Matan Atad
One Sequence to Segment Them All: Efficient Data Augmentation for CT and MRI Cross-Domain 3D Spine Segmentation
May 04, 2026Abstract:Deep learning-based medical image segmentation is increasingly used to support clinical diagnosis and develop new treatment strategies. However, model performance remains limited by the scarcity of high-quality annotated data and insufficient generalization across imaging protocols. This limitation is particularly evident in MRI and CT, where models are typically trained on a single acquisition sequence and exhibit reduced robustness when applied to unseen sequences or contrasts. Although data augmentation is widely used to improve general robustness on medical images, its impact on cross-modality generalization has not been quantitatively explored. In this work, we study a targeted set of data augmentation techniques designed to improve cross-modality transfer. We train three spine segmentation models, each on a single-modality/sequence dataset, and evaluate them across seven out-of-distribution datasets (spanning CT and MRI), reflecting a realistic single-sequence training and multi-sequence/contrast/modality deployment scenario. Our results demonstrate substantial performance gains on unseen domains (average Dice gain of 155 %) while preserving in-domain accuracy (average Dice decrease of 0.008 %), including effective transfer between CT and MRI. To mitigate the computational cost typically associated with strong data augmentation, we implement GPU-optimized augmentations that maintain, and even improve, training efficiency by approximately 10 %. We release our approach as an open-source toolbox, enabling seamless integration into commonly used frameworks such as nnUNet and MONAI. These augmentations significantly enhance robustness to heterogeneous clinical imaging scenarios without compromising training speed.
VERIDAH: Solving Enumeration Anomaly Aware Vertebra Labeling across Imaging Sequences
Jan 20, 2026Abstract:The human spine commonly consists of seven cervical, twelve thoracic, and five lumbar vertebrae. However, enumeration anomalies may result in individuals having eleven or thirteen thoracic vertebrae and four or six lumbar vertebrae. Although the identification of enumeration anomalies has potential clinical implications for chronic back pain and operation planning, the thoracolumbar junction is often poorly assessed and rarely described in clinical reports. Additionally, even though multiple deep-learning-based vertebra labeling algorithms exist, there is a lack of methods to automatically label enumeration anomalies. Our work closes that gap by introducing "Vertebra Identification with Anomaly Handling" (VERIDAH), a novel vertebra labeling algorithm based on multiple classification heads combined with a weighted vertebra sequence prediction algorithm. We show that our approach surpasses existing models on T2w TSE sagittal (98.30% vs. 94.24% of subjects with all vertebrae correctly labeled, p < 0.001) and CT imaging (99.18% vs. 77.26% of subjects with all vertebrae correctly labeled, p < 0.001) and works in arbitrary field-of-view images. VERIDAH correctly labeled the presence 2 Möller et al. of thoracic enumeration anomalies in 87.80% and 96.30% of T2w and CT images, respectively, and lumbar enumeration anomalies in 94.48% and 97.22% for T2w and CT, respectively. Our code and models are available at: https://github.com/Hendrik-code/spineps.
Rule-based Key-Point Extraction for MR-Guided Biomechanical Digital Twins of the Spine
Aug 20, 2025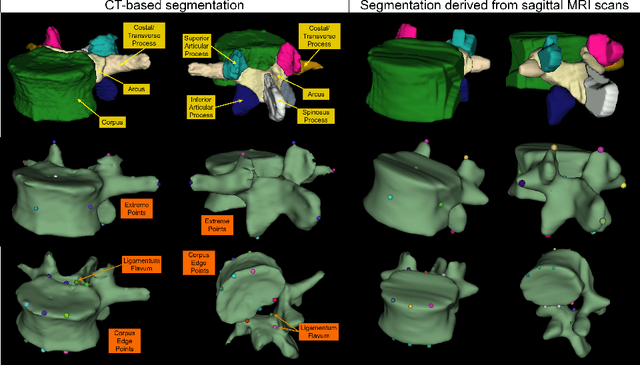

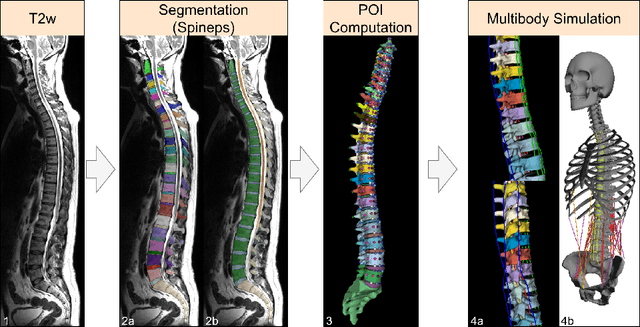

Abstract:Digital twins offer a powerful framework for subject-specific simulation and clinical decision support, yet their development often hinges on accurate, individualized anatomical modeling. In this work, we present a rule-based approach for subpixel-accurate key-point extraction from MRI, adapted from prior CT-based methods. Our approach incorporates robust image alignment and vertebra-specific orientation estimation to generate anatomically meaningful landmarks that serve as boundary conditions and force application points, like muscle and ligament insertions in biomechanical models. These models enable the simulation of spinal mechanics considering the subject's individual anatomy, and thus support the development of tailored approaches in clinical diagnostics and treatment planning. By leveraging MR imaging, our method is radiation-free and well-suited for large-scale studies and use in underrepresented populations. This work contributes to the digital twin ecosystem by bridging the gap between precise medical image analysis with biomechanical simulation, and aligns with key themes in personalized modeling for healthcare.
Automated Thoracolumbar Stump Rib Detection and Analysis in a Large CT Cohort
May 08, 2025



Abstract:Thoracolumbar stump ribs are one of the essential indicators of thoracolumbar transitional vertebrae or enumeration anomalies. While some studies manually assess these anomalies and describe the ribs qualitatively, this study aims to automate thoracolumbar stump rib detection and analyze their morphology quantitatively. To this end, we train a high-resolution deep-learning model for rib segmentation and show significant improvements compared to existing models (Dice score 0.997 vs. 0.779, p-value < 0.01). In addition, we use an iterative algorithm and piece-wise linear interpolation to assess the length of the ribs, showing a success rate of 98.2%. When analyzing morphological features, we show that stump ribs articulate more posteriorly at the vertebrae (-19.2 +- 3.8 vs -13.8 +- 2.5, p-value < 0.01), are thinner (260.6 +- 103.4 vs. 563.6 +- 127.1, p-value < 0.01), and are oriented more downwards and sideways within the first centimeters in contrast to full-length ribs. We show that with partially visible ribs, these features can achieve an F1-score of 0.84 in differentiating stump ribs from regular ones. We publish the model weights and masks for public use.
MAGO-SP: Detection and Correction of Water-Fat Swaps in Magnitude-Only VIBE MRI
Feb 20, 2025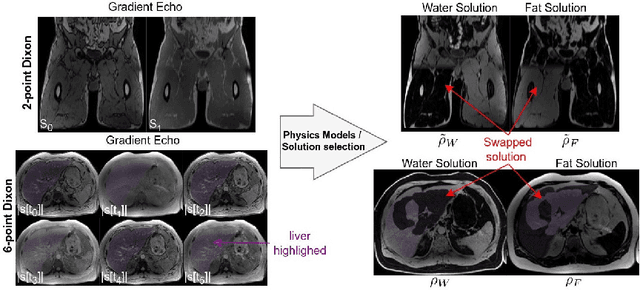
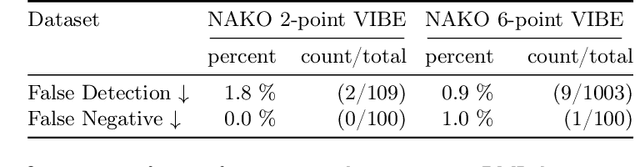
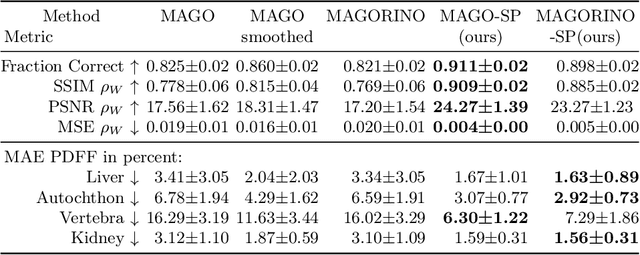
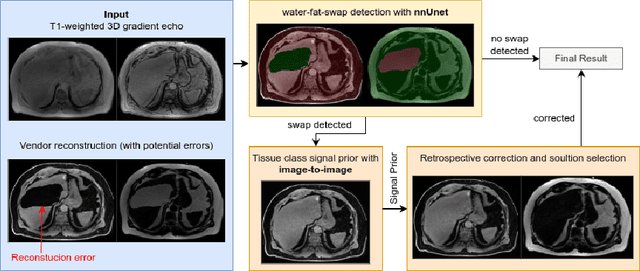

Abstract:Volume Interpolated Breath-Hold Examination (VIBE) MRI generates images suitable for water and fat signal composition estimation. While the two-point VIBE provides water-fat-separated images, the six-point VIBE allows estimation of the effective transversal relaxation rate R2* and the proton density fat fraction (PDFF), which are imaging markers for health and disease. Ambiguity during signal reconstruction can lead to water-fat swaps. This shortcoming challenges the application of VIBE-MRI for automated PDFF analyses of large-scale clinical data and of population studies. This study develops an automated pipeline to detect and correct water-fat swaps in non-contrast-enhanced VIBE images. Our three-step pipeline begins with training a segmentation network to classify volumes as "fat-like" or "water-like," using synthetic water-fat swaps generated by merging fat and water volumes with Perlin noise. Next, a denoising diffusion image-to-image network predicts water volumes as signal priors for correction. Finally, we integrate this prior into a physics-constrained model to recover accurate water and fat signals. Our approach achieves a < 1% error rate in water-fat swap detection for a 6-point VIBE. Notably, swaps disproportionately affect individuals in the Underweight and Class 3 Obesity BMI categories. Our correction algorithm ensures accurate solution selection in chemical phase MRIs, enabling reliable PDFF estimation. This forms a solid technical foundation for automated large-scale population imaging analysis.
PARASIDE: An Automatic Paranasal Sinus Segmentation and Structure Analysis Tool for MRI
Jan 24, 2025



Abstract:Chronic rhinosinusitis (CRS) is a common and persistent sinus imflammation that affects 5 - 12\% of the general population. It significantly impacts quality of life and is often difficult to assess due to its subjective nature in clinical evaluation. We introduce PARASIDE, an automatic tool for segmenting air and soft tissue volumes of the structures of the sinus maxillaris, frontalis, sphenodalis and ethmoidalis in T1 MRI. By utilizing that segmentation, we can quantify feature relations that have been observed only manually and subjectively before. We performed an exemplary study and showed both volume and intensity relations between structures and radiology reports. While the soft tissue segmentation is good, the automated annotations of the air volumes are excellent. The average intensity over air structures are consistently below those of the soft tissues, close to perfect separability. Healthy subjects exhibit lower soft tissue volumes and lower intensities. Our developed system is the first automated whole nasal segmentation of 16 structures, and capable of calculating medical relevant features such as the Lund-Mackay score.
Detecting Unforeseen Data Properties with Diffusion Autoencoder Embeddings using Spine MRI data
Oct 14, 2024



Abstract:Deep learning has made significant strides in medical imaging, leveraging the use of large datasets to improve diagnostics and prognostics. However, large datasets often come with inherent errors through subject selection and acquisition. In this paper, we investigate the use of Diffusion Autoencoder (DAE) embeddings for uncovering and understanding data characteristics and biases, including biases for protected variables like sex and data abnormalities indicative of unwanted protocol variations. We use sagittal T2-weighted magnetic resonance (MR) images of the neck, chest, and lumbar region from 11186 German National Cohort (NAKO) participants. We compare DAE embeddings with existing generative models like StyleGAN and Variational Autoencoder. Evaluations on a large-scale dataset consisting of sagittal T2-weighted MR images of three spine regions show that DAE embeddings effectively separate protected variables such as sex and age. Furthermore, we used t-SNE visualization to identify unwanted variations in imaging protocols, revealing differences in head positioning. Our embedding can identify samples where a sex predictor will have issues learning the correct sex. Our findings highlight the potential of using advanced embedding techniques like DAEs to detect data quality issues and biases in medical imaging datasets. Identifying such hidden relations can enhance the reliability and fairness of deep learning models in healthcare applications, ultimately improving patient care and outcomes.
Don't You (Project Around Discs)? Neural Network Surrogate and Projected Gradient Descent for Calibrating an Intervertebral Disc Finite Element Model
Aug 12, 2024



Abstract:Accurate calibration of finite element (FE) models of human intervertebral discs (IVDs) is essential for their reliability and application in diagnosing and planning treatments for spinal conditions. Traditional calibration methods are computationally intensive, requiring iterative, derivative-free optimization algorithms that often take hours or days to converge. This study addresses these challenges by introducing a novel, efficient, and effective calibration method for an L4-L5 IVD FE model using a neural network (NN) surrogate. The NN surrogate predicts simulation outcomes with high accuracy, outperforming other machine learning models, and significantly reduces the computational cost associated with traditional FE simulations. Next, a Projected Gradient Descent (PGD) approach guided by gradients of the NN surrogate is proposed to efficiently calibrate FE models. Our method explicitly enforces feasibility with a projection step, thus maintaining material bounds throughout the optimization process. The proposed method is evaluated against state-of-the-art Genetic Algorithm (GA) and inverse model baselines on synthetic and in vitro experimental datasets. Our approach demonstrates superior performance on synthetic data, achieving a Mean Absolute Error (MAE) of 0.06 compared to the baselines' MAE of 0.18 and 0.54, respectively. On experimental specimens, our method outperforms the baseline in 5 out of 6 cases. Most importantly, our approach reduces calibration time to under three seconds, compared to up to 8 days per sample required by traditional calibration. Such efficiency paves the way for applying more complex FE models, enabling accurate patient-specific simulations and advancing spinal treatment planning.
Counterfactual Explanations for Medical Image Classification and Regression using Diffusion Autoencoder
Aug 02, 2024



Abstract:Counterfactual explanations (CEs) aim to enhance the interpretability of machine learning models by illustrating how alterations in input features would affect the resulting predictions. Common CE approaches require an additional model and are typically constrained to binary counterfactuals. In contrast, we propose a novel method that operates directly on the latent space of a generative model, specifically a Diffusion Autoencoder (DAE). This approach offers inherent interpretability by enabling the generation of CEs and the continuous visualization of the model's internal representation across decision boundaries. Our method leverages the DAE's ability to encode images into a semantically rich latent space in an unsupervised manner, eliminating the need for labeled data or separate feature extraction models. We show that these latent representations are helpful for medical condition classification and the ordinal regression of severity pathologies, such as vertebral compression fractures (VCF) and diabetic retinopathy (DR). Beyond binary CEs, our method supports the visualization of ordinal CEs using a linear model, providing deeper insights into the model's decision-making process and enhancing interpretability. Experiments across various medical imaging datasets demonstrate the method's advantages in interpretability and versatility. The linear manifold of the DAE's latent space allows for meaningful interpolation and manipulation, making it a powerful tool for exploring medical image properties. Our code is available at https://github.com/matanat/dae_counterfactual.
TotalVibeSegmentator: Full Torso Segmentation for the NAKO and UK Biobank in Volumetric Interpolated Breath-hold Examination Body Images
May 31, 2024



Abstract:Objectives: To present a publicly available torso segmentation network for large epidemiology datasets on volumetric interpolated breath-hold examination (VIBE) images. Materials & Methods: We extracted preliminary segmentations from TotalSegmentator, spine, and body composition networks for VIBE images, then improved them iteratively and retrained a nnUNet network. Using subsets of NAKO (85 subjects) and UK Biobank (16 subjects), we evaluated with Dice-score on a holdout set (12 subjects) and existing organ segmentation approach (1000 subjects), generating 71 semantic segmentation types for VIBE images. We provide an additional network for the vertebra segments 22 individual vertebra types. Results: We achieved an average Dice score of 0.89 +- 0.07 overall 71 segmentation labels. We scored > 0.90 Dice-score on the abdominal organs except for the pancreas with a Dice of 0.70. Conclusion: Our work offers a detailed and refined publicly available full torso segmentation on VIBE images.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge