting Wildfire Spread Using Remote Sensing Data
Mar 18, 2025



Abstract:Facing the escalating threat of global wildfires, numerous computer vision techniques using remote sensing data have been applied in this area. However, the selection of deep learning methods for wildfire prediction remains uncertain due to the lack of comparative analysis in a quantitative and explainable manner, crucial for improving prevention measures and refining models. This study aims to thoroughly compare the performance, efficiency, and explainability of four prevalent deep learning architectures: Autoencoder, ResNet, UNet, and Transformer-based Swin-UNet. Employing a real-world dataset that includes nearly a decade of remote sensing data from California, U.S., these models predict the spread of wildfires for the following day. Through detailed quantitative comparison analysis, we discovered that Transformer-based Swin-UNet and UNet generally outperform Autoencoder and ResNet, particularly due to the advanced attention mechanisms in Transformer-based Swin-UNet and the efficient use of skip connections in both UNet and Transformer-based Swin-UNet, which contribute to superior predictive accuracy and model interpretability. Then we applied XAI techniques on all four models, this not only enhances the clarity and trustworthiness of models but also promotes focused improvements in wildfire prediction capabilities. The XAI analysis reveals that UNet and Transformer-based Swin-UNet are able to focus on critical features such as 'Previous Fire Mask', 'Drought', and 'Vegetation' more effectively than the other two models, while also maintaining balanced attention to the remaining features, leading to their superior performance. The insights from our thorough comparative analysis offer substantial implications for future model design and also provide guidance for model selection in different scenarios.
K-space Diffusion Model Based MR Reconstruction Method for Simultaneous Multislice Imaging
Jan 06, 2025Abstract:Simultaneous Multi-Slice(SMS) is a magnetic resonance imaging (MRI) technique which excites several slices concurrently using multiband radiofrequency pulses to reduce scanning time. However, due to its variable data structure and difficulty in acquisition, it is challenging to integrate SMS data as training data into deep learning frameworks.This study proposed a novel k-space diffusion model of SMS reconstruction that does not utilize SMS data for training. Instead, it incorporates Slice GRAPPA during the sampling process to reconstruct SMS data from different acquisition modes.Our results demonstrated that this method outperforms traditional SMS reconstruction methods and can achieve higher acceleration factors without in-plane aliasing.
Image Synthesis with Class-Aware Semantic Diffusion Models for Surgical Scene Segmentation
Oct 31, 2024



Abstract:Surgical scene segmentation is essential for enhancing surgical precision, yet it is frequently compromised by the scarcity and imbalance of available data. To address these challenges, semantic image synthesis methods based on generative adversarial networks and diffusion models have been developed. However, these models often yield non-diverse images and fail to capture small, critical tissue classes, limiting their effectiveness. In response, we propose the Class-Aware Semantic Diffusion Model (CASDM), a novel approach which utilizes segmentation maps as conditions for image synthesis to tackle data scarcity and imbalance. Novel class-aware mean squared error and class-aware self-perceptual loss functions have been defined to prioritize critical, less visible classes, thereby enhancing image quality and relevance. Furthermore, to our knowledge, we are the first to generate multi-class segmentation maps using text prompts in a novel fashion to specify their contents. These maps are then used by CASDM to generate surgical scene images, enhancing datasets for training and validating segmentation models. Our evaluation, which assesses both image quality and downstream segmentation performance, demonstrates the strong effectiveness and generalisability of CASDM in producing realistic image-map pairs, significantly advancing surgical scene segmentation across diverse and challenging datasets.
Quantum Neural Network for Accelerated Magnetic Resonance Imaging
Oct 12, 2024



Abstract:Magnetic resonance image reconstruction starting from undersampled k-space data requires the recovery of many potential nonlinear features, which is very difficult for algorithms to recover these features. In recent years, the development of quantum computing has discovered that quantum convolution can improve network accuracy, possibly due to potential quantum advantages. This article proposes a hybrid neural network containing quantum and classical networks for fast magnetic resonance imaging, and conducts experiments on a quantum computer simulation system. The experimental results indicate that the hybrid network has achieved excellent reconstruction results, and also confirm the feasibility of applying hybrid quantum-classical neural networks into the image reconstruction of rapid magnetic resonance imaging.
Optimized Magnetic Resonance Fingerprinting Using Ziv-Zakai Bound
Oct 10, 2024



Abstract:Magnetic Resonance Fingerprinting (MRF) has emerged as a promising quantitative imaging technique within the field of Magnetic Resonance Imaging (MRI), offers comprehensive insights into tissue properties by simultaneously acquiring multiple tissue parameter maps in a single acquisition. Sequence optimization is crucial for improving the accuracy and efficiency of MRF. In this work, a novel framework for MRF sequence optimization is proposed based on the Ziv-Zakai bound (ZZB). Unlike the Cram\'er-Rao bound (CRB), which aims to enhance the quality of a single fingerprint signal with deterministic parameters, ZZB provides insights into evaluating the minimum mismatch probability for pairs of fingerprint signals within the specified parameter range in MRF. Specifically, the explicit ZZB is derived to establish a lower bound for the discrimination error in the fingerprint signal matching process within MRF. This bound illuminates the intrinsic limitations of MRF sequences, thereby fostering a deeper understanding of existing sequence performance. Subsequently, an optimal experiment design problem based on ZZB was formulated to ascertain the optimal scheme of acquisition parameters, maximizing discrimination power of MRF between different tissue types. Preliminary numerical experiments show that the optimized ZZB scheme outperforms both the conventional and CRB schemes in terms of the reconstruction accuracy of multiple parameter maps.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge