Junzhou Huang
Graph Convolutional Module for Temporal Action Localization in Videos
Dec 01, 2021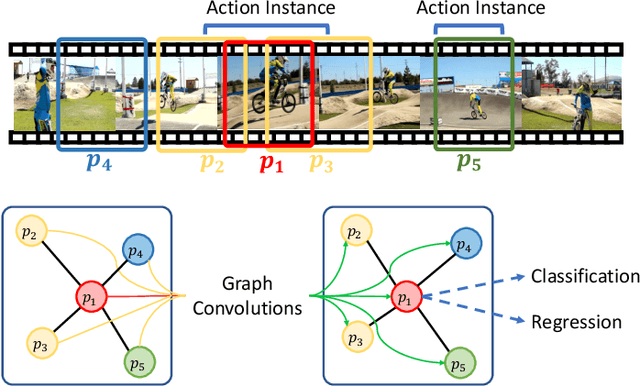
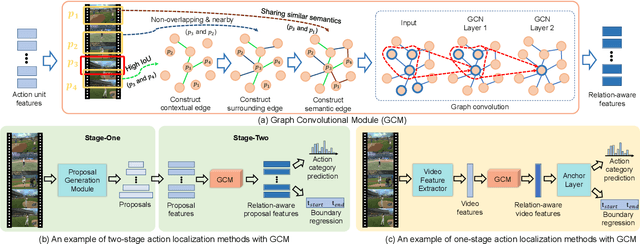
Abstract:Temporal action localization has long been researched in computer vision. Existing state-of-the-art action localization methods divide each video into multiple action units (i.e., proposals in two-stage methods and segments in one-stage methods) and then perform action recognition/regression on each of them individually, without explicitly exploiting their relations during learning. In this paper, we claim that the relations between action units play an important role in action localization, and a more powerful action detector should not only capture the local content of each action unit but also allow a wider field of view on the context related to it. To this end, we propose a general graph convolutional module (GCM) that can be easily plugged into existing action localization methods, including two-stage and one-stage paradigms. To be specific, we first construct a graph, where each action unit is represented as a node and their relations between two action units as an edge. Here, we use two types of relations, one for capturing the temporal connections between different action units, and the other one for characterizing their semantic relationship. Particularly for the temporal connections in two-stage methods, we further explore two different kinds of edges, one connecting the overlapping action units and the other one connecting surrounding but disjointed units. Upon the graph we built, we then apply graph convolutional networks (GCNs) to model the relations among different action units, which is able to learn more informative representations to enhance action localization. Experimental results show that our GCM consistently improves the performance of existing action localization methods, including two-stage methods (e.g., CBR and R-C3D) and one-stage methods (e.g., D-SSAD), verifying the generality and effectiveness of our GCM.
CoDiM: Learning with Noisy Labels via Contrastive Semi-Supervised Learning
Nov 23, 2021
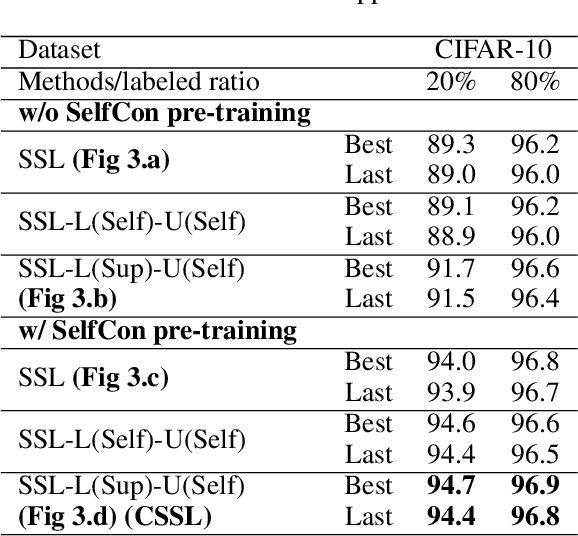
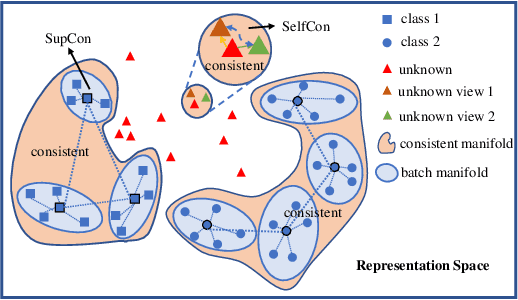
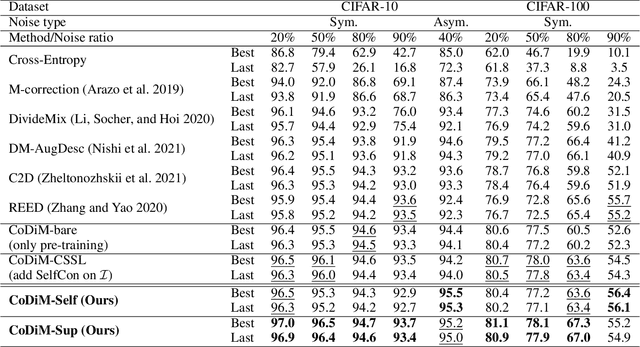
Abstract:Labels are costly and sometimes unreliable. Noisy label learning, semi-supervised learning, and contrastive learning are three different strategies for designing learning processes requiring less annotation cost. Semi-supervised learning and contrastive learning have been recently demonstrated to improve learning strategies that address datasets with noisy labels. Still, the inner connections between these fields as well as the potential to combine their strengths together have only started to emerge. In this paper, we explore further ways and advantages to fuse them. Specifically, we propose CSSL, a unified Contrastive Semi-Supervised Learning algorithm, and CoDiM (Contrastive DivideMix), a novel algorithm for learning with noisy labels. CSSL leverages the power of classical semi-supervised learning and contrastive learning technologies and is further adapted to CoDiM, which learns robustly from multiple types and levels of label noise. We show that CoDiM brings consistent improvements and achieves state-of-the-art results on multiple benchmarks.
Local Augmentation for Graph Neural Networks
Sep 08, 2021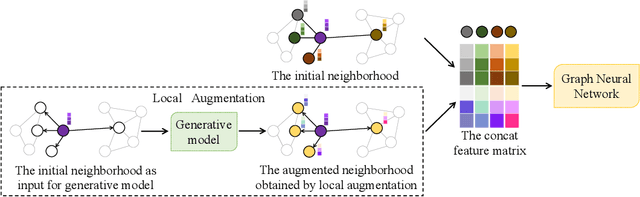
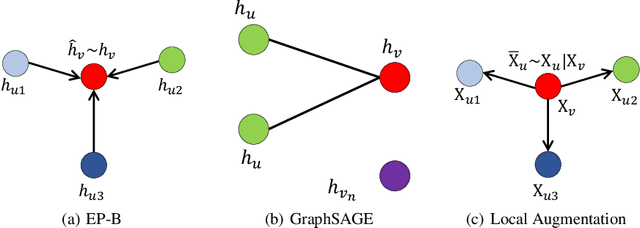
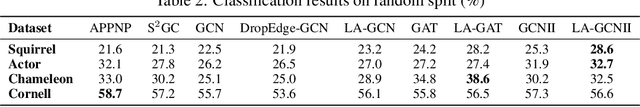
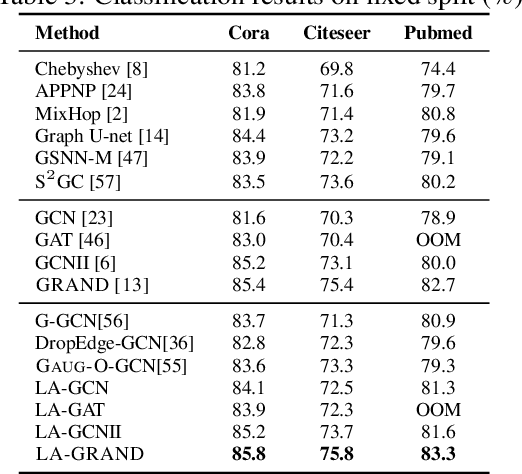
Abstract:Data augmentation has been widely used in image data and linguistic data but remains under-explored on graph-structured data. Existing methods focus on augmenting the graph data from a global perspective and largely fall into two genres: structural manipulation and adversarial training with feature noise injection. However, the structural manipulation approach suffers information loss issues while the adversarial training approach may downgrade the feature quality by injecting noise. In this work, we introduce the local augmentation, which enhances node features by its local subgraph structures. Specifically, we model the data argumentation as a feature generation process. Given the central node's feature, our local augmentation approach learns the conditional distribution of its neighbors' features and generates the neighbors' optimal feature to boost the performance of downstream tasks. Based on the local augmentation, we further design a novel framework: LA-GNN, which can apply to any GNN models in a plug-and-play manner. Extensive experiments and analyses show that local augmentation consistently yields performance improvement for various GNN architectures across a diverse set of benchmarks. Code is available at https://github.com/Soughing0823/LAGNN.
TVT: Transferable Vision Transformer for Unsupervised Domain Adaptation
Aug 12, 2021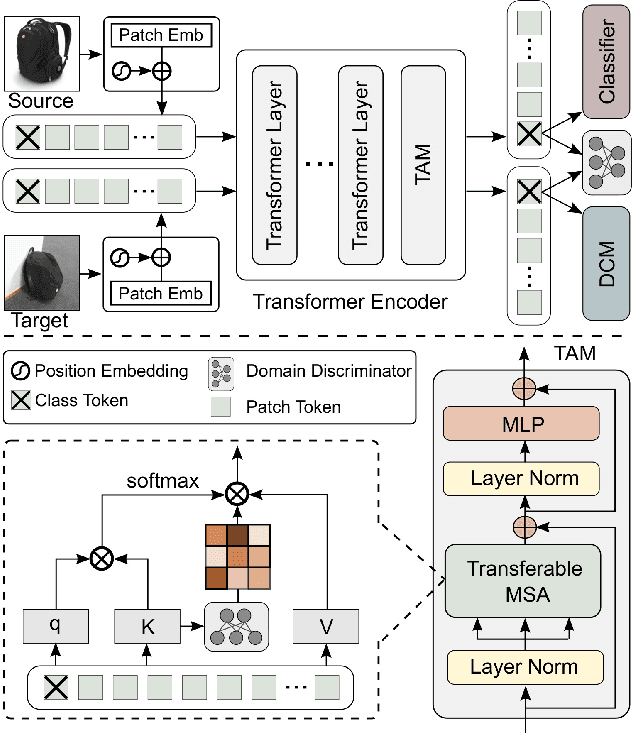
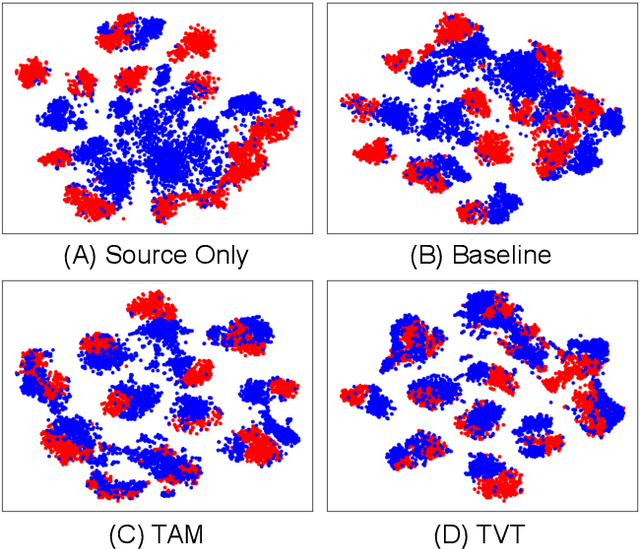
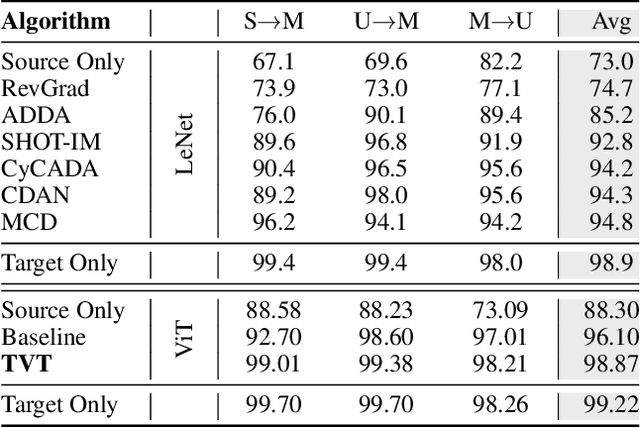
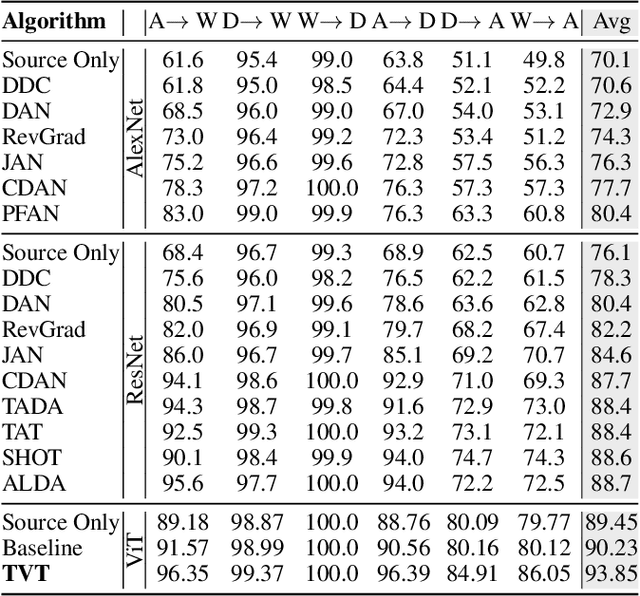

Abstract:Unsupervised domain adaptation (UDA) aims to transfer the knowledge learnt from a labeled source domain to an unlabeled target domain. Previous work is mainly built upon convolutional neural networks (CNNs) to learn domain-invariant representations. With the recent exponential increase in applying Vision Transformer (ViT) to vision tasks, the capability of ViT in adapting cross-domain knowledge, however, remains unexplored in the literature. To fill this gap, this paper first comprehensively investigates the transferability of ViT on a variety of domain adaptation tasks. Surprisingly, ViT demonstrates superior transferability over its CNNs-based counterparts with a large margin, while the performance can be further improved by incorporating adversarial adaptation. Notwithstanding, directly using CNNs-based adaptation strategies fails to take the advantage of ViT's intrinsic merits (e.g., attention mechanism and sequential image representation) which play an important role in knowledge transfer. To remedy this, we propose an unified framework, namely Transferable Vision Transformer (TVT), to fully exploit the transferability of ViT for domain adaptation. Specifically, we delicately devise a novel and effective unit, which we term Transferability Adaption Module (TAM). By injecting learned transferabilities into attention blocks, TAM compels ViT focus on both transferable and discriminative features. Besides, we leverage discriminative clustering to enhance feature diversity and separation which are undermined during adversarial domain alignment. To verify its versatility, we perform extensive studies of TVT on four benchmarks and the experimental results demonstrate that TVT attains significant improvements compared to existing state-of-the-art UDA methods.
Frustratingly Easy Transferability Estimation
Jun 17, 2021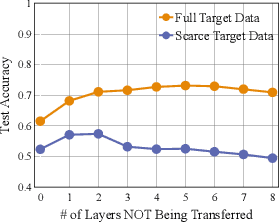
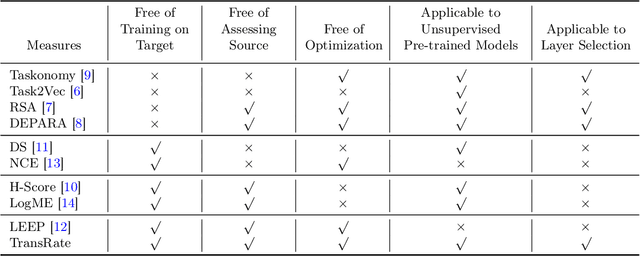
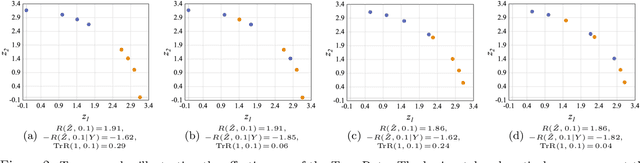
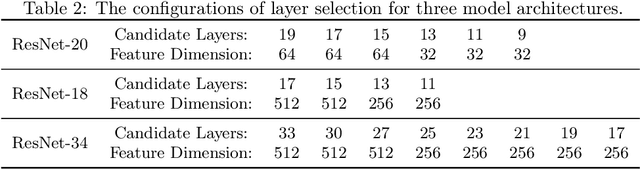
Abstract:Transferability estimation has been an essential tool in selecting a pre-trained model and the layers of it to transfer, so as to maximize the performance on a target task and prevent negative transfer. Existing estimation algorithms either require intensive training on target tasks or have difficulties in evaluating the transferability between layers. We propose a simple, efficient, and effective transferability measure named TransRate. With single pass through the target data, TransRate measures the transferability as the mutual information between the features of target examples extracted by a pre-trained model and labels of them. We overcome the challenge of efficient mutual information estimation by resorting to coding rate that serves as an effective alternative to entropy. TransRate is theoretically analyzed to be closely related to the performance after transfer learning. Despite its extraordinary simplicity in 10 lines of codes, TransRate performs remarkably well in extensive evaluations on 22 pre-trained models and 16 downstream tasks.
PI-GNN: A Novel Perspective on Semi-Supervised Node Classification against Noisy Labels
Jun 14, 2021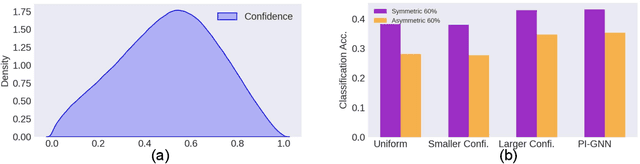
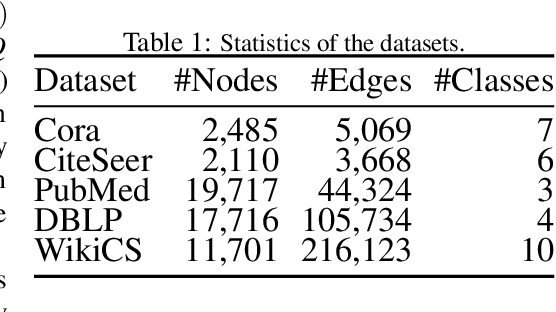
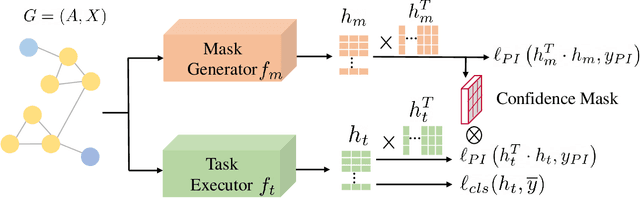
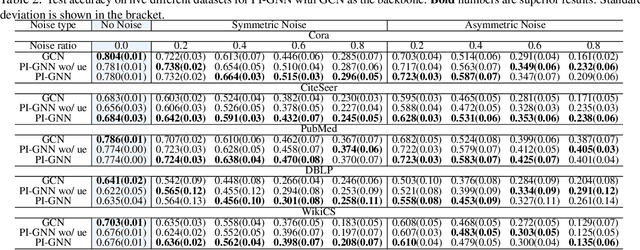
Abstract:Semi-supervised node classification, as a fundamental problem in graph learning, leverages unlabeled nodes along with a small portion of labeled nodes for training. Existing methods rely heavily on high-quality labels, which, however, are expensive to obtain in real-world applications since certain noises are inevitably involved during the labeling process. It hence poses an unavoidable challenge for the learning algorithm to generalize well. In this paper, we propose a novel robust learning objective dubbed pairwise interactions (PI) for the model, such as Graph Neural Network (GNN) to combat noisy labels. Unlike classic robust training approaches that operate on the pointwise interactions between node and class label pairs, PI explicitly forces the embeddings for node pairs that hold a positive PI label to be close to each other, which can be applied to both labeled and unlabeled nodes. We design several instantiations for PI labels based on the graph structure and the node class labels, and further propose a new uncertainty-aware training technique to mitigate the negative effect of the sub-optimal PI labels. Extensive experiments on different datasets and GNN architectures demonstrate the effectiveness of PI, yielding a promising improvement over the state-of-the-art methods.
Energy-Based Learning for Cooperative Games, with Applications to Feature/Data/Model Valuations
Jun 05, 2021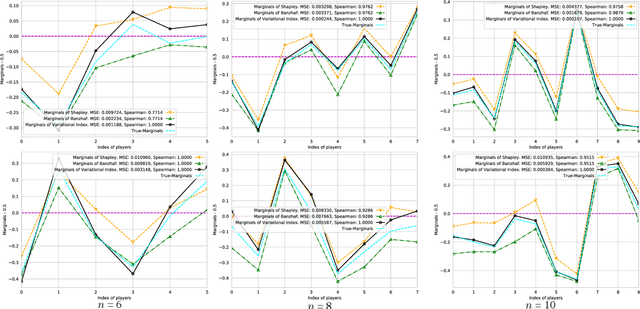
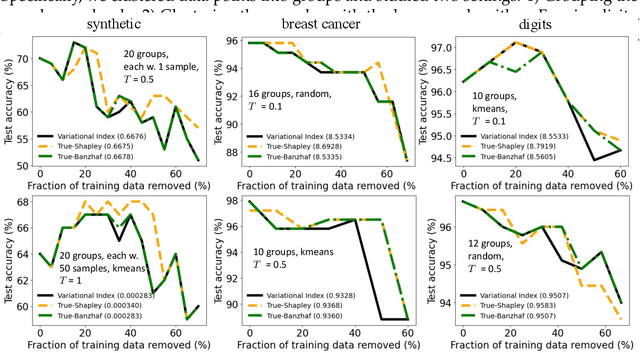
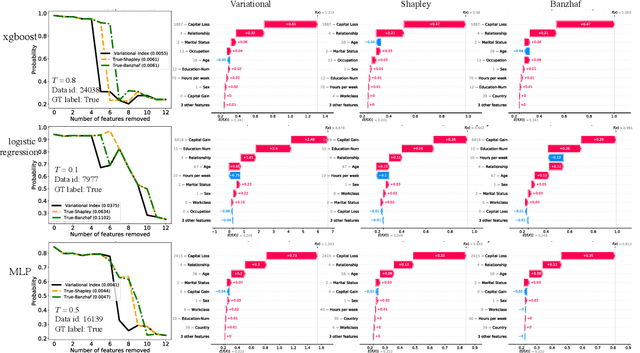
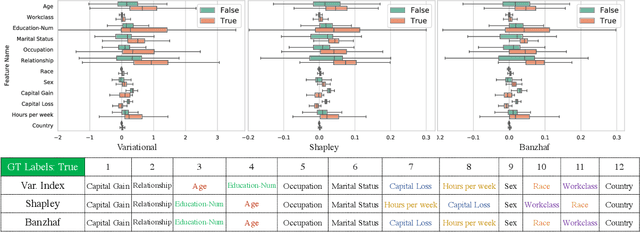
Abstract:Valuation problems, such as attribution-based feature interpretation, data valuation and model valuation for ensembles, become increasingly more important in many machine learning applications. Such problems are commonly solved by well-known game-theoretic criteria, such as Shapley value or Banzhaf index. In this work, we present a novel energy-based treatment for cooperative games, with a theoretical justification by the maximum entropy framework. Surprisingly, by conducting variational inference of the energy-based model, we recover various game-theoretic valuation criteria, such as Shapley value and Banzhaf index, through conducting one-step gradient ascent for maximizing the mean-field ELBO objective. This observation also verifies the rationality of existing criteria, as they are all trying to decouple the correlations among the players through the mean-field approach. By running gradient ascent for multiple steps, we achieve a trajectory of the valuations, among which we define the valuation with the best conceivable decoupling error as the Variational Index. We experimentally demonstrate that the proposed Variational Index enjoys intriguing properties on certain synthetic and real-world valuation problems.
EBM-Fold: Fully-Differentiable Protein Folding Powered by Energy-based Models
May 31, 2021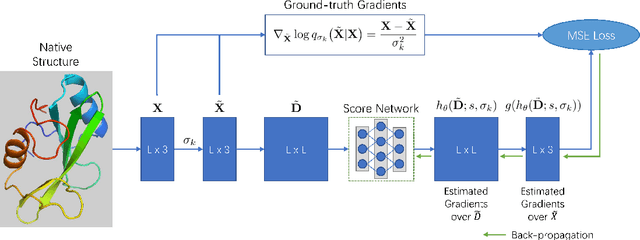

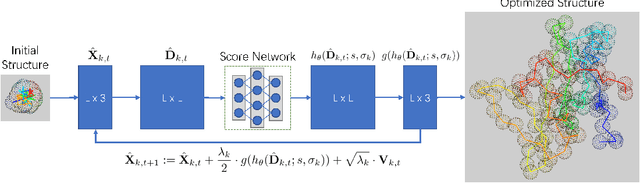
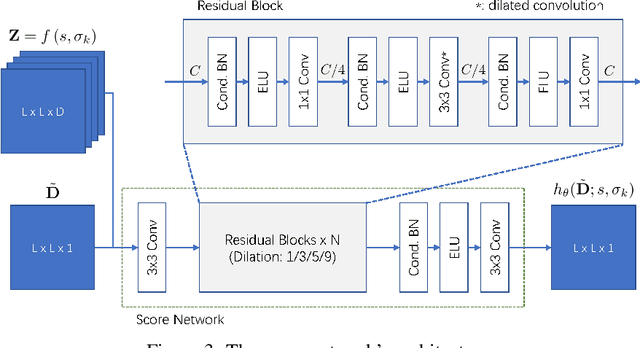
Abstract:Accurate protein structure prediction from amino-acid sequences is critical to better understanding the protein function. Recent advances in this area largely benefit from more precise inter-residue distance and orientation predictions, powered by deep neural networks. However, the structure optimization procedure is still dominated by traditional tools, e.g. Rosetta, where the structure is solved via minimizing a pre-defined statistical energy function (with optional prediction-based restraints). Such energy function may not be optimal in formulating the whole conformation space of proteins. In this paper, we propose a fully-differentiable approach for protein structure optimization, guided by a data-driven generative network. This network is trained in a denoising manner, attempting to predict the correction signal from corrupted distance matrices between Ca atoms. Once the network is well trained, Langevin dynamics based sampling is adopted to gradually optimize structures from random initialization. Extensive experiments demonstrate that our EBM-Fold approach can efficiently produce high-quality decoys, compared against traditional Rosetta-based structure optimization routines.
tFold-TR: Combining Deep Learning Enhanced Hybrid Potential Energy for Template-Based Modeling Structure Refinement
May 30, 2021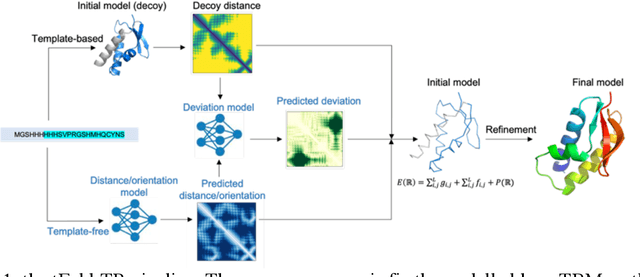
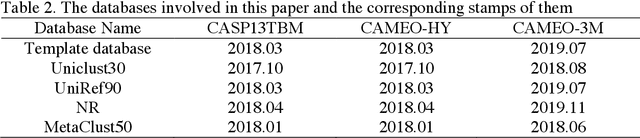


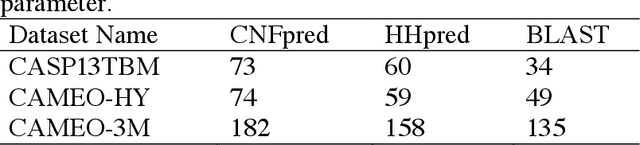
Abstract:Protein structure prediction has been a grand challenge for over 50 years, owing to its broad scientific and application interests. There are two primary types of modeling algorithms, template-free modeling and template-based modeling. The latter one is suitable for easy prediction tasks and is widely adopted in computer-aided drug discoveries for drug design and screening. Although it has been several decades since its first edition, the current template-based modeling approach suffers from two critical problems: 1) there are many missing regions in the template-query sequence alignment, and 2) the accuracy of the distance pairs from different regions of the template varies, and this information is not well introduced into the modeling. To solve these two problems, we propose a structural optimization process based on template modeling, introducing two neural network models to predict the distance information of the missing regions and the accuracy of the distance pairs of different regions in the template modeling structure. The predicted distances and residue pairwise-specific deviations are incorporated into the potential energy function for structural optimization, which significantly improves the qualities of the original template modeling decoys.
Learning Graphon Autoencoders for Generative Graph Modeling
May 29, 2021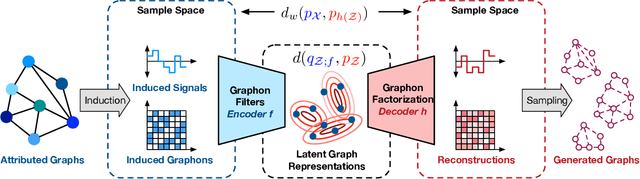
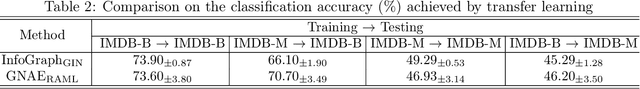
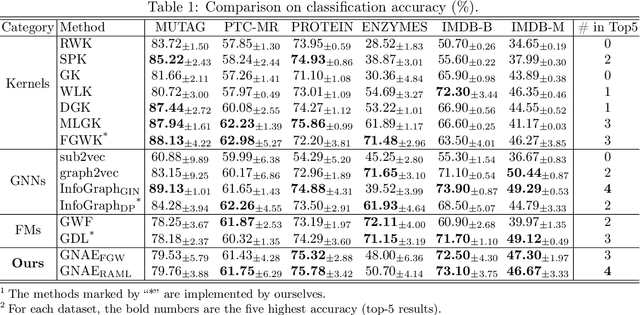
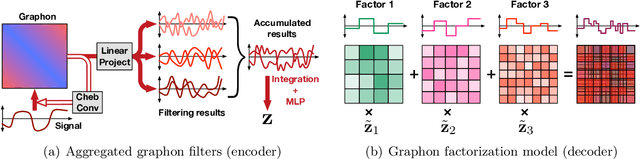

Abstract:Graphon is a nonparametric model that generates graphs with arbitrary sizes and can be induced from graphs easily. Based on this model, we propose a novel algorithmic framework called \textit{graphon autoencoder} to build an interpretable and scalable graph generative model. This framework treats observed graphs as induced graphons in functional space and derives their latent representations by an encoder that aggregates Chebshev graphon filters. A linear graphon factorization model works as a decoder, leveraging the latent representations to reconstruct the induced graphons (and the corresponding observed graphs). We develop an efficient learning algorithm to learn the encoder and the decoder, minimizing the Wasserstein distance between the model and data distributions. This algorithm takes the KL divergence of the graph distributions conditioned on different graphons as the underlying distance and leads to a reward-augmented maximum likelihood estimation. The graphon autoencoder provides a new paradigm to represent and generate graphs, which has good generalizability and transferability.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge