Kaixin Xu
Joint Architecture-Token-Bitwidth Multi-Axis Optimization of Vision Transformers for Semiconductor IC Packaging
May 03, 2026Abstract:Vision Transformers (ViTs) have achieved strong performance in visual recognition, yet their deployment in resource-constrained industrial environments remains limited. Some main challenges are their high computational cost, memory requirement, and energy consumption. While individual efficiency techniques such as neural architecture search (NAS), token compression, and low-precision inference have been extensively studied, most prior work targets only a single optimization axis, limiting overall deployment gains while preserving accuracy. In this paper, we present one of the first holistic frameworks that jointly optimizes three complementary axes: architecture, token, and bit-width. Specifically, the framework identifies compact backbones via Neural Architecture Search (AutoFormer), reduces information processing via token merging (ToMe), and accelerates per-operation execution via fp16 mixed-precision inference. Starting from a DeiT-B/16 baseline, we first analyze accuracy-efficiency trade-offs on ImageNet-1K under aggressive compression. Then, we apply the selected configurations to a real-world in-house 3D X-ray semiconductor defect classification dataset for IC chip packaging inspection. Results show that the proposed multi-axis framework achieves more than 10 times improvement in throughput along with over 10 times reductions in parameter count, FLOPs, and energy consumption, while maintaining the required accuracy on the downstream industrial task. To the best of our knowledge, this is among the earliest works to jointly optimize architecture, token, and bit-width dimensions in ViTs and the first such resource-efficient, deployment-focused study tailored to semiconductor manufacturing.
Select-then-Solve: Paradigm Routing as Inference-Time Optimization for LLM Agents
Apr 08, 2026Abstract:When an LLM-based agent improves on a task, is the gain from the model itself or from the reasoning paradigm wrapped around it? We study this question by comparing six inference-time paradigms, namely Direct, CoT, ReAct, Plan-Execute, Reflection, and ReCode, across four frontier LLMs and ten benchmarks, yielding roughly 18,000 runs. We find that reasoning structure helps dramatically on some tasks but hurts on others: ReAct improves over Direct by 44pp on GAIA, while CoT degrades performance by 15pp on HumanEval. No single paradigm dominates, and oracle per-task selection beats the best fixed paradigm by 17.1pp on average. Motivated by this complementarity, we propose a select-then-solve approach: before answering each task, a lightweight embedding-based router selects the most suitable paradigm. Across four models, the router improves average accuracy from 47.6% to 53.1%, outperforming the best fixed paradigm at 50.3% by 2.8pp and recovering up to 37% of the oracle gap. In contrast, zero-shot self-routing only works for GPT-5 at 67.1% and fails for weaker models, all trailing the learned router. Our results argue that reasoning paradigm selection should be a per-task decision made by a learned router, not a fixed architectural choice.
Ego to World: Collaborative Spatial Reasoning in Embodied Systems via Reinforcement Learning
Mar 16, 2026Abstract:Understanding the world from distributed, partial viewpoints is a fundamental challenge for embodied multi-agent systems. Each agent perceives the environment through an ego-centric view that is often limited by occlusion and ambiguity. To study this problem, we introduce the Ego-to-World (E2W) benchmark, which evaluates a vision-language model's ability to fuse heterogeneous viewpoints across three tasks: (i) global counting, (ii) relational location reasoning, and (iii) action-oriented grasping that requires predicting view-specific image coordinates. To address this setting, we propose CoRL, a two-stage framework that combines Chain-of-Thought supervised fine-tuning with reinforcement learning using Group-Relative Policy Optimization. Its core component, the Cross-View Spatial Reward (CVSR), provides dense task-aligned feedback by linking reasoning steps to visual evidence, ensuring coherent cross-view entity resolution, and guiding the model toward correct final predictions. Experiments on E2W show that CoRL consistently surpasses strong proprietary and open-source baselines on both reasoning and perception-grounding metrics, while ablations further confirm the necessity of each CVSR component. Beyond that, CoRL generalizes to external spatial reasoning benchmarks and enables effective real-world multi-robot manipulation with calibrated multi-camera rigs, demonstrating cross-view localization and successful grasp-and-place execution. Together, E2W and CoRL provide a principled foundation for learning world-centric scene understanding from distributed, ego-centric observations, advancing collaborative embodied AI.
RPO:Reinforcement Fine-Tuning with Partial Reasoning Optimization
Jan 27, 2026Abstract:Within the domain of large language models, reinforcement fine-tuning algorithms necessitate the generation of a complete reasoning trajectory beginning from the input query, which incurs significant computational overhead during the rollout phase of training. To address this issue, we analyze the impact of different segments of the reasoning path on the correctness of the final result and, based on these insights, propose Reinforcement Fine-Tuning with Partial Reasoning Optimization (RPO), a plug-and-play reinforcement fine-tuning algorithm. Unlike traditional reinforcement fine-tuning algorithms that generate full reasoning paths, RPO trains the model by generating suffixes of the reasoning path using experience cache. During the rollout phase of training, RPO reduces token generation in this phase by approximately 95%, greatly lowering the theoretical time overhead. Compared with full-path reinforcement fine-tuning algorithms, RPO reduces the training time of the 1.5B model by 90% and the 7B model by 72%. At the same time, it can be integrated with typical algorithms such as GRPO and DAPO, enabling them to achieve training acceleration while maintaining performance comparable to the original algorithms. Our code is open-sourced at https://github.com/yhz5613813/RPO.
Dynamic Deep Graph Learning for Incomplete Multi-View Clustering with Masked Graph Reconstruction Loss
Nov 14, 2025Abstract:The prevalence of real-world multi-view data makes incomplete multi-view clustering (IMVC) a crucial research. The rapid development of Graph Neural Networks (GNNs) has established them as one of the mainstream approaches for multi-view clustering. Despite significant progress in GNNs-based IMVC, some challenges remain: (1) Most methods rely on the K-Nearest Neighbors (KNN) algorithm to construct static graphs from raw data, which introduces noise and diminishes the robustness of the graph topology. (2) Existing methods typically utilize the Mean Squared Error (MSE) loss between the reconstructed graph and the sparse adjacency graph directly as the graph reconstruction loss, leading to substantial gradient noise during optimization. To address these issues, we propose a novel \textbf{D}ynamic Deep \textbf{G}raph Learning for \textbf{I}ncomplete \textbf{M}ulti-\textbf{V}iew \textbf{C}lustering with \textbf{M}asked Graph Reconstruction Loss (DGIMVCM). Firstly, we construct a missing-robust global graph from the raw data. A graph convolutional embedding layer is then designed to extract primary features and refined dynamic view-specific graph structures, leveraging the global graph for imputation of missing views. This process is complemented by graph structure contrastive learning, which identifies consistency among view-specific graph structures. Secondly, a graph self-attention encoder is introduced to extract high-level representations based on the imputed primary features and view-specific graphs, and is optimized with a masked graph reconstruction loss to mitigate gradient noise during optimization. Finally, a clustering module is constructed and optimized through a pseudo-label self-supervised training mechanism. Extensive experiments on multiple datasets validate the effectiveness and superiority of DGIMVCM.
Learning Primitive Embodied World Models: Towards Scalable Robotic Learning
Aug 28, 2025



Abstract:While video-generation-based embodied world models have gained increasing attention, their reliance on large-scale embodied interaction data remains a key bottleneck. The scarcity, difficulty of collection, and high dimensionality of embodied data fundamentally limit the alignment granularity between language and actions and exacerbate the challenge of long-horizon video generation--hindering generative models from achieving a "GPT moment" in the embodied domain. There is a naive observation: the diversity of embodied data far exceeds the relatively small space of possible primitive motions. Based on this insight, we propose a novel paradigm for world modeling--Primitive Embodied World Models (PEWM). By restricting video generation to fixed short horizons, our approach 1) enables fine-grained alignment between linguistic concepts and visual representations of robotic actions, 2) reduces learning complexity, 3) improves data efficiency in embodied data collection, and 4) decreases inference latency. By equipping with a modular Vision-Language Model (VLM) planner and a Start-Goal heatmap Guidance mechanism (SGG), PEWM further enables flexible closed-loop control and supports compositional generalization of primitive-level policies over extended, complex tasks. Our framework leverages the spatiotemporal vision priors in video models and the semantic awareness of VLMs to bridge the gap between fine-grained physical interaction and high-level reasoning, paving the way toward scalable, interpretable, and general-purpose embodied intelligence.
A Spatial-Physics Informed Model for 3D Spiral Sample Scanned by SQUID Microscopy
Jul 16, 2025

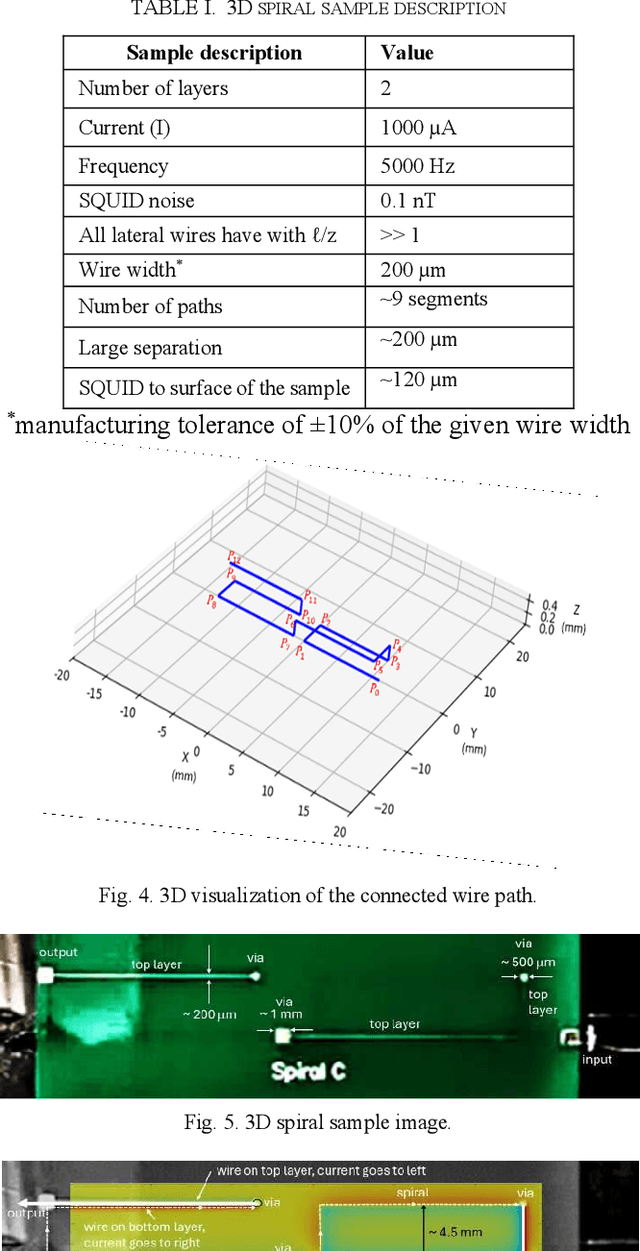

Abstract:The development of advanced packaging is essential in the semiconductor manufacturing industry. However, non-destructive testing (NDT) of advanced packaging becomes increasingly challenging due to the depth and complexity of the layers involved. In such a scenario, Magnetic field imaging (MFI) enables the imaging of magnetic fields generated by currents. For MFI to be effective in NDT, the magnetic fields must be converted into current density. This conversion has typically relied solely on a Fast Fourier Transform (FFT) for magnetic field inversion; however, the existing approach does not consider eddy current effects or image misalignment in the test setup. In this paper, we present a spatial-physics informed model (SPIM) designed for a 3D spiral sample scanned using Superconducting QUantum Interference Device (SQUID) microscopy. The SPIM encompasses three key components: i) magnetic image enhancement by aligning all the "sharp" wire field signals to mitigate the eddy current effect using both in-phase (I-channel) and quadrature-phase (Q-channel) images; (ii) magnetic image alignment that addresses skew effects caused by any misalignment of the scanning SQUID microscope relative to the wire segments; and (iii) an inversion method for converting magnetic fields to magnetic currents by integrating the Biot-Savart Law with FFT. The results show that the SPIM improves I-channel sharpness by 0.3% and reduces Q-channel sharpness by 25%. Also, we were able to remove rotational and skew misalignments of 0.30 in a real image. Overall, SPIM highlights the potential of combining spatial analysis with physics-driven models in practical applications.
* copyright 2025 IEEE. Personal use of this material is permitted. Permission from IEEE must be obtained for all other uses, in any current or future media, including reprinting/republishing this material for advertising or promotional purposes, creating new collective works, for resale or redistribution to servers or lists, or reuse of any copyrighted component of this work in other works
Global Graph Propagation with Hierarchical Information Transfer for Incomplete Contrastive Multi-view Clustering
Feb 26, 2025



Abstract:Incomplete multi-view clustering has become one of the important research problems due to the extensive missing multi-view data in the real world. Although the existing methods have made great progress, there are still some problems: 1) most methods cannot effectively mine the information hidden in the missing data; 2) most methods typically divide representation learning and clustering into two separate stages, but this may affect the clustering performance as the clustering results directly depend on the learned representation. To address these problems, we propose a novel incomplete multi-view clustering method with hierarchical information transfer. Firstly, we design the view-specific Graph Convolutional Networks (GCN) to obtain the representation encoding the graph structure, which is then fused into the consensus representation. Secondly, considering that one layer of GCN transfers one-order neighbor node information, the global graph propagation with the consensus representation is proposed to handle the missing data and learn deep representation. Finally, we design a weight-sharing pseudo-classifier with contrastive learning to obtain an end-to-end framework that combines view-specific representation learning, global graph propagation with hierarchical information transfer, and contrastive clustering for joint optimization. Extensive experiments conducted on several commonly-used datasets demonstrate the effectiveness and superiority of our method in comparison with other state-of-the-art approaches. The code is available at https://github.com/KelvinXuu/GHICMC.
DM3D: Distortion-Minimized Weight Pruning for Lossless 3D Object Detection
Jul 02, 2024



Abstract:Applying deep neural networks to 3D point cloud processing has attracted increasing attention due to its advanced performance in many areas, such as AR/VR, autonomous driving, and robotics. However, as neural network models and 3D point clouds expand in size, it becomes a crucial challenge to reduce the computational and memory overhead to meet latency and energy constraints in real-world applications. Although existing approaches have proposed to reduce both computational cost and memory footprint, most of them only address the spatial redundancy in inputs, i.e. removing the redundancy of background points in 3D data. In this paper, we propose a novel post-training weight pruning scheme for 3D object detection that is (1) orthogonal to all existing point cloud sparsifying methods, which determines redundant parameters in the pretrained model that lead to minimal distortion in both locality and confidence (detection distortion); and (2) a universal plug-and-play pruning framework that works with arbitrary 3D detection model. This framework aims to minimize detection distortion of network output to maximally maintain detection precision, by identifying layer-wise sparsity based on second-order Taylor approximation of the distortion. Albeit utilizing second-order information, we introduced a lightweight scheme to efficiently acquire Hessian information, and subsequently perform dynamic programming to solve the layer-wise sparsity. Extensive experiments on KITTI, Nuscenes and ONCE datasets demonstrate that our approach is able to maintain and even boost the detection precision on pruned model under noticeable computation reduction (FLOPs). Noticeably, we achieve over 3.89x, 3.72x FLOPs reduction on CenterPoint and PVRCNN model, respectively, without mAP decrease, significantly improving the state-of-the-art.
LPViT: Low-Power Semi-structured Pruning for Vision Transformers
Jul 02, 2024



Abstract:Vision transformers have emerged as a promising alternative to convolutional neural networks for various image analysis tasks, offering comparable or superior performance. However, one significant drawback of ViTs is their resource-intensive nature, leading to increased memory footprint, computation complexity, and power consumption. To democratize this high-performance technology and make it more environmentally friendly, it is essential to compress ViT models, reducing their resource requirements while maintaining high performance. In this paper, we introduce a new block-structured pruning to address the resource-intensive issue for ViTs, offering a balanced trade-off between accuracy and hardware acceleration. Unlike unstructured pruning or channel-wise structured pruning, block pruning leverages the block-wise structure of linear layers, resulting in more efficient matrix multiplications. To optimize this pruning scheme, our paper proposes a novel hardware-aware learning objective that simultaneously maximizes speedup and minimizes power consumption during inference, tailored to the block sparsity structure. This objective eliminates the need for empirical look-up tables and focuses solely on reducing parametrized layer connections. Moreover, our paper provides a lightweight algorithm to achieve post-training pruning for ViTs, utilizing second-order Taylor approximation and empirical optimization to solve the proposed hardware-aware objective. Extensive experiments on ImageNet are conducted across various ViT architectures, including DeiT-B and DeiT-S, demonstrating competitive performance with other pruning methods and achieving a remarkable balance between accuracy preservation and power savings. Especially, we achieve up to 3.93x and 1.79x speedups on dedicated hardware and GPUs respectively for DeiT-B, and also observe an inference power reduction by 1.4x on real-world GPUs.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge