Pheng Ann Heng
Federated Learning Enables Big Data for Rare Cancer Boundary Detection
Apr 25, 2022Abstract:Although machine learning (ML) has shown promise in numerous domains, there are concerns about generalizability to out-of-sample data. This is currently addressed by centrally sharing ample, and importantly diverse, data from multiple sites. However, such centralization is challenging to scale (or even not feasible) due to various limitations. Federated ML (FL) provides an alternative to train accurate and generalizable ML models, by only sharing numerical model updates. Here we present findings from the largest FL study to-date, involving data from 71 healthcare institutions across 6 continents, to generate an automatic tumor boundary detector for the rare disease of glioblastoma, utilizing the largest dataset of such patients ever used in the literature (25,256 MRI scans from 6,314 patients). We demonstrate a 33% improvement over a publicly trained model to delineate the surgically targetable tumor, and 23% improvement over the tumor's entire extent. We anticipate our study to: 1) enable more studies in healthcare informed by large and diverse data, ensuring meaningful results for rare diseases and underrepresented populations, 2) facilitate further quantitative analyses for glioblastoma via performance optimization of our consensus model for eventual public release, and 3) demonstrate the effectiveness of FL at such scale and task complexity as a paradigm shift for multi-site collaborations, alleviating the need for data sharing.
LHNN: Lattice Hypergraph Neural Network for VLSI Congestion Prediction
Mar 24, 2022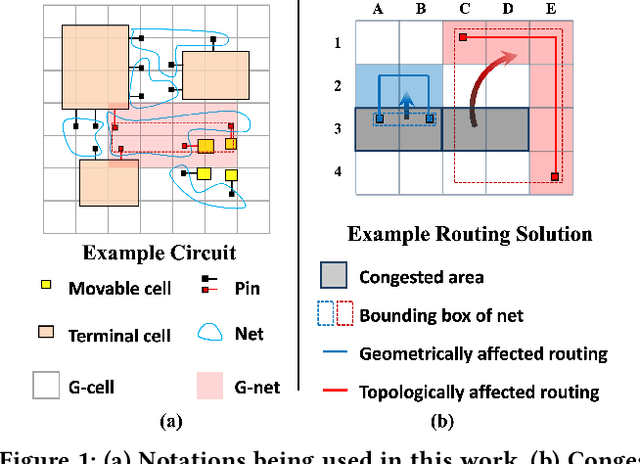
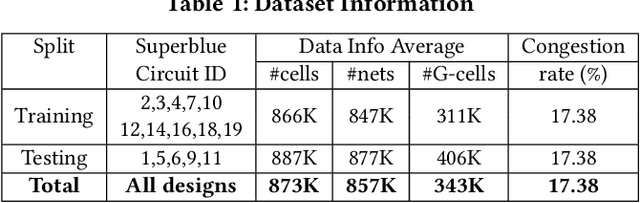
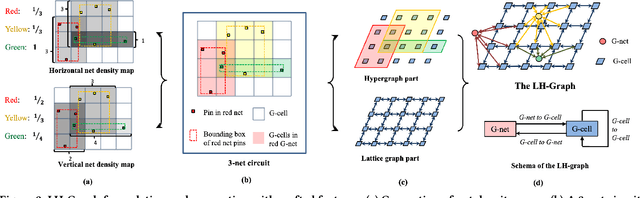
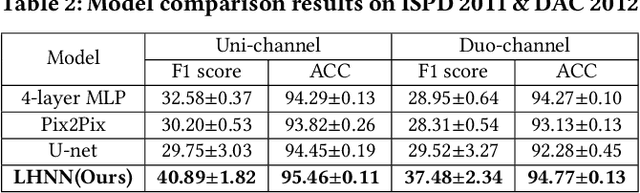
Abstract:Precise congestion prediction from a placement solution plays a crucial role in circuit placement. This work proposes the lattice hypergraph (LH-graph), a novel graph formulation for circuits, which preserves netlist data during the whole learning process, and enables the congestion information propagated geometrically and topologically. Based on the formulation, we further developed a heterogeneous graph neural network architecture LHNN, jointing the routing demand regression to support the congestion spot classification. LHNN constantly achieves more than 35% improvements compared with U-nets and Pix2Pix on the F1 score. We expect our work shall highlight essential procedures using machine learning for congestion prediction.
Comparative Validation of Machine Learning Algorithms for Surgical Workflow and Skill Analysis with the HeiChole Benchmark
Sep 30, 2021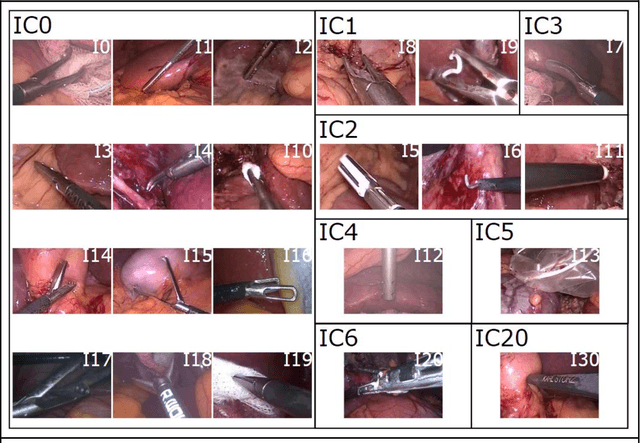
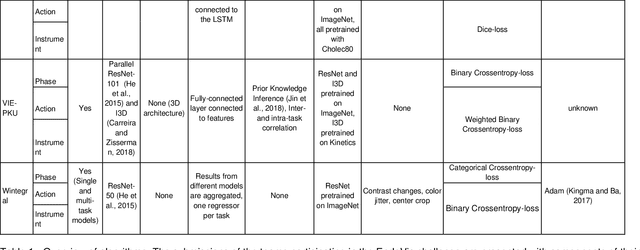
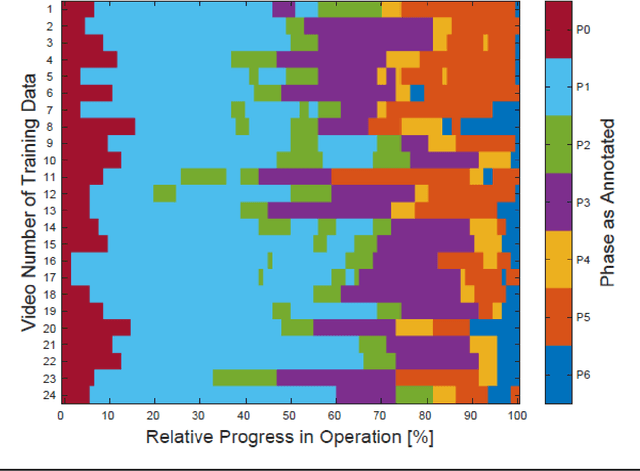
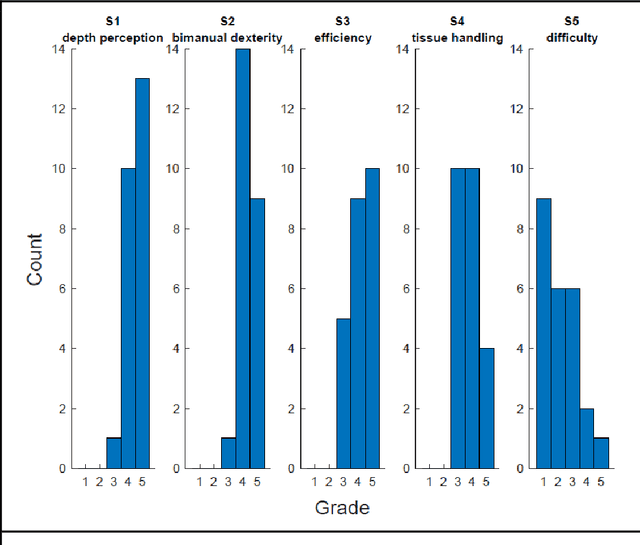
Abstract:PURPOSE: Surgical workflow and skill analysis are key technologies for the next generation of cognitive surgical assistance systems. These systems could increase the safety of the operation through context-sensitive warnings and semi-autonomous robotic assistance or improve training of surgeons via data-driven feedback. In surgical workflow analysis up to 91% average precision has been reported for phase recognition on an open data single-center dataset. In this work we investigated the generalizability of phase recognition algorithms in a multi-center setting including more difficult recognition tasks such as surgical action and surgical skill. METHODS: To achieve this goal, a dataset with 33 laparoscopic cholecystectomy videos from three surgical centers with a total operation time of 22 hours was created. Labels included annotation of seven surgical phases with 250 phase transitions, 5514 occurences of four surgical actions, 6980 occurences of 21 surgical instruments from seven instrument categories and 495 skill classifications in five skill dimensions. The dataset was used in the 2019 Endoscopic Vision challenge, sub-challenge for surgical workflow and skill analysis. Here, 12 teams submitted their machine learning algorithms for recognition of phase, action, instrument and/or skill assessment. RESULTS: F1-scores were achieved for phase recognition between 23.9% and 67.7% (n=9 teams), for instrument presence detection between 38.5% and 63.8% (n=8 teams), but for action recognition only between 21.8% and 23.3% (n=5 teams). The average absolute error for skill assessment was 0.78 (n=1 team). CONCLUSION: Surgical workflow and skill analysis are promising technologies to support the surgical team, but are not solved yet, as shown by our comparison of algorithms. This novel benchmark can be used for comparable evaluation and validation of future work.
Balancing Accuracy and Fairness for Interactive Recommendation with Reinforcement Learning
Jun 25, 2021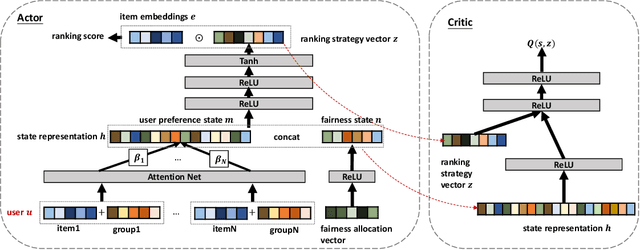
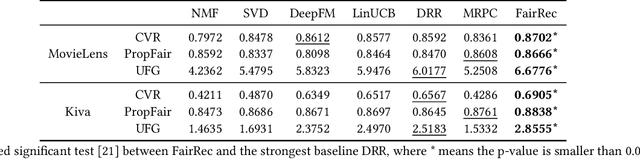
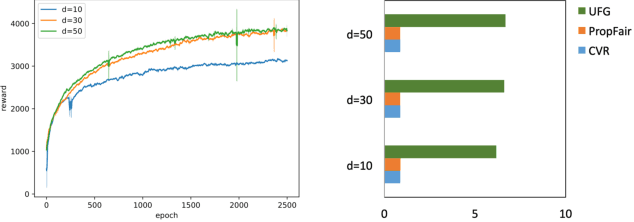
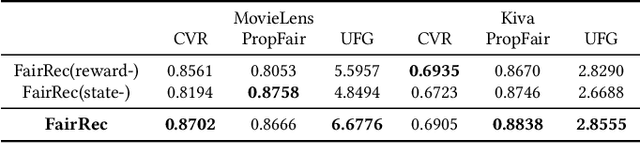
Abstract:Fairness in recommendation has attracted increasing attention due to bias and discrimination possibly caused by traditional recommenders. In Interactive Recommender Systems (IRS), user preferences and the system's fairness status are constantly changing over time. Existing fairness-aware recommenders mainly consider fairness in static settings. Directly applying existing methods to IRS will result in poor recommendation. To resolve this problem, we propose a reinforcement learning based framework, FairRec, to dynamically maintain a long-term balance between accuracy and fairness in IRS. User preferences and the system's fairness status are jointly compressed into the state representation to generate recommendations. FairRec aims at maximizing our designed cumulative reward that combines accuracy and fairness. Extensive experiments validate that FairRec can improve fairness, while preserving good recommendation quality.
Cascaded Robust Learning at Imperfect Labels for Chest X-ray Segmentation
Apr 05, 2021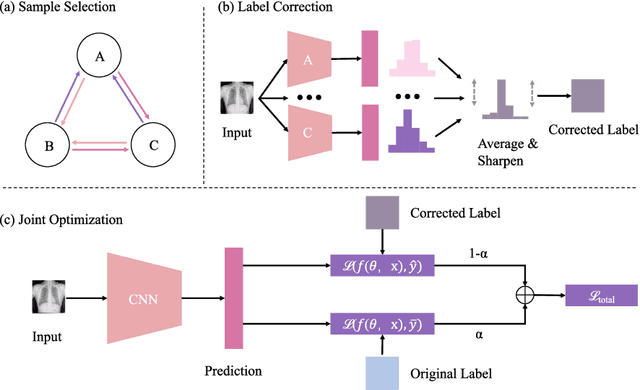
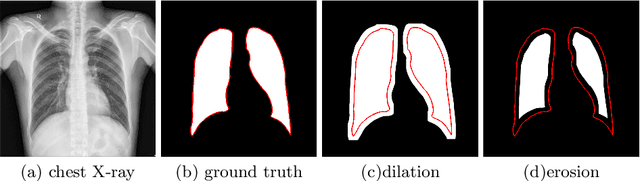
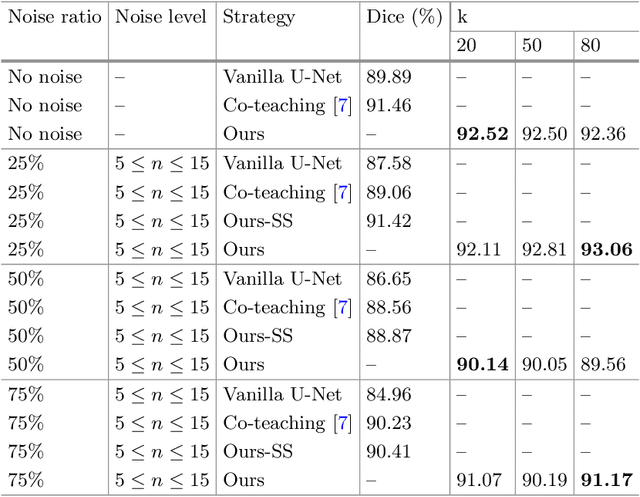

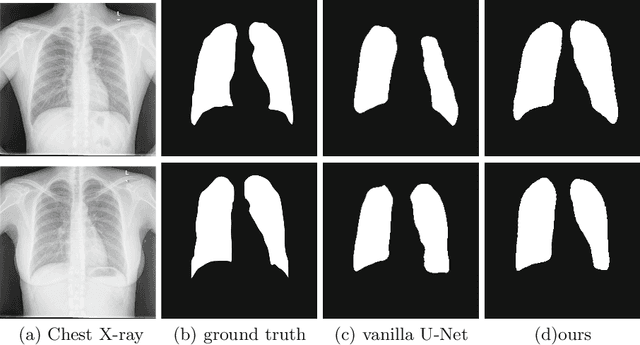
Abstract:The superior performance of CNN on medical image analysis heavily depends on the annotation quality, such as the number of labeled image, the source of image, and the expert experience. The annotation requires great expertise and labour. To deal with the high inter-rater variability, the study of imperfect label has great significance in medical image segmentation tasks. In this paper, we present a novel cascaded robust learning framework for chest X-ray segmentation with imperfect annotation. Our model consists of three independent network, which can effectively learn useful information from the peer networks. The framework includes two stages. In the first stage, we select the clean annotated samples via a model committee setting, the networks are trained by minimizing a segmentation loss using the selected clean samples. In the second stage, we design a joint optimization framework with label correction to gradually correct the wrong annotation and improve the network performance. We conduct experiments on the public chest X-ray image datasets collected by Shenzhen Hospital. The results show that our methods could achieve a significant improvement on the accuracy in segmentation tasks compared to the previous methods.
Global Guidance Network for Breast Lesion Segmentation in Ultrasound Images
Apr 05, 2021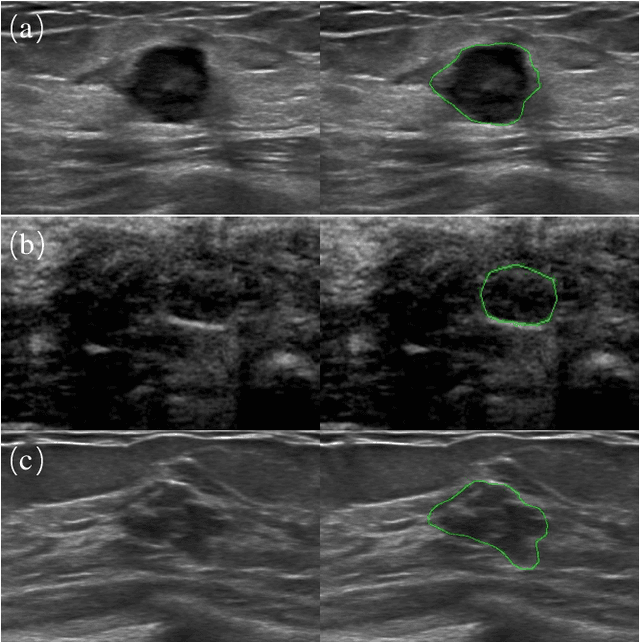
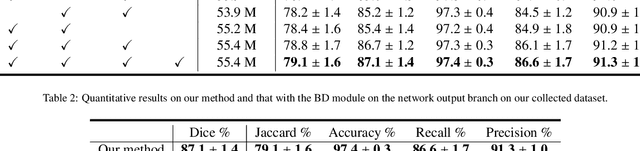
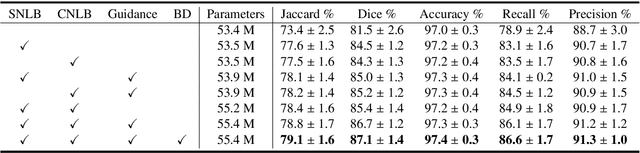
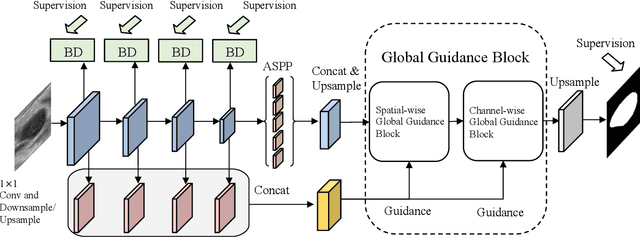

Abstract:Automatic breast lesion segmentation in ultrasound helps to diagnose breast cancer, which is one of the dreadful diseases that affect women globally. Segmenting breast regions accurately from ultrasound image is a challenging task due to the inherent speckle artifacts, blurry breast lesion boundaries, and inhomogeneous intensity distributions inside the breast lesion regions. Recently, convolutional neural networks (CNNs) have demonstrated remarkable results in medical image segmentation tasks. However, the convolutional operations in a CNN often focus on local regions, which suffer from limited capabilities in capturing long-range dependencies of the input ultrasound image, resulting in degraded breast lesion segmentation accuracy. In this paper, we develop a deep convolutional neural network equipped with a global guidance block (GGB) and breast lesion boundary detection (BD) modules for boosting the breast ultrasound lesion segmentation. The GGB utilizes the multi-layer integrated feature map as a guidance information to learn the long-range non-local dependencies from both spatial and channel domains. The BD modules learn additional breast lesion boundary map to enhance the boundary quality of a segmentation result refinement. Experimental results on a public dataset and a collected dataset show that our network outperforms other medical image segmentation methods and the recent semantic segmentation methods on breast ultrasound lesion segmentation. Moreover, we also show the application of our network on the ultrasound prostate segmentation, in which our method better identifies prostate regions than state-of-the-art networks.
Semi-supervised Medical Image Classification with Relation-driven Self-ensembling Model
May 15, 2020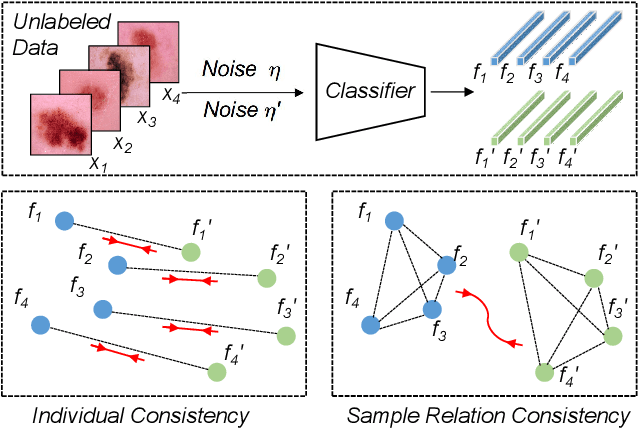
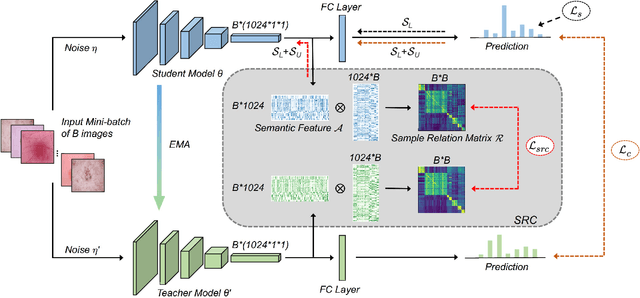
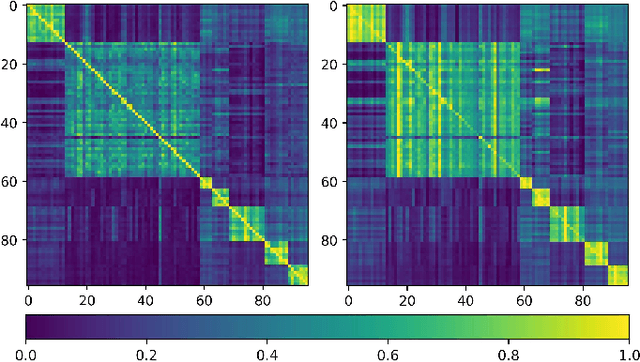
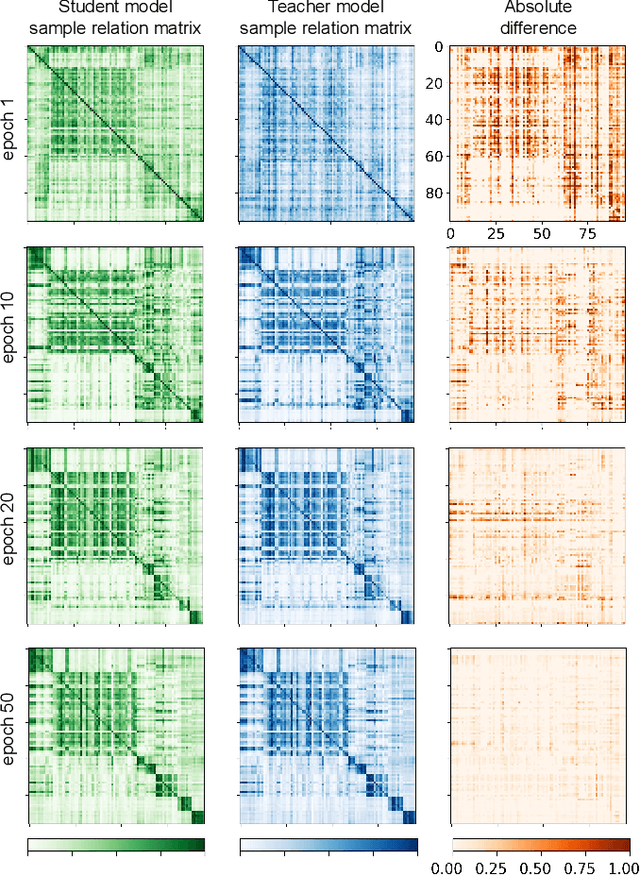
Abstract:Training deep neural networks usually requires a large amount of labeled data to obtain good performance. However, in medical image analysis, obtaining high-quality labels for the data is laborious and expensive, as accurately annotating medical images demands expertise knowledge of the clinicians. In this paper, we present a novel relation-driven semi-supervised framework for medical image classification. It is a consistency-based method which exploits the unlabeled data by encouraging the prediction consistency of given input under perturbations, and leverages a self-ensembling model to produce high-quality consistency targets for the unlabeled data. Considering that human diagnosis often refers to previous analogous cases to make reliable decisions, we introduce a novel sample relation consistency (SRC) paradigm to effectively exploit unlabeled data by modeling the relationship information among different samples. Superior to existing consistency-based methods which simply enforce consistency of individual predictions, our framework explicitly enforces the consistency of semantic relation among different samples under perturbations, encouraging the model to explore extra semantic information from unlabeled data. We have conducted extensive experiments to evaluate our method on two public benchmark medical image classification datasets, i.e.,skin lesion diagnosis with ISIC 2018 challenge and thorax disease classification with ChestX-ray14. Our method outperforms many state-of-the-art semi-supervised learning methods on both single-label and multi-label image classification scenarios.
MS-Net: Multi-Site Network for Improving Prostate Segmentation with Heterogeneous MRI Data
Feb 19, 2020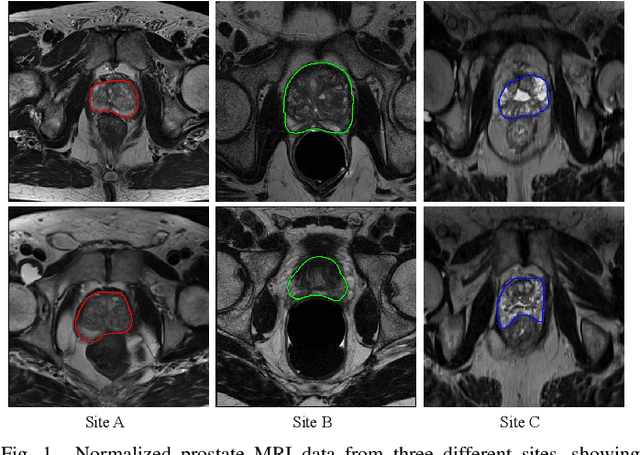
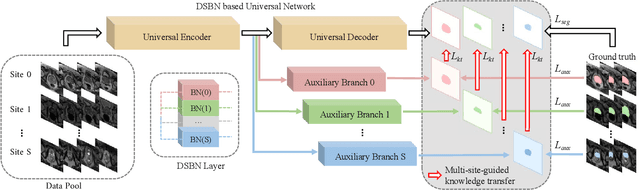
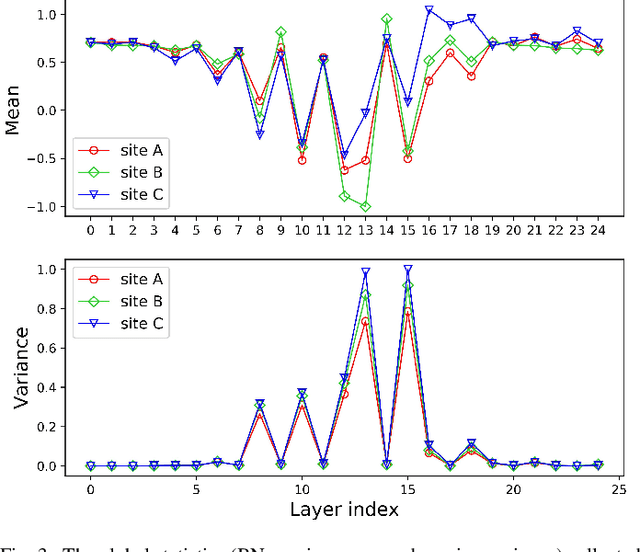
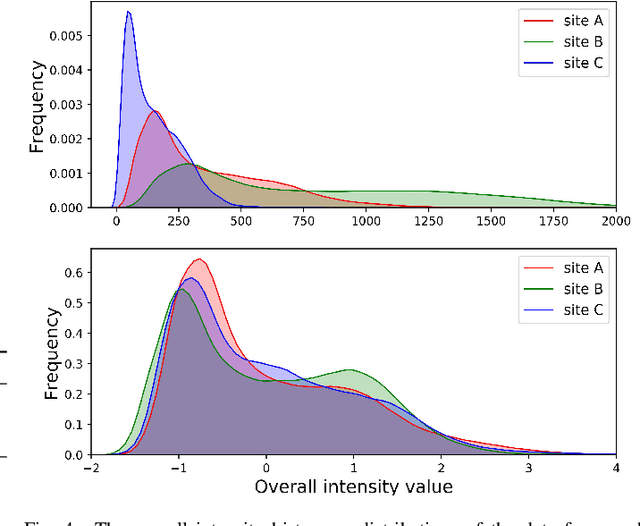
Abstract:Automated prostate segmentation in MRI is highly demanded for computer-assisted diagnosis. Recently, a variety of deep learning methods have achieved remarkable progress in this task, usually relying on large amounts of training data. Due to the nature of scarcity for medical images, it is important to effectively aggregate data from multiple sites for robust model training, to alleviate the insufficiency of single-site samples. However, the prostate MRIs from different sites present heterogeneity due to the differences in scanners and imaging protocols, raising challenges for effective ways of aggregating multi-site data for network training. In this paper, we propose a novel multi-site network (MS-Net) for improving prostate segmentation by learning robust representations, leveraging multiple sources of data. To compensate for the inter-site heterogeneity of different MRI datasets, we develop Domain-Specific Batch Normalization layers in the network backbone, enabling the network to estimate statistics and perform feature normalization for each site separately. Considering the difficulty of capturing the shared knowledge from multiple datasets, a novel learning paradigm, i.e., Multi-site-guided Knowledge Transfer, is proposed to enhance the kernels to extract more generic representations from multi-site data. Extensive experiments on three heterogeneous prostate MRI datasets demonstrate that our MS-Net improves the performance across all datasets consistently, and outperforms state-of-the-art methods for multi-site learning.
Unsupervised Bidirectional Cross-Modality Adaptation via Deeply Synergistic Image and Feature Alignment for Medical Image Segmentation
Feb 06, 2020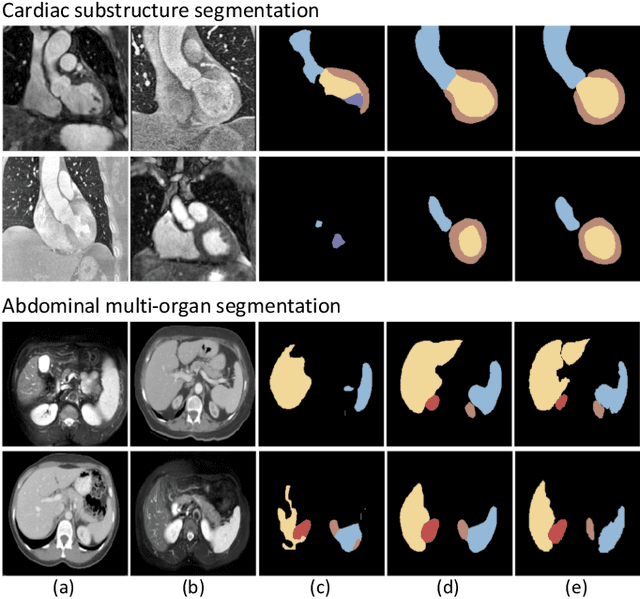
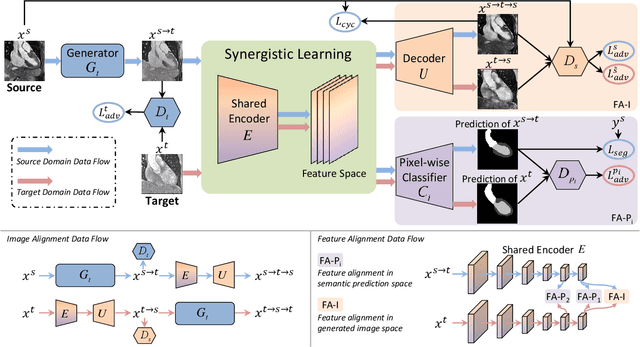
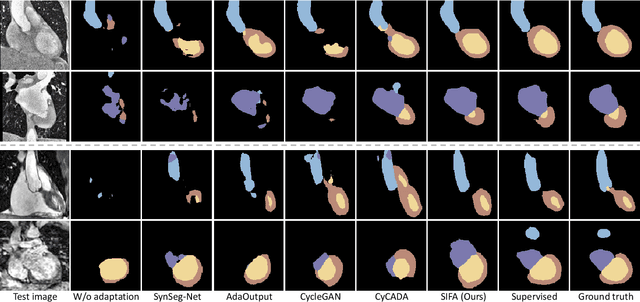

Abstract:Unsupervised domain adaptation has increasingly gained interest in medical image computing, aiming to tackle the performance degradation of deep neural networks when being deployed to unseen data with heterogeneous characteristics. In this work, we present a novel unsupervised domain adaptation framework, named as Synergistic Image and Feature Alignment (SIFA), to effectively adapt a segmentation network to an unlabeled target domain. Our proposed SIFA conducts synergistic alignment of domains from both image and feature perspectives. In particular, we simultaneously transform the appearance of images across domains and enhance domain-invariance of the extracted features by leveraging adversarial learning in multiple aspects and with a deeply supervised mechanism. The feature encoder is shared between both adaptive perspectives to leverage their mutual benefits via end-to-end learning. We have extensively evaluated our method with cardiac substructure segmentation and abdominal multi-organ segmentation for bidirectional cross-modality adaptation between MRI and CT images. Experimental results on two different tasks demonstrate that our SIFA method is effective in improving segmentation performance on unlabeled target images, and outperforms the state-of-the-art domain adaptation approaches by a large margin.
Unpaired Multi-modal Segmentation via Knowledge Distillation
Jan 06, 2020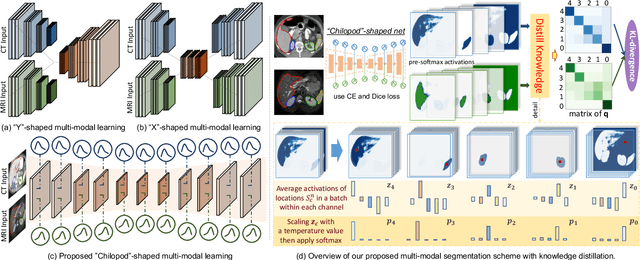
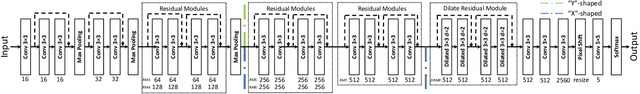
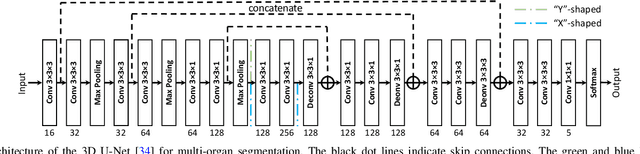
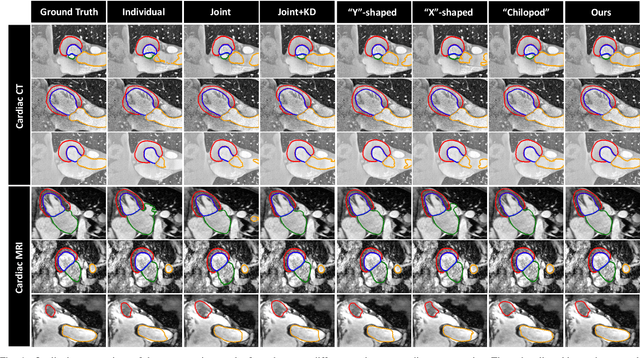
Abstract:Multi-modal learning is typically performed with network architectures containing modality-specific layers and shared layers, utilizing co-registered images of different modalities. We propose a novel learning scheme for unpaired cross-modality image segmentation, with a highly compact architecture achieving superior segmentation accuracy. In our method, we heavily reuse network parameters, by sharing all convolutional kernels across CT and MRI, and only employ modality-specific internal normalization layers which compute respective statistics. To effectively train such a highly compact model, we introduce a novel loss term inspired by knowledge distillation, by explicitly constraining the KL-divergence of our derived prediction distributions between modalities. We have extensively validated our approach on two multi-class segmentation problems: i) cardiac structure segmentation, and ii) abdominal organ segmentation. Different network settings, i.e., 2D dilated network and 3D U-net, are utilized to investigate our method's general efficacy. Experimental results on both tasks demonstrate that our novel multi-modal learning scheme consistently outperforms single-modal training and previous multi-modal approaches.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge