Hanchen Wang
RIDA: A Robust Attack Framework on Incomplete Graphs
Jul 25, 2024Abstract:Graph Neural Networks (GNNs) are vital in data science but are increasingly susceptible to adversarial attacks. To help researchers develop more robust GNN models, it's essential to focus on designing strong attack models as foundational benchmarks and guiding references. Among adversarial attacks, gray-box poisoning attacks are noteworthy due to their effectiveness and fewer constraints. These attacks exploit GNNs' need for retraining on updated data, thereby impacting their performance by perturbing these datasets. However, current research overlooks the real-world scenario of incomplete graphs.To address this gap, we introduce the Robust Incomplete Deep Attack Framework (RIDA). It is the first algorithm for robust gray-box poisoning attacks on incomplete graphs. The approach innovatively aggregates distant vertex information and ensures powerful data utilization.Extensive tests against 9 SOTA baselines on 3 real-world datasets demonstrate RIDA's superiority in handling incompleteness and high attack performance on the incomplete graph.
APS-USCT: Ultrasound Computed Tomography on Sparse Data via AI-Physic Synergy
Jul 18, 2024



Abstract:Ultrasound computed tomography (USCT) is a promising technique that achieves superior medical imaging reconstruction resolution by fully leveraging waveform information, outperforming conventional ultrasound methods. Despite its advantages, high-quality USCT reconstruction relies on extensive data acquisition by a large number of transducers, leading to increased costs, computational demands, extended patient scanning times, and manufacturing complexities. To mitigate these issues, we propose a new USCT method called APS-USCT, which facilitates imaging with sparse data, substantially reducing dependence on high-cost dense data acquisition. Our APS-USCT method consists of two primary components: APS-wave and APS-FWI. The APS-wave component, an encoder-decoder system, preprocesses the waveform data, converting sparse data into dense waveforms to augment sample density prior to reconstruction. The APS-FWI component, utilizing the InversionNet, directly reconstructs the speed of sound (SOS) from the ultrasound waveform data. We further improve the model's performance by incorporating Squeeze-and-Excitation (SE) Blocks and source encoding techniques. Testing our method on a breast cancer dataset yielded promising results. It demonstrated outstanding performance with an average Structural Similarity Index (SSIM) of 0.8431. Notably, over 82% of samples achieved an SSIM above 0.8, with nearly 61% exceeding 0.85, highlighting the significant potential of our approach in improving USCT image reconstruction by efficiently utilizing sparse data.
STG-Mamba: Spatial-Temporal Graph Learning via Selective State Space Model
Mar 31, 2024



Abstract:Spatial-Temporal Graph (STG) data is characterized as dynamic, heterogenous, and non-stationary, leading to the continuous challenge of spatial-temporal graph learning. In the past few years, various GNN-based methods have been proposed to solely focus on mimicking the relationships among node individuals of the STG network, ignoring the significance of modeling the intrinsic features that exist in STG system over time. In contrast, modern Selective State Space Models (SSSMs) present a new approach which treat STG Network as a system, and meticulously explore the STG system's dynamic state evolution across temporal dimension. In this work, we introduce Spatial-Temporal Graph Mamba (STG-Mamba) as the first exploration of leveraging the powerful selective state space models for STG learning by treating STG Network as a system, and employing the Graph Selective State Space Block (GS3B) to precisely characterize the dynamic evolution of STG networks. STG-Mamba is formulated as an Encoder-Decoder architecture, which takes GS3B as the basic module, for efficient sequential data modeling. Furthermore, to strengthen GNN's ability of modeling STG data under the setting of SSSMs, we propose Kalman Filtering Graph Neural Networks (KFGN) for adaptive graph structure upgrading. KFGN smoothly fits in the context of selective state space evolution, and at the same time keeps linear complexity. Extensive empirical studies are conducted on three benchmark STG forecasting datasets, demonstrating the performance superiority and computational efficiency of STG-Mamba. It not only surpasses existing state-of-the-art methods in terms of STG forecasting performance, but also effectively alleviate the computational bottleneck of large-scale graph networks in reducing the computational cost of FLOPs and test inference time.
Unsupervised Discovery of Steerable Factors When Graph Deep Generative Models Are Entangled
Jan 29, 2024Abstract:Deep generative models (DGMs) have been widely developed for graph data. However, much less investigation has been carried out on understanding the latent space of such pretrained graph DGMs. These understandings possess the potential to provide constructive guidelines for crucial tasks, such as graph controllable generation. Thus in this work, we are interested in studying this problem and propose GraphCG, a method for the unsupervised discovery of steerable factors in the latent space of pretrained graph DGMs. We first examine the representation space of three pretrained graph DGMs with six disentanglement metrics, and we observe that the pretrained representation space is entangled. Motivated by this observation, GraphCG learns the steerable factors via maximizing the mutual information between semantic-rich directions, where the controlled graph moving along the same direction will share the same steerable factors. We quantitatively verify that GraphCG outperforms four competitive baselines on two graph DGMs pretrained on two molecule datasets. Additionally, we qualitatively illustrate seven steerable factors learned by GraphCG on five pretrained DGMs over five graph datasets, including two for molecules and three for point clouds.
A Physics-guided Generative AI Toolkit for Geophysical Monitoring
Jan 06, 2024Abstract:Full-waveform inversion (FWI) plays a vital role in geoscience to explore the subsurface. It utilizes the seismic wave to image the subsurface velocity map. As the machine learning (ML) technique evolves, the data-driven approaches using ML for FWI tasks have emerged, offering enhanced accuracy and reduced computational cost compared to traditional physics-based methods. However, a common challenge in geoscience, the unprivileged data, severely limits ML effectiveness. The issue becomes even worse during model pruning, a step essential in geoscience due to environmental complexities. To tackle this, we introduce the EdGeo toolkit, which employs a diffusion-based model guided by physics principles to generate high-fidelity velocity maps. The toolkit uses the acoustic wave equation to generate corresponding seismic waveform data, facilitating the fine-tuning of pruned ML models. Our results demonstrate significant improvements in SSIM scores and reduction in both MAE and MSE across various pruning ratios. Notably, the ML model fine-tuned using data generated by EdGeo yields superior quality of velocity maps, especially in representing unprivileged features, outperforming other existing methods.
Learned Full Waveform Inversion Incorporating Task Information for Ultrasound Computed Tomography
Aug 30, 2023Abstract:Ultrasound computed tomography (USCT) is an emerging imaging modality that holds great promise for breast imaging. Full-waveform inversion (FWI)-based image reconstruction methods incorporate accurate wave physics to produce high spatial resolution quantitative images of speed of sound or other acoustic properties of the breast tissues from USCT measurement data. However, the high computational cost of FWI reconstruction represents a significant burden for its widespread application in a clinical setting. The research reported here investigates the use of a convolutional neural network (CNN) to learn a mapping from USCT waveform data to speed of sound estimates. The CNN was trained using a supervised approach with a task-informed loss function aiming at preserving features of the image that are relevant to the detection of lesions. A large set of anatomically and physiologically realistic numerical breast phantoms (NBPs) and corresponding simulated USCT measurements was employed during training. Once trained, the CNN can perform real-time FWI image reconstruction from USCT waveform data. The performance of the proposed method was assessed and compared against FWI using a hold-out sample of 41 NBPs and corresponding USCT data. Accuracy was measured using relative mean square error (RMSE), structural self-similarity index measure (SSIM), and lesion detection performance (DICE score). This numerical experiment demonstrates that a supervised learning model can achieve accuracy comparable to FWI in terms of RMSE and SSIM, and better performance in terms of task performance, while significantly reducing computational time.
Does Full Waveform Inversion Benefit from Big Data?
Jul 28, 2023Abstract:This paper investigates the impact of big data on deep learning models for full waveform inversion (FWI). While it is well known that big data can boost the performance of deep learning models in many tasks, its effectiveness has not been validated for FWI. To address this gap, we present an empirical study that investigates how deep learning models in FWI behave when trained on OpenFWI, a collection of large-scale, multi-structural datasets published recently. Particularly, we train and evaluate the FWI models on a combination of 10 2D subsets in OpenFWI that contain 470K data pairs in total. Our experiments demonstrate that larger datasets lead to better performance and generalization of deep learning models for FWI. We further demonstrate that model capacity needs to scale in accordance with data size for optimal improvement.
$\mathbf{\mathbb{E}^{FWI}}$: Multi-parameter Benchmark Datasets for Elastic Full Waveform Inversion of Geophysical Properties
Jun 21, 2023



Abstract:Elastic geophysical properties (such as P- and S-wave velocities) are of great importance to various subsurface applications like CO$_2$ sequestration and energy exploration (e.g., hydrogen and geothermal). Elastic full waveform inversion (FWI) is widely applied for characterizing reservoir properties. In this paper, we introduce $\mathbf{\mathbb{E}^{FWI}}$, a comprehensive benchmark dataset that is specifically designed for elastic FWI. $\mathbf{\mathbb{E}^{FWI}}$ encompasses 8 distinct datasets that cover diverse subsurface geologic structures (flat, curve, faults, etc). The benchmark results produced by three different deep learning methods are provided. In contrast to our previously presented dataset (pressure recordings) for acoustic FWI (referred to as OpenFWI), the seismic dataset in $\mathbf{\mathbb{E}^{FWI}}$ has both vertical and horizontal components. Moreover, the velocity maps in $\mathbf{\mathbb{E}^{FWI}}$ incorporate both P- and S-wave velocities. While the multicomponent data and the added S-wave velocity make the data more realistic, more challenges are introduced regarding the convergence and computational cost of the inversion. We conduct comprehensive numerical experiments to explore the relationship between P-wave and S-wave velocities in seismic data. The relation between P- and S-wave velocities provides crucial insights into the subsurface properties such as lithology, porosity, fluid content, etc. We anticipate that $\mathbf{\mathbb{E}^{FWI}}$ will facilitate future research on multiparameter inversions and stimulate endeavors in several critical research topics of carbon-zero and new energy exploration. All datasets, codes and relevant information can be accessed through our website at https://efwi-lanl.github.io/
Solving Seismic Wave Equations on Variable Velocity Models with Fourier Neural Operator
Sep 25, 2022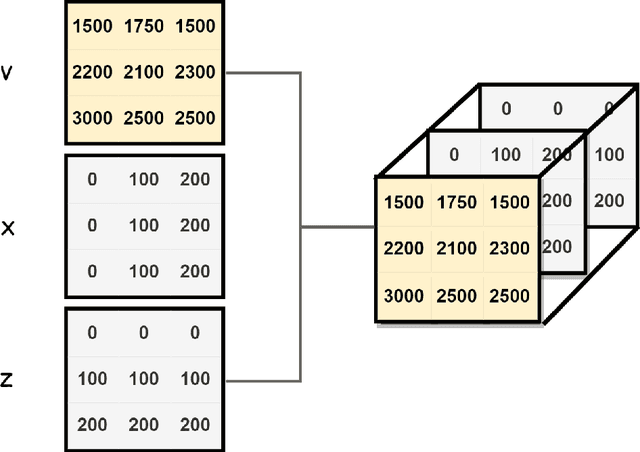
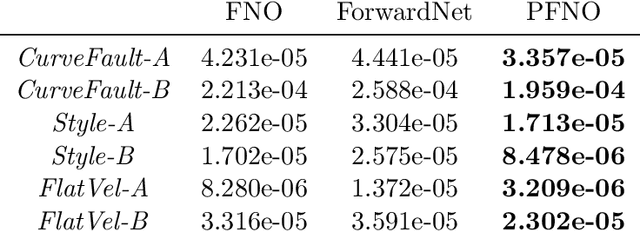
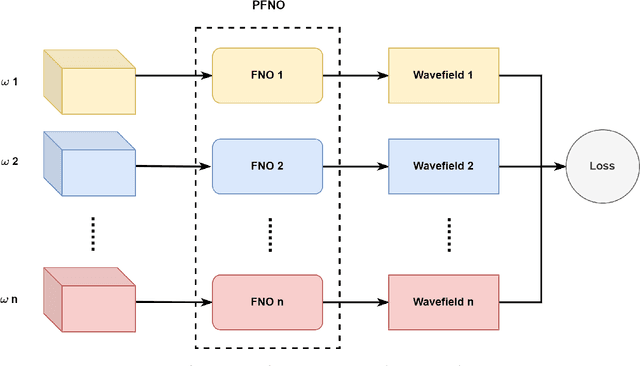

Abstract:In the study of subsurface seismic imaging, solving the acoustic wave equation is a pivotal component in existing models. With the advancement of deep learning, neural networks are applied to numerically solve partial differential equations by learning the mapping between the inputs and the solution of the equation, the wave equation in particular, since traditional methods can be time consuming if numerous instances are to be solved. Previous works that concentrate on solving the wave equation by neural networks consider either a single velocity model or multiple simple velocity models, which is restricted in practice. Therefore, inspired by the idea of operator learning, this work leverages the Fourier neural operator (FNO) to effectively learn the frequency domain seismic wavefields under the context of variable velocity models. Moreover, we propose a new framework paralleled Fourier neural operator (PFNO) for efficiently training the FNO-based solver given multiple source locations and frequencies. Numerical experiments demonstrate the high accuracy of both FNO and PFNO with complicated velocity models in the OpenFWI datasets. Furthermore, the cross-dataset generalization test verifies that PFNO adapts to out-of-distribution velocity models. Also, PFNO has robust performance in the presence of random noise in the labels. Finally, PFNO admits higher computational efficiency on large-scale testing datasets, compared with the traditional finite-difference method. The aforementioned advantages endow the FNO-based solver with the potential to build powerful models for research on seismic waves.
Evaluating Self-Supervised Learning for Molecular Graph Embeddings
Jun 16, 2022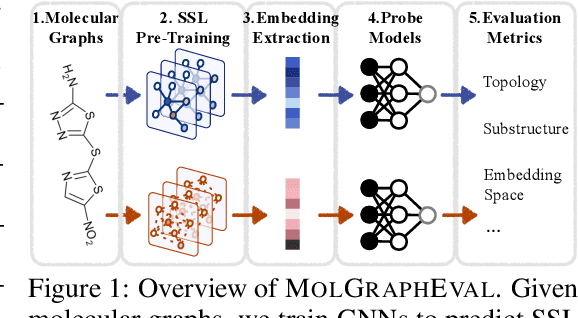
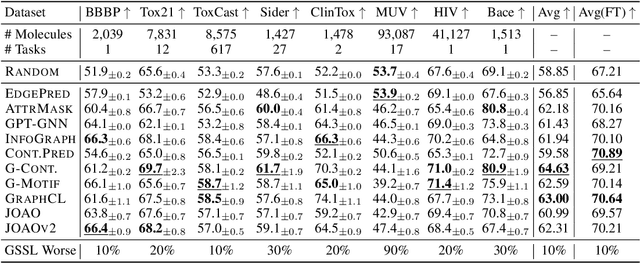
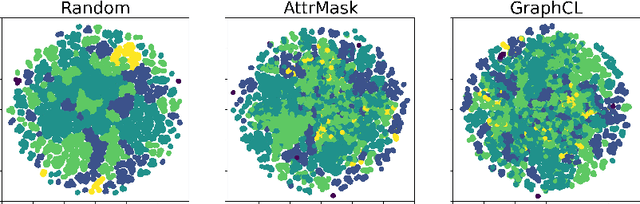
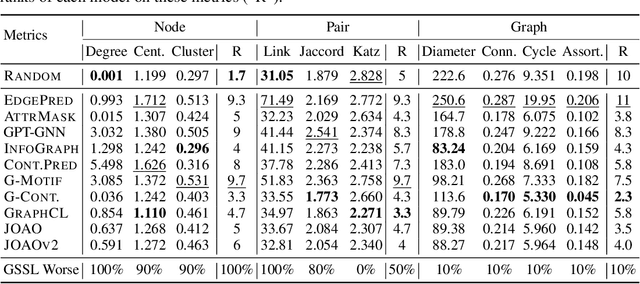
Abstract:Graph Self-Supervised Learning (GSSL) paves the way for learning graph embeddings without expert annotation, which is particularly impactful for molecular graphs since the number of possible molecules is enormous and labels are expensive to obtain. However, by design, GSSL methods are not trained to perform well on one downstream task but aim for transferability to many, making evaluating them less straightforward. As a step toward obtaining profiles of molecular graph embeddings with diverse and interpretable attributes, we introduce Molecular Graph Representation Evaluation (MolGraphEval), a suite of probe tasks, categorised into (i) topological-, (ii) substructure-, and (iii) embedding space properties. By benchmarking existing GSSL methods on both existing downstream datasets and MolGraphEval, we discover surprising discrepancies between conclusions drawn from existing datasets alone versus more fine-grained probing, suggesting that current evaluation protocols do not provide the whole picture. Our modular, automated end-to-end GSSL pipeline code will be released upon acceptance, including standardised graph loading, experiment management, and embedding evaluation.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge