Satrajit Chakrabarty
Synthetic Volumetric Data Generation Enables Zero-Shot Generalization of Foundation Models in 3D Medical Image Segmentation
Jan 18, 2026Abstract:Foundation models such as Segment Anything Model 2 (SAM 2) exhibit strong generalization on natural images and videos but perform poorly on medical data due to differences in appearance statistics, imaging physics, and three-dimensional structure. To address this gap, we introduce SynthFM-3D, an analytical framework that mathematically models 3D variability in anatomy, contrast, boundary definition, and noise to generate synthetic data for training promptable segmentation models without real annotations. We fine-tuned SAM 2 on 10,000 SynthFM-3D volumes and evaluated it on eleven anatomical structures across three medical imaging modalities (CT, MR, ultrasound) from five public datasets. SynthFM-3D training led to consistent and statistically significant Dice score improvements over the pretrained SAM 2 baseline, demonstrating stronger zero-shot generalization across modalities. When compared with the supervised SAM-Med3D model on unseen cardiac ultrasound data, SynthFM-3D achieved 2-3x higher Dice scores, establishing analytical 3D data modeling as an effective pathway to modality-agnostic medical segmentation.
Analysis of the MICCAI Brain Tumor Segmentation -- Metastases (BraTS-METS) 2025 Lighthouse Challenge: Brain Metastasis Segmentation on Pre- and Post-treatment MRI
Apr 16, 2025Abstract:Despite continuous advancements in cancer treatment, brain metastatic disease remains a significant complication of primary cancer and is associated with an unfavorable prognosis. One approach for improving diagnosis, management, and outcomes is to implement algorithms based on artificial intelligence for the automated segmentation of both pre- and post-treatment MRI brain images. Such algorithms rely on volumetric criteria for lesion identification and treatment response assessment, which are still not available in clinical practice. Therefore, it is critical to establish tools for rapid volumetric segmentations methods that can be translated to clinical practice and that are trained on high quality annotated data. The BraTS-METS 2025 Lighthouse Challenge aims to address this critical need by establishing inter-rater and intra-rater variability in dataset annotation by generating high quality annotated datasets from four individual instances of segmentation by neuroradiologists while being recorded on video (two instances doing "from scratch" and two instances after AI pre-segmentation). This high-quality annotated dataset will be used for testing phase in 2025 Lighthouse challenge and will be publicly released at the completion of the challenge. The 2025 Lighthouse challenge will also release the 2023 and 2024 segmented datasets that were annotated using an established pipeline of pre-segmentation, student annotation, two neuroradiologists checking, and one neuroradiologist finalizing the process. It builds upon its previous edition by including post-treatment cases in the dataset. Using these high-quality annotated datasets, the 2025 Lighthouse challenge plans to test benchmark algorithms for automated segmentation of pre-and post-treatment brain metastases (BM), trained on diverse and multi-institutional datasets of MRI images obtained from patients with brain metastases.
SynthFM: Training Modality-agnostic Foundation Models for Medical Image Segmentation without Real Medical Data
Apr 11, 2025



Abstract:Foundation models like the Segment Anything Model (SAM) excel in zero-shot segmentation for natural images but struggle with medical image segmentation due to differences in texture, contrast, and noise. Annotating medical images is costly and requires domain expertise, limiting large-scale annotated data availability. To address this, we propose SynthFM, a synthetic data generation framework that mimics the complexities of medical images, enabling foundation models to adapt without real medical data. Using SAM's pretrained encoder and training the decoder from scratch on SynthFM's dataset, we evaluated our method on 11 anatomical structures across 9 datasets (CT, MRI, and Ultrasound). SynthFM outperformed zero-shot baselines like SAM and MedSAM, achieving superior results under different prompt settings and on out-of-distribution datasets.
QCResUNet: Joint Subject-level and Voxel-level Segmentation Quality Prediction
Dec 10, 2024Abstract:Deep learning has made significant strides in automated brain tumor segmentation from magnetic resonance imaging (MRI) scans in recent years. However, the reliability of these tools is hampered by the presence of poor-quality segmentation outliers, particularly in out-of-distribution samples, making their implementation in clinical practice difficult. Therefore, there is a need for quality control (QC) to screen the quality of the segmentation results. Although numerous automatic QC methods have been developed for segmentation quality screening, most were designed for cardiac MRI segmentation, which involves a single modality and a single tissue type. Furthermore, most prior works only provided subject-level predictions of segmentation quality and did not identify erroneous parts segmentation that may require refinement. To address these limitations, we proposed a novel multi-task deep learning architecture, termed QCResUNet, which produces subject-level segmentation-quality measures as well as voxel-level segmentation error maps for each available tissue class. To validate the effectiveness of the proposed method, we conducted experiments on assessing its performance on evaluating the quality of two distinct segmentation tasks. First, we aimed to assess the quality of brain tumor segmentation results. For this task, we performed experiments on one internal and two external datasets. Second, we aimed to evaluate the segmentation quality of cardiac Magnetic Resonance Imaging (MRI) data from the Automated Cardiac Diagnosis Challenge. The proposed method achieved high performance in predicting subject-level segmentation-quality metrics and accurately identifying segmentation errors on a voxel basis. This has the potential to be used to guide human-in-the-loop feedback to improve segmentations in clinical settings.
Is SAM 2 Better than SAM in Medical Image Segmentation?
Aug 08, 2024Abstract:Segment Anything Model (SAM) demonstrated impressive performance in zero-shot promptable segmentation on natural images. The recently released Segment Anything Model 2 (SAM 2) model claims to have better performance than SAM on images while extending the model's capabilities to video segmentation. It is important to evaluate the recent model's ability in medical image segmentation in a zero-shot promptable manner. In this work, we performed extensive studies with multiple datasets from different imaging modalities to compare the performance between SAM and SAM 2. We used two point prompt strategies: (i) single positive prompt near the centroid of the target structure and (ii) additional positive prompts placed randomly within the target structure. The evaluation included 21 unique organ-modality combinations including abdominal structures, cardiac structures, and fetal head images acquired from publicly available MRI, CT, and Ultrasound datasets. The preliminary results, based on 2D images, indicate that while SAM 2 may perform slightly better in a few cases, but it does not in general surpass SAM for medical image segmentation. Especially when the contrast is lower like in CT, Ultrasound images, SAM 2 performs poorly than SAM. For MRI images, SAM 2 performs at par or better than SAM. Similar to SAM, SAM 2 also suffers from over-segmentation issue especially when the boundaries of the to-be-segmented organ is fuzzy in nature.
Framing image registration as a landmark detection problem for better representation of clinical relevance
Jul 31, 2023



Abstract:Nowadays, registration methods are typically evaluated based on sub-resolution tracking error differences. In an effort to reinfuse this evaluation process with clinical relevance, we propose to reframe image registration as a landmark detection problem. Ideally, landmark-specific detection thresholds are derived from an inter-rater analysis. To approximate this costly process, we propose to compute hit rate curves based on the distribution of errors of a sub-sample inter-rater analysis. Therefore, we suggest deriving thresholds from the error distribution using the formula: median + delta * median absolute deviation. The method promises differentiation of previously indistinguishable registration algorithms and further enables assessing the clinical significance in algorithm development.
MRI-based classification of IDH mutation and 1p/19q codeletion status of gliomas using a 2.5D hybrid multi-task convolutional neural network
Oct 07, 2022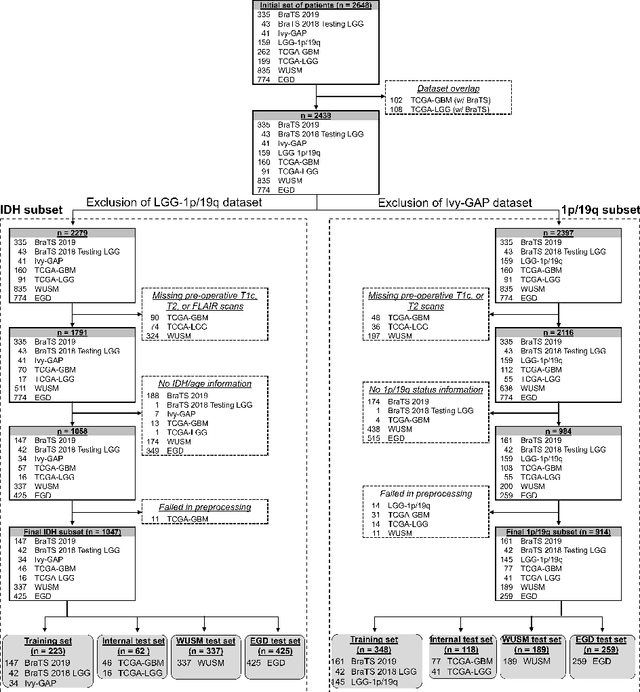
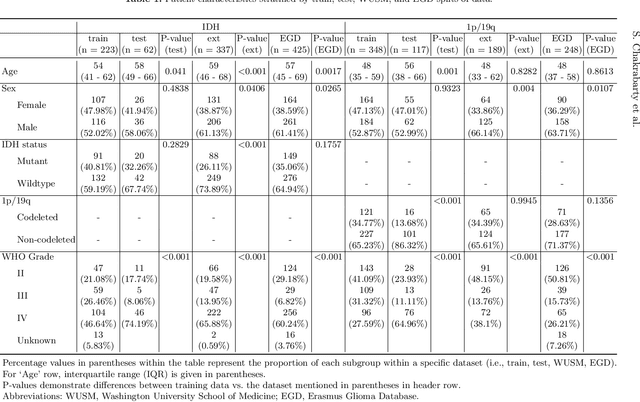
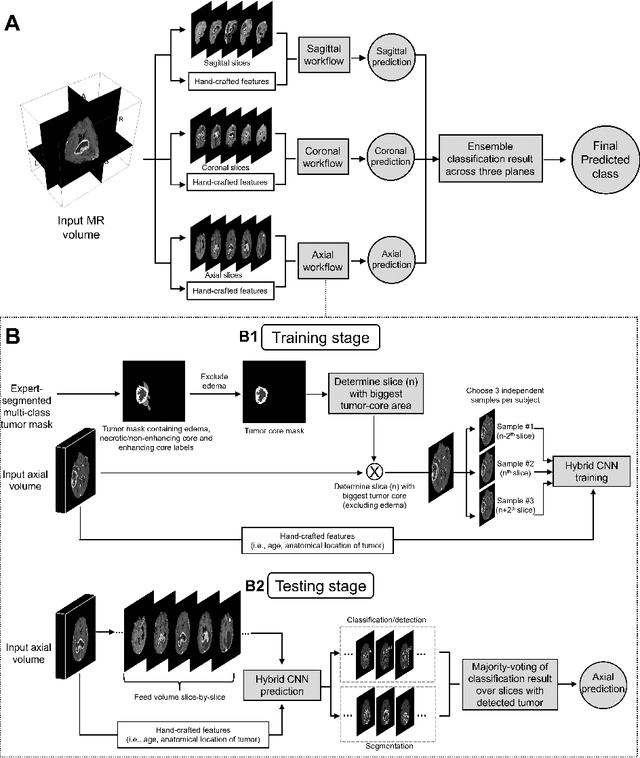
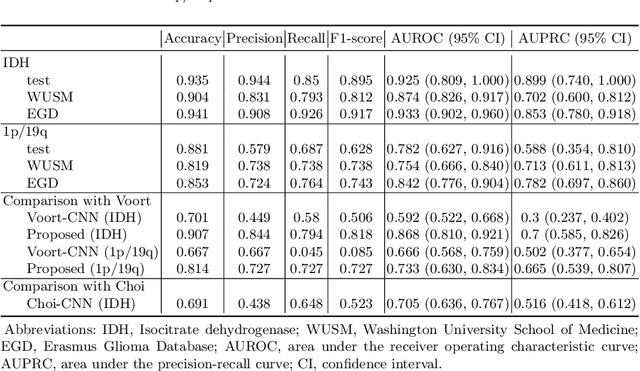
Abstract:Isocitrate dehydrogenase (IDH) mutation and 1p/19q codeletion status are important prognostic markers for glioma. Currently, they are determined using invasive procedures. Our goal was to develop artificial intelligence-based methods to non-invasively determine these molecular alterations from MRI. For this purpose, pre-operative MRI scans of 2648 patients with gliomas (grade II-IV) were collected from Washington University School of Medicine (WUSM; n = 835) and publicly available datasets viz. Brain Tumor Segmentation (BraTS; n = 378), LGG 1p/19q (n = 159), Ivy Glioblastoma Atlas Project (Ivy GAP; n = 41), The Cancer Genome Atlas (TCGA; n = 461), and the Erasmus Glioma Database (EGD; n = 774). A 2.5D hybrid convolutional neural network was proposed to simultaneously localize the tumor and classify its molecular status by leveraging imaging features from MR scans and prior knowledge features from clinical records and tumor location. The models were tested on one internal (TCGA) and two external (WUSM and EGD) test sets. For IDH, the best-performing model achieved areas under the receiver operating characteristic (AUROC) of 0.925, 0.874, 0.933 and areas under the precision-recall curves (AUPRC) of 0.899, 0.702, 0.853 on the internal, WUSM, and EGD test sets, respectively. For 1p/19q, the best model achieved AUROCs of 0.782, 0.754, 0.842, and AUPRCs of 0.588, 0.713, 0.782, on those three data-splits, respectively. The high accuracy of the model on unseen data showcases its generalization capabilities and suggests its potential to perform a 'virtual biopsy' for tailoring treatment planning and overall clinical management of gliomas.
Integrative Imaging Informatics for Cancer Research: Workflow Automation for Neuro-oncology (I3CR-WANO)
Oct 06, 2022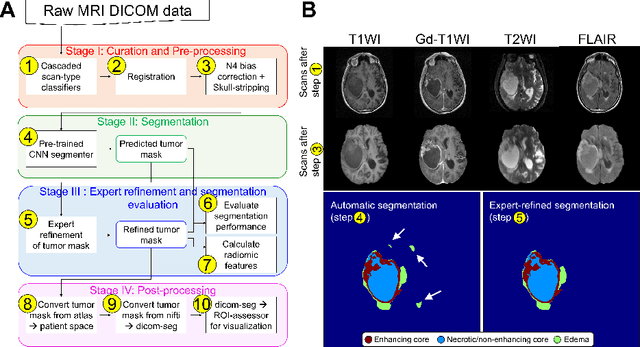
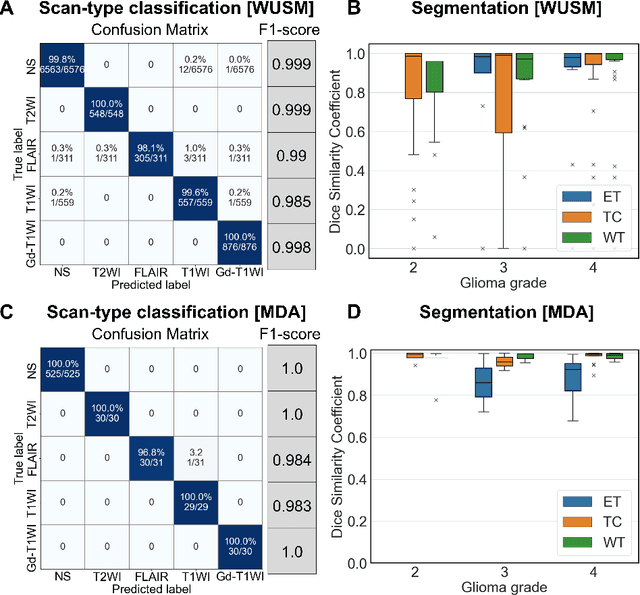
Abstract:Efforts to utilize growing volumes of clinical imaging data to generate tumor evaluations continue to require significant manual data wrangling owing to the data heterogeneity. Here, we propose an artificial intelligence-based solution for the aggregation and processing of multisequence neuro-oncology MRI data to extract quantitative tumor measurements. Our end-to-end framework i) classifies MRI sequences using an ensemble classifier, ii) preprocesses the data in a reproducible manner, iii) delineates tumor tissue subtypes using convolutional neural networks, and iv) extracts diverse radiomic features. Moreover, it is robust to missing sequences and adopts an expert-in-the-loop approach, where the segmentation results may be manually refined by radiologists. Following the implementation of the framework in Docker containers, it was applied to two retrospective glioma datasets collected from the Washington University School of Medicine (WUSM; n = 384) and the M.D. Anderson Cancer Center (MDA; n = 30) comprising preoperative MRI scans from patients with pathologically confirmed gliomas. The scan-type classifier yielded an accuracy of over 99%, correctly identifying sequences from 380/384 and 30/30 sessions from the WUSM and MDA datasets, respectively. Segmentation performance was quantified using the Dice Similarity Coefficient between the predicted and expert-refined tumor masks. Mean Dice scores were 0.882 ($\pm$0.244) and 0.977 ($\pm$0.04) for whole tumor segmentation for WUSM and MDA, respectively. This streamlined framework automatically curated, processed, and segmented raw MRI data of patients with varying grades of gliomas, enabling the curation of large-scale neuro-oncology datasets and demonstrating a high potential for integration as an assistive tool in clinical practice.
The Brain Tumor Sequence Registration Challenge: Establishing Correspondence between Pre-Operative and Follow-up MRI scans of diffuse glioma patients
Dec 13, 2021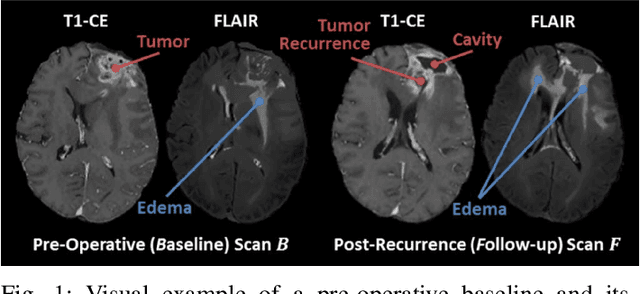
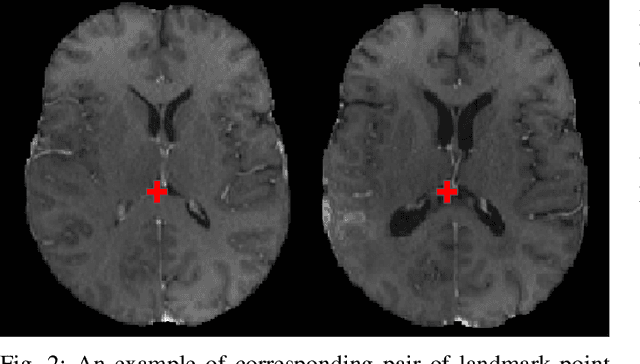
Abstract:Registration of longitudinal brain Magnetic Resonance Imaging (MRI) scans containing pathologies is challenging due to tissue appearance changes, and still an unsolved problem. This paper describes the first Brain Tumor Sequence Registration (BraTS-Reg) challenge, focusing on estimating correspondences between pre-operative and follow-up scans of the same patient diagnosed with a brain diffuse glioma. The BraTS-Reg challenge intends to establish a public benchmark environment for deformable registration algorithms. The associated dataset comprises de-identified multi-institutional multi-parametric MRI (mpMRI) data, curated for each scan's size and resolution, according to a common anatomical template. Clinical experts have generated extensive annotations of landmarks points within the scans, descriptive of distinct anatomical locations across the temporal domain. The training data along with these ground truth annotations will be released to participants to design and develop their registration algorithms, whereas the annotations for the validation and the testing data will be withheld by the organizers and used to evaluate the containerized algorithms of the participants. Each submitted algorithm will be quantitatively evaluated using several metrics, such as the Median Absolute Error (MAE), Robustness, and the Jacobian determinant.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge