Veronika A. Zimmer
Unbalanced optimal transport for robust longitudinal lesion evolution with registration-aware and appearance-guided priors
Feb 10, 2026Abstract:Evaluating lesion evolution in longitudinal CT scans of can cer patients is essential for assessing treatment response, yet establishing reliable lesion correspondence across time remains challenging. Standard bipartite matchers, which rely on geometric proximity, struggle when lesions appear, disappear, merge, or split. We propose a registration-aware matcher based on unbalanced optimal transport (UOT) that accommodates unequal lesion mass and adapts priors to patient-level tumor-load changes. Our transport cost blends (i) size-normalized geometry, (ii) local registration trust from the deformation-field Jacobian, and (iii) optional patch-level appearance consistency. The resulting transport plan is sparsified by relative pruning, yielding one-to-one matches as well as new, disappearing, merging, and splitting lesions without retraining or heuristic rules. On longitudinal CT data, our approach achieves consistently higher edge-detection precision and recall, improved lesion-state recall, and superior lesion-graph component F1 scores versus distance-only baselines.
From Model Based to Learned Regularization in Medical Image Registration: A Comprehensive Review
Dec 20, 2024Abstract:Image registration is fundamental in medical imaging applications, such as disease progression analysis or radiation therapy planning. The primary objective of image registration is to precisely capture the deformation between two or more images, typically achieved by minimizing an optimization problem. Due to its inherent ill-posedness, regularization is a key component in driving the solution toward anatomically meaningful deformations. A wide range of regularization methods has been proposed for both conventional and deep learning-based registration. However, the appropriate application of regularization techniques often depends on the specific registration problem, and no one-fits-all method exists. Despite its importance, regularization is often overlooked or addressed with default approaches, assuming existing methods are sufficient. A comprehensive and structured review remains missing. This review addresses this gap by introducing a novel taxonomy that systematically categorizes the diverse range of proposed regularization methods. It highlights the emerging field of learned regularization, which leverages data-driven techniques to automatically derive deformation properties from the data. Moreover, this review examines the transfer of regularization methods from conventional to learning-based registration, identifies open challenges, and outlines future research directions. By emphasizing the critical role of regularization in image registration, we hope to inspire the research community to reconsider regularization strategies in modern registration algorithms and to explore this rapidly evolving field further.
General Vision Encoder Features as Guidance in Medical Image Registration
Jul 18, 2024Abstract:General vision encoders like DINOv2 and SAM have recently transformed computer vision. Even though they are trained on natural images, such encoder models have excelled in medical imaging, e.g., in classification, segmentation, and registration. However, no in-depth comparison of different state-of-the-art general vision encoders for medical registration is available. In this work, we investigate how well general vision encoder features can be used in the dissimilarity metrics for medical image registration. We explore two encoders that were trained on natural images as well as one that was fine-tuned on medical data. We apply the features within the well-established B-spline FFD registration framework. In extensive experiments on cardiac cine MRI data, we find that using features as additional guidance for conventional metrics improves the registration quality. The code is available at github.com/compai-lab/2024-miccai-koegl.
Data-Driven Tissue- and Subject-Specific Elastic Regularization for Medical Image Registration
Jul 05, 2024



Abstract:Physics-inspired regularization is desired for intra-patient image registration since it can effectively capture the biomechanical characteristics of anatomical structures. However, a major challenge lies in the reliance on physical parameters: Parameter estimations vary widely across the literature, and the physical properties themselves are inherently subject-specific. In this work, we introduce a novel data-driven method that leverages hypernetworks to learn the tissue-dependent elasticity parameters of an elastic regularizer. Notably, our approach facilitates the estimation of patient-specific parameters without the need to retrain the network. We evaluate our method on three publicly available 2D and 3D lung CT and cardiac MR datasets. We find that with our proposed subject-specific tissue-dependent regularization, a higher registration quality is achieved across all datasets compared to using a global regularizer. The code is available at https://github.com/compai-lab/2024-miccai-reithmeir.
Learning Physics-Inspired Regularization for Medical Image Registration with Hypernetworks
Nov 14, 2023Abstract:Medical image registration aims at identifying the spatial deformation between images of the same anatomical region and is fundamental to image-based diagnostics and therapy. To date, the majority of the deep learning-based registration methods employ regularizers that enforce global spatial smoothness, e.g., the diffusion regularizer. However, such regularizers are not tailored to the data and might not be capable of reflecting the complex underlying deformation. In contrast, physics-inspired regularizers promote physically plausible deformations. One such regularizer is the linear elastic regularizer which models the deformation of elastic material. These regularizers are driven by parameters that define the material's physical properties. For biological tissue, a wide range of estimations of such parameters can be found in the literature and it remains an open challenge to identify suitable parameter values for successful registration. To overcome this problem and to incorporate physical properties into learning-based registration, we propose to use a hypernetwork that learns the effect of the physical parameters of a physics-inspired regularizer on the resulting spatial deformation field. In particular, we adapt the HyperMorph framework to learn the effect of the two elasticity parameters of the linear elastic regularizer. Our approach enables the efficient discovery of suitable, data-specific physical parameters at test time.
3D Arterial Segmentation via Single 2D Projections and Depth Supervision in Contrast-Enhanced CT Images
Sep 15, 2023Abstract:Automated segmentation of the blood vessels in 3D volumes is an essential step for the quantitative diagnosis and treatment of many vascular diseases. 3D vessel segmentation is being actively investigated in existing works, mostly in deep learning approaches. However, training 3D deep networks requires large amounts of manual 3D annotations from experts, which are laborious to obtain. This is especially the case for 3D vessel segmentation, as vessels are sparse yet spread out over many slices and disconnected when visualized in 2D slices. In this work, we propose a novel method to segment the 3D peripancreatic arteries solely from one annotated 2D projection per training image with depth supervision. We perform extensive experiments on the segmentation of peripancreatic arteries on 3D contrast-enhanced CT images and demonstrate how well we capture the rich depth information from 2D projections. We demonstrate that by annotating a single, randomly chosen projection for each training sample, we obtain comparable performance to annotating multiple 2D projections, thereby reducing the annotation effort. Furthermore, by mapping the 2D labels to the 3D space using depth information and incorporating this into training, we almost close the performance gap between 3D supervision and 2D supervision. Our code is available at: https://github.com/alinafdima/3Dseg-mip-depth.
MAD: Modality Agnostic Distance Measure for Image Registration
Sep 06, 2023Abstract:Multi-modal image registration is a crucial pre-processing step in many medical applications. However, it is a challenging task due to the complex intensity relationships between different imaging modalities, which can result in large discrepancy in image appearance. The success of multi-modal image registration, whether it is conventional or learning based, is predicated upon the choice of an appropriate distance (or similarity) measure. Particularly, deep learning registration algorithms lack in accuracy or even fail completely when attempting to register data from an "unseen" modality. In this work, we present Modality Agnostic Distance (MAD), a deep image distance}] measure that utilises random convolutions to learn the inherent geometry of the images while being robust to large appearance changes. Random convolutions are geometry-preserving modules which we use to simulate an infinite number of synthetic modalities alleviating the need for aligned paired data during training. We can therefore train MAD on a mono-modal dataset and successfully apply it to a multi-modal dataset. We demonstrate that not only can MAD affinely register multi-modal images successfully, but it has also a larger capture range than traditional measures such as Mutual Information and Normalised Gradient Fields.
A skeletonization algorithm for gradient-based optimization
Sep 05, 2023Abstract:The skeleton of a digital image is a compact representation of its topology, geometry, and scale. It has utility in many computer vision applications, such as image description, segmentation, and registration. However, skeletonization has only seen limited use in contemporary deep learning solutions. Most existing skeletonization algorithms are not differentiable, making it impossible to integrate them with gradient-based optimization. Compatible algorithms based on morphological operations and neural networks have been proposed, but their results often deviate from the geometry and topology of the true medial axis. This work introduces the first three-dimensional skeletonization algorithm that is both compatible with gradient-based optimization and preserves an object's topology. Our method is exclusively based on matrix additions and multiplications, convolutional operations, basic non-linear functions, and sampling from a uniform probability distribution, allowing it to be easily implemented in any major deep learning library. In benchmarking experiments, we prove the advantages of our skeletonization algorithm compared to non-differentiable, morphological, and neural-network-based baselines. Finally, we demonstrate the utility of our algorithm by integrating it with two medical image processing applications that use gradient-based optimization: deep-learning-based blood vessel segmentation, and multimodal registration of the mandible in computed tomography and magnetic resonance images.
Constructing Population-Specific Atlases from Whole Body MRI: Application to the UKBB
Aug 28, 2023Abstract:Population atlases are commonly utilised in medical imaging to facilitate the investigation of variability across populations. Such atlases enable the mapping of medical images into a common coordinate system, promoting comparability and enabling the study of inter-subject differences. Constructing such atlases becomes particularly challenging when working with highly heterogeneous datasets, such as whole-body images, where subjects show significant anatomical variations. In this work, we propose a pipeline for generating a standardised whole-body atlas for a highly heterogeneous population by partitioning the population into meaningful subgroups. We create six whole-body atlases that represent a healthy population average using magnetic resonance (MR) images from the UK Biobank dataset. We furthermore unbias them, and this way obtain a realistic representation of the population. In addition to the anatomical atlases, we generate probabilistic atlases that capture the distributions of abdominal fat and five abdominal organs across the population. We demonstrate different applications of these atlases, using the differences between subjects with medical conditions such as diabetes and cardiovascular diseases and healthy subjects from the atlas space. With this work, we make the constructed anatomical and label atlases publically available and anticipate them to support medical research conducted on whole-body MR images.
ICoNIK: Generating Respiratory-Resolved Abdominal MR Reconstructions Using Neural Implicit Representations in k-Space
Aug 17, 2023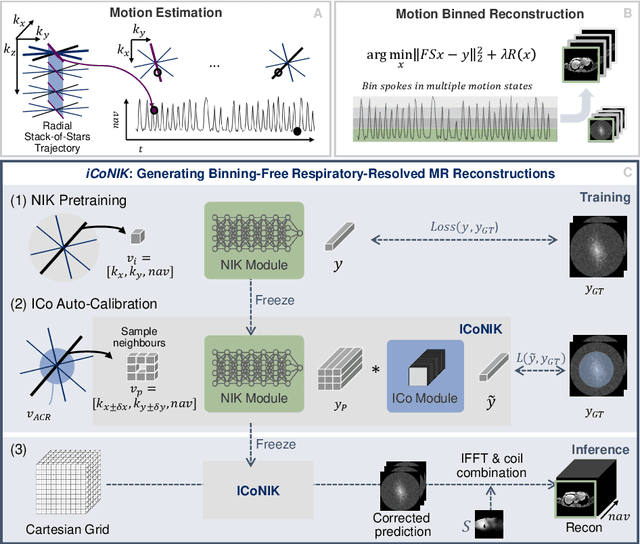
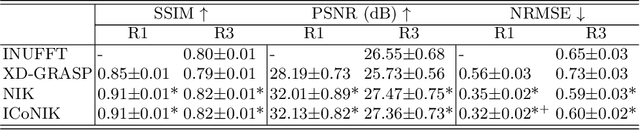
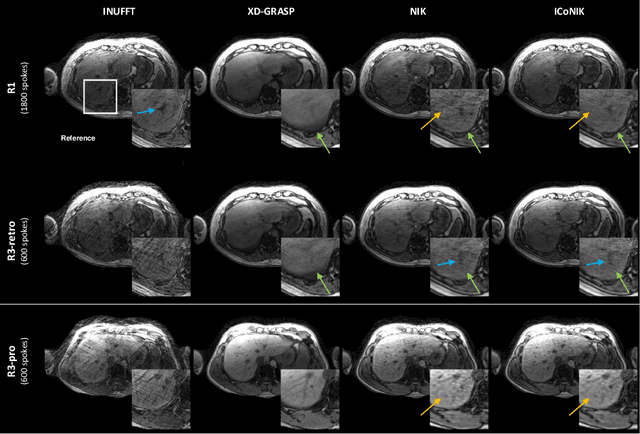
Abstract:Motion-resolved reconstruction for abdominal magnetic resonance imaging (MRI) remains a challenge due to the trade-off between residual motion blurring caused by discretized motion states and undersampling artefacts. In this work, we propose to generate blurring-free motion-resolved abdominal reconstructions by learning a neural implicit representation directly in k-space (NIK). Using measured sampling points and a data-derived respiratory navigator signal, we train a network to generate continuous signal values. To aid the regularization of sparsely sampled regions, we introduce an additional informed correction layer (ICo), which leverages information from neighboring regions to correct NIK's prediction. Our proposed generative reconstruction methods, NIK and ICoNIK, outperform standard motion-resolved reconstruction techniques and provide a promising solution to address motion artefacts in abdominal MRI.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge