Jan S. Kirschke
Department of Diagnostic and Interventional Neuroradiology, School of Medicine, Klinikum rechts der Isar, Technical University of Munich, Germany
SPINEPS -- Automatic Whole Spine Segmentation of T2-weighted MR images using a Two-Phase Approach to Multi-class Semantic and Instance Segmentation
Feb 26, 2024



Abstract:Purpose. To present SPINEPS, an open-source deep learning approach for semantic and instance segmentation of 14 spinal structures (ten vertebra substructures, intervertebral discs, spinal cord, spinal canal, and sacrum) in whole body T2w MRI. Methods. During this HIPPA-compliant, retrospective study, we utilized the public SPIDER dataset (218 subjects, 63% female) and a subset of the German National Cohort (1423 subjects, mean age 53, 49% female) for training and evaluation. We combined CT and T2w segmentations to train models that segment 14 spinal structures in T2w sagittal scans both semantically and instance-wise. Performance evaluation metrics included Dice similarity coefficient, average symmetrical surface distance, panoptic quality, segmentation quality, and recognition quality. Statistical significance was assessed using the Wilcoxon signed-rank test. An in-house dataset was used to qualitatively evaluate out-of-distribution samples. Results. On the public dataset, our approach outperformed the baseline (instance-wise vertebra dice score 0.929 vs. 0.907, p-value<0.001). Training on auto-generated annotations and evaluating on manually corrected test data from the GNC yielded global dice scores of 0.900 for vertebrae, 0.960 for intervertebral discs, and 0.947 for the spinal canal. Incorporating the SPIDER dataset during training increased these scores to 0.920, 0.967, 0.958, respectively. Conclusions. The proposed segmentation approach offers robust segmentation of 14 spinal structures in T2w sagittal images, including the spinal cord, spinal canal, intervertebral discs, endplate, sacrum, and vertebrae. The approach yields both a semantic and instance mask as output, thus being easy to utilize. This marks the first publicly available algorithm for whole spine segmentation in sagittal T2w MR imaging.
Benchmarking the CoW with the TopCoW Challenge: Topology-Aware Anatomical Segmentation of the Circle of Willis for CTA and MRA
Dec 29, 2023



Abstract:The Circle of Willis (CoW) is an important network of arteries connecting major circulations of the brain. Its vascular architecture is believed to affect the risk, severity, and clinical outcome of serious neuro-vascular diseases. However, characterizing the highly variable CoW anatomy is still a manual and time-consuming expert task. The CoW is usually imaged by two angiographic imaging modalities, magnetic resonance angiography (MRA) and computed tomography angiography (CTA), but there exist limited public datasets with annotations on CoW anatomy, especially for CTA. Therefore we organized the TopCoW Challenge in 2023 with the release of an annotated CoW dataset and invited submissions worldwide for the CoW segmentation task, which attracted over 140 registered participants from four continents. TopCoW dataset was the first public dataset with voxel-level annotations for CoW's 13 vessel components, made possible by virtual-reality (VR) technology. It was also the first dataset with paired MRA and CTA from the same patients. TopCoW challenge aimed to tackle the CoW characterization problem as a multiclass anatomical segmentation task with an emphasis on topological metrics. The top performing teams managed to segment many CoW components to Dice scores around 90%, but with lower scores for communicating arteries and rare variants. There were also topological mistakes for predictions with high Dice scores. Additional topological analysis revealed further areas for improvement in detecting certain CoW components and matching CoW variant's topology accurately. TopCoW represented a first attempt at benchmarking the CoW anatomical segmentation task for MRA and CTA, both morphologically and topologically.
Panoptica -- instance-wise evaluation of 3D semantic and instance segmentation maps
Dec 05, 2023



Abstract:This paper introduces panoptica, a versatile and performance-optimized package designed for computing instance-wise segmentation quality metrics from 2D and 3D segmentation maps. panoptica addresses the limitations of existing metrics and provides a modular framework that complements the original intersection over union-based panoptic quality with other metrics, such as the distance metric Average Symmetric Surface Distance. The package is open-source, implemented in Python, and accompanied by comprehensive documentation and tutorials. panoptica employs a three-step metrics computation process to cover diverse use cases. The efficacy of panoptica is demonstrated on various real-world biomedical datasets, where an instance-wise evaluation is instrumental for an accurate representation of the underlying clinical task. Overall, we envision panoptica as a valuable tool facilitating in-depth evaluation of segmentation methods.
3D Arterial Segmentation via Single 2D Projections and Depth Supervision in Contrast-Enhanced CT Images
Sep 15, 2023Abstract:Automated segmentation of the blood vessels in 3D volumes is an essential step for the quantitative diagnosis and treatment of many vascular diseases. 3D vessel segmentation is being actively investigated in existing works, mostly in deep learning approaches. However, training 3D deep networks requires large amounts of manual 3D annotations from experts, which are laborious to obtain. This is especially the case for 3D vessel segmentation, as vessels are sparse yet spread out over many slices and disconnected when visualized in 2D slices. In this work, we propose a novel method to segment the 3D peripancreatic arteries solely from one annotated 2D projection per training image with depth supervision. We perform extensive experiments on the segmentation of peripancreatic arteries on 3D contrast-enhanced CT images and demonstrate how well we capture the rich depth information from 2D projections. We demonstrate that by annotating a single, randomly chosen projection for each training sample, we obtain comparable performance to annotating multiple 2D projections, thereby reducing the annotation effort. Furthermore, by mapping the 2D labels to the 3D space using depth information and incorporating this into training, we almost close the performance gap between 3D supervision and 2D supervision. Our code is available at: https://github.com/alinafdima/3Dseg-mip-depth.
Framing image registration as a landmark detection problem for better representation of clinical relevance
Jul 31, 2023



Abstract:Nowadays, registration methods are typically evaluated based on sub-resolution tracking error differences. In an effort to reinfuse this evaluation process with clinical relevance, we propose to reframe image registration as a landmark detection problem. Ideally, landmark-specific detection thresholds are derived from an inter-rater analysis. To approximate this costly process, we propose to compute hit rate curves based on the distribution of errors of a sub-sample inter-rater analysis. Therefore, we suggest deriving thresholds from the error distribution using the formula: median + delta * median absolute deviation. The method promises differentiation of previously indistinguishable registration algorithms and further enables assessing the clinical significance in algorithm development.
Inter-Rater Uncertainty Quantification in Medical Image Segmentation via Rater-Specific Bayesian Neural Networks
Jun 28, 2023



Abstract:Automated medical image segmentation inherently involves a certain degree of uncertainty. One key factor contributing to this uncertainty is the ambiguity that can arise in determining the boundaries of a target region of interest, primarily due to variations in image appearance. On top of this, even among experts in the field, different opinions can emerge regarding the precise definition of specific anatomical structures. This work specifically addresses the modeling of segmentation uncertainty, known as inter-rater uncertainty. Its primary objective is to explore and analyze the variability in segmentation outcomes that can occur when multiple experts in medical imaging interpret and annotate the same images. We introduce a novel Bayesian neural network-based architecture to estimate inter-rater uncertainty in medical image segmentation. Our approach has three key advancements. Firstly, we introduce a one-encoder-multi-decoder architecture specifically tailored for uncertainty estimation, enabling us to capture the rater-specific representation of each expert involved. Secondly, we propose Bayesian modeling for the new architecture, allowing efficient capture of the inter-rater distribution, particularly in scenarios with limited annotations. Lastly, we enhance the rater-specific representation by integrating an attention module into each decoder. This module facilitates focused and refined segmentation results for each rater. We conduct extensive evaluations using synthetic and real-world datasets to validate our technical innovations rigorously. Our method surpasses existing baseline methods in five out of seven diverse tasks on the publicly available \emph{QUBIQ} dataset, considering two evaluation metrics encompassing different uncertainty aspects. Our codes, models, and the new dataset are available through our GitHub repository: https://github.com/HaoWang420/bOEMD-net .
Primitive Simultaneous Optimization of Similarity Metrics for Image Registration
Apr 04, 2023



Abstract:Even though simultaneous optimization of similarity metrics represents a standard procedure in the field of semantic segmentation, surprisingly, this does not hold true for image registration. To close this unexpected gap in the literature, we investigate in a complex multi-modal 3D setting whether simultaneous optimization of registration metrics, here implemented by means of primitive summation, can benefit image registration. We evaluate two challenging datasets containing collections of pre- to post-operative and pre- to intra-operative Magnetic Resonance Imaging (MRI) of glioma. Employing the proposed optimization we demonstrate improved registration accuracy in terms of Target Registration Error (TRE) on expert neuroradiologists' landmark annotations.
Multi-contrast MRI Super-resolution via Implicit Neural Representations
Mar 27, 2023Abstract:Clinical routine and retrospective cohorts commonly include multi-parametric Magnetic Resonance Imaging; however, they are mostly acquired in different anisotropic 2D views due to signal-to-noise-ratio and scan-time constraints. Thus acquired views suffer from poor out-of-plane resolution and affect downstream volumetric image analysis that typically requires isotropic 3D scans. Combining different views of multi-contrast scans into high-resolution isotropic 3D scans is challenging due to the lack of a large training cohort, which calls for a subject-specific framework.This work proposes a novel solution to this problem leveraging Implicit Neural Representations (INR). Our proposed INR jointly learns two different contrasts of complementary views in a continuous spatial function and benefits from exchanging anatomical information between them. Trained within minutes on a single commodity GPU, our model provides realistic super-resolution across different pairs of contrasts in our experiments with three datasets. Using Mutual Information (MI) as a metric, we find that our model converges to an optimum MI amongst sequences, achieving anatomically faithful reconstruction. Code is available at: https://github.com/jqmcginnis/multi_contrast_inr.
Semantic Latent Space Regression of Diffusion Autoencoders for Vertebral Fracture Grading
Mar 21, 2023



Abstract:Vertebral fractures are a consequence of osteoporosis, with significant health implications for affected patients. Unfortunately, grading their severity using CT exams is hard and subjective, motivating automated grading methods. However, current approaches are hindered by imbalance and scarcity of data and a lack of interpretability. To address these challenges, this paper proposes a novel approach that leverages unlabelled data to train a generative Diffusion Autoencoder (DAE) model as an unsupervised feature extractor. We model fracture grading as a continuous regression, which is more reflective of the smooth progression of fractures. Specifically, we use a binary, supervised fracture classifier to construct a hyperplane in the DAE's latent space. We then regress the severity of the fracture as a function of the distance to this hyperplane, calibrating the results to the Genant scale. Importantly, the generative nature of our method allows us to visualize different grades of a given vertebra, providing interpretability and insight into the features that contribute to automated grading.
ISLES 2022: A multi-center magnetic resonance imaging stroke lesion segmentation dataset
Jun 14, 2022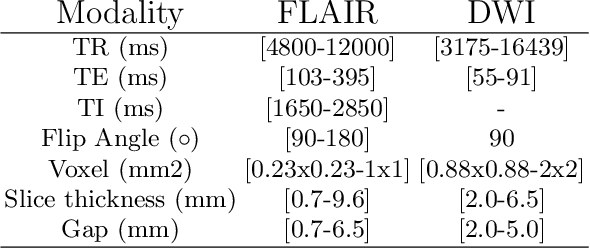
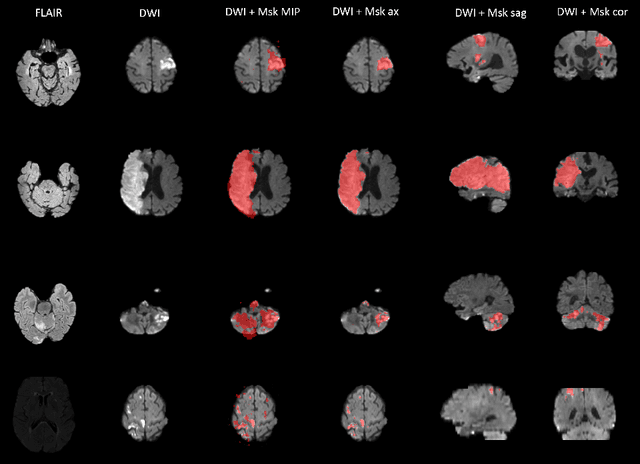

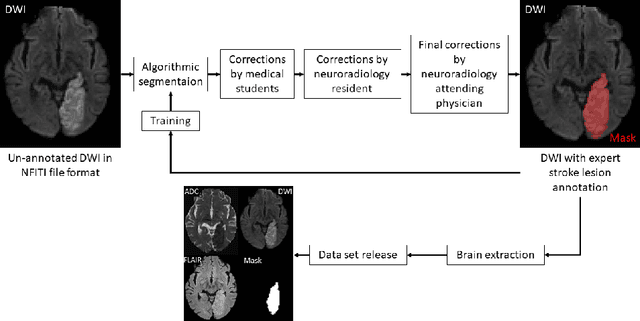
Abstract:Magnetic resonance imaging (MRI) is a central modality for stroke imaging. It is used upon patient admission to make treatment decisions such as selecting patients for intravenous thrombolysis or endovascular therapy. MRI is later used in the duration of hospital stay to predict outcome by visualizing infarct core size and location. Furthermore, it may be used to characterize stroke etiology, e.g. differentiation between (cardio)-embolic and non-embolic stroke. Computer based automated medical image processing is increasingly finding its way into clinical routine. Previous iterations of the Ischemic Stroke Lesion Segmentation (ISLES) challenge have aided in the generation of identifying benchmark methods for acute and sub-acute ischemic stroke lesion segmentation. Here we introduce an expert-annotated, multicenter MRI dataset for segmentation of acute to subacute stroke lesions. This dataset comprises 400 multi-vendor MRI cases with high variability in stroke lesion size, quantity and location. It is split into a training dataset of n=250 and a test dataset of n=150. All training data will be made publicly available. The test dataset will be used for model validation only and will not be released to the public. This dataset serves as the foundation of the ISLES 2022 challenge with the goal of finding algorithmic methods to enable the development and benchmarking of robust and accurate segmentation algorithms for ischemic stroke.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge