Matt Kusner
Uncertainty Quantification of Pre-Trained and Fine-Tuned Surrogate Models using Conformal Prediction
Aug 19, 2024



Abstract:Data-driven surrogate models have shown immense potential as quick, inexpensive approximations to complex numerical and experimental modelling tasks. However, most surrogate models characterising physical systems do not quantify their uncertainty, rendering their predictions unreliable, and needing further validation. Though Bayesian approximations offer some solace in estimating the error associated with these models, they cannot provide they cannot provide guarantees, and the quality of their inferences depends on the availability of prior information and good approximations to posteriors for complex problems. This is particularly pertinent to multi-variable or spatio-temporal problems. Our work constructs and formalises a conformal prediction framework that satisfies marginal coverage for spatio-temporal predictions in a model-agnostic manner, requiring near-zero computational costs. The paper provides an extensive empirical study of the application of the framework to ascertain valid error bars that provide guaranteed coverage across the surrogate model's domain of operation. The application scope of our work extends across a large range of spatio-temporal models, ranging from solving partial differential equations to weather forecasting. Through the applications, the paper looks at providing statistically valid error bars for deterministic models, as well as crafting guarantees to the error bars of probabilistic models. The paper concludes with a viable conformal prediction formalisation that provides guaranteed coverage of the surrogate model, regardless of model architecture, and its training regime and is unbothered by the curse of dimensionality.
Valid Error Bars for Neural Weather Models using Conformal Prediction
Jun 20, 2024



Abstract:Neural weather models have shown immense potential as inexpensive and accurate alternatives to physics-based models. However, most models trained to perform weather forecasting do not quantify the uncertainty associated with their forecasts. This limits the trust in the model and the usefulness of the forecasts. In this work we construct and formalise a conformal prediction framework as a post-processing method for estimating this uncertainty. The method is model-agnostic and gives calibrated error bounds for all variables, lead times and spatial locations. No modifications are required to the model and the computational cost is negligible compared to model training. We demonstrate the usefulness of the conformal prediction framework on a limited area neural weather model for the Nordic region. We further explore the advantages of the framework for deterministic and probabilistic models.
Plasma Surrogate Modelling using Fourier Neural Operators
Nov 10, 2023Abstract:Predicting plasma evolution within a Tokamak reactor is crucial to realizing the goal of sustainable fusion. Capabilities in forecasting the spatio-temporal evolution of plasma rapidly and accurately allow us to quickly iterate over design and control strategies on current Tokamak devices and future reactors. Modelling plasma evolution using numerical solvers is often expensive, consuming many hours on supercomputers, and hence, we need alternative inexpensive surrogate models. We demonstrate accurate predictions of plasma evolution both in simulation and experimental domains using deep learning-based surrogate modelling tools, viz., Fourier Neural Operators (FNO). We show that FNO has a speedup of six orders of magnitude over traditional solvers in predicting the plasma dynamics simulated from magnetohydrodynamic models, while maintaining a high accuracy (MSE $\approx$ $10^{-5}$). Our modified version of the FNO is capable of solving multi-variable Partial Differential Equations (PDE), and can capture the dependence among the different variables in a single model. FNOs can also predict plasma evolution on real-world experimental data observed by the cameras positioned within the MAST Tokamak, i.e., cameras looking across the central solenoid and the divertor in the Tokamak. We show that FNOs are able to accurately forecast the evolution of plasma and have the potential to be deployed for real-time monitoring. We also illustrate their capability in forecasting the plasma shape, the locations of interactions of the plasma with the central solenoid and the divertor for the full duration of the plasma shot within MAST. The FNO offers a viable alternative for surrogate modelling as it is quick to train and infer, and requires fewer data points, while being able to do zero-shot super-resolution and getting high-fidelity solutions.
Causal Inference with Treatment Measurement Error: A Nonparametric Instrumental Variable Approach
Jun 18, 2022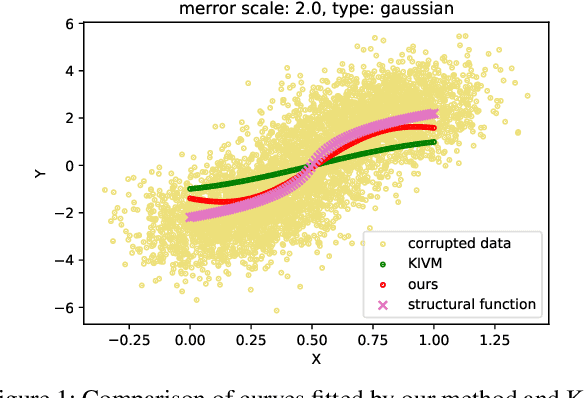
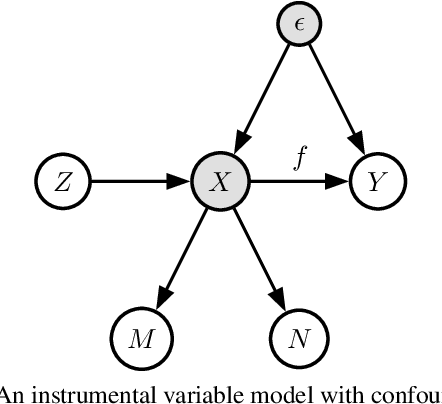
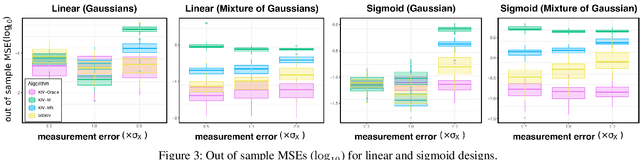
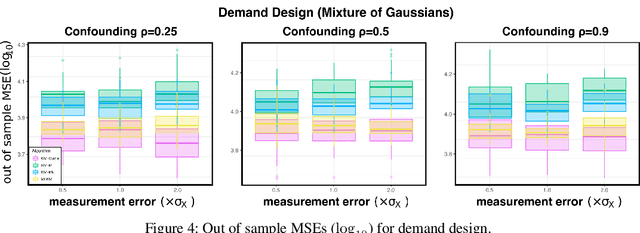
Abstract:We propose a kernel-based nonparametric estimator for the causal effect when the cause is corrupted by error. We do so by generalizing estimation in the instrumental variable setting. Despite significant work on regression with measurement error, additionally handling unobserved confounding in the continuous setting is non-trivial: we have seen little prior work. As a by-product of our investigation, we clarify a connection between mean embeddings and characteristic functions, and how learning one simultaneously allows one to learn the other. This opens the way for kernel method research to leverage existing results in characteristic function estimation. Finally, we empirically show that our proposed method, MEKIV, improves over baselines and is robust under changes in the strength of measurement error and to the type of error distributions.
Evaluating Self-Supervised Learning for Molecular Graph Embeddings
Jun 16, 2022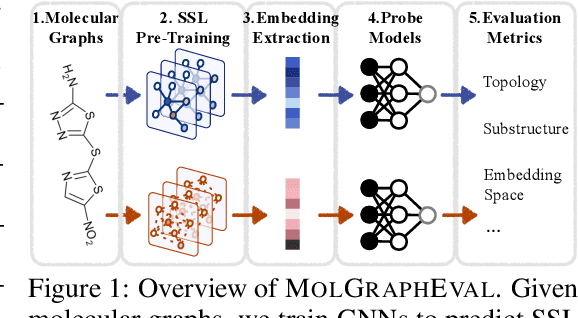
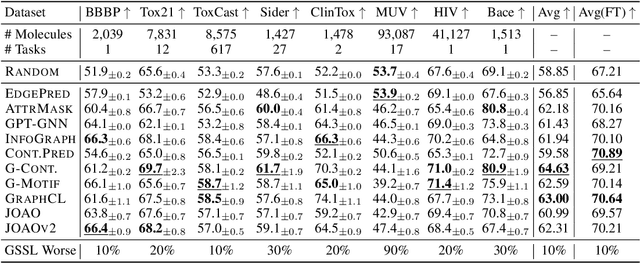
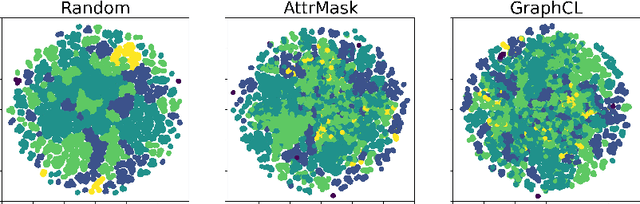
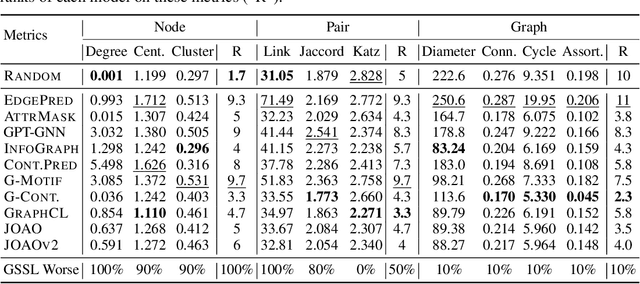
Abstract:Graph Self-Supervised Learning (GSSL) paves the way for learning graph embeddings without expert annotation, which is particularly impactful for molecular graphs since the number of possible molecules is enormous and labels are expensive to obtain. However, by design, GSSL methods are not trained to perform well on one downstream task but aim for transferability to many, making evaluating them less straightforward. As a step toward obtaining profiles of molecular graph embeddings with diverse and interpretable attributes, we introduce Molecular Graph Representation Evaluation (MolGraphEval), a suite of probe tasks, categorised into (i) topological-, (ii) substructure-, and (iii) embedding space properties. By benchmarking existing GSSL methods on both existing downstream datasets and MolGraphEval, we discover surprising discrepancies between conclusions drawn from existing datasets alone versus more fine-grained probing, suggesting that current evaluation protocols do not provide the whole picture. Our modular, automated end-to-end GSSL pipeline code will be released upon acceptance, including standardised graph loading, experiment management, and embedding evaluation.
Stochastic Causal Programming for Bounding Treatment Effects
Feb 22, 2022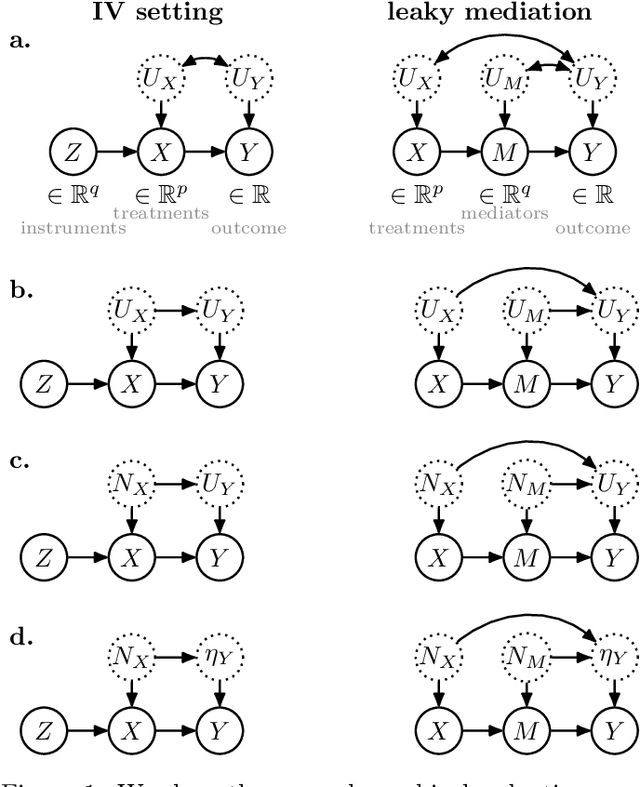
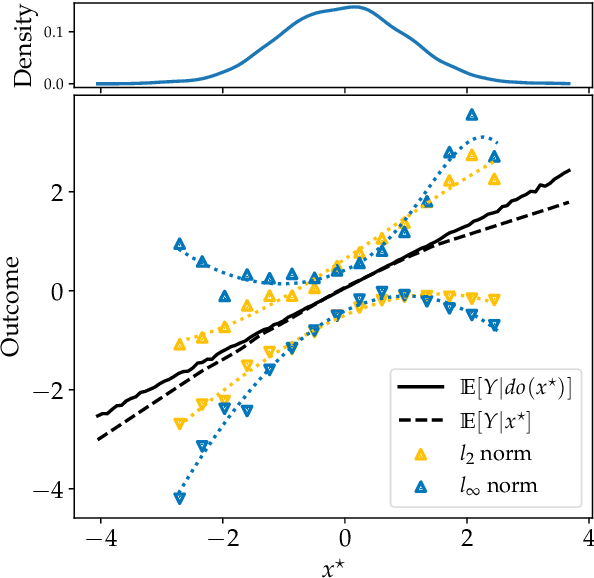
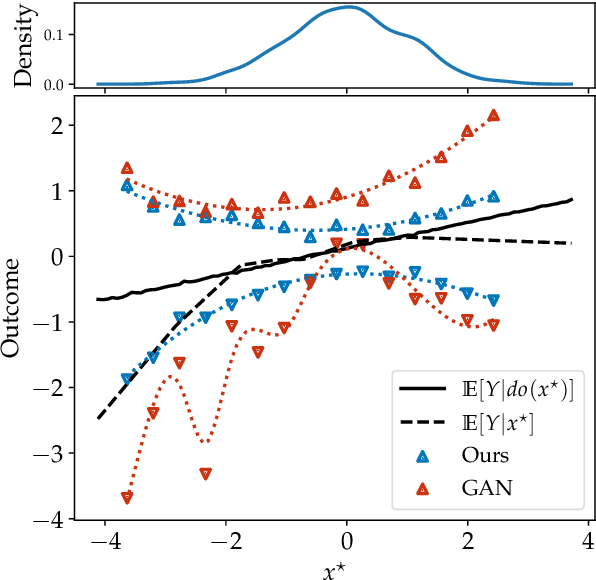
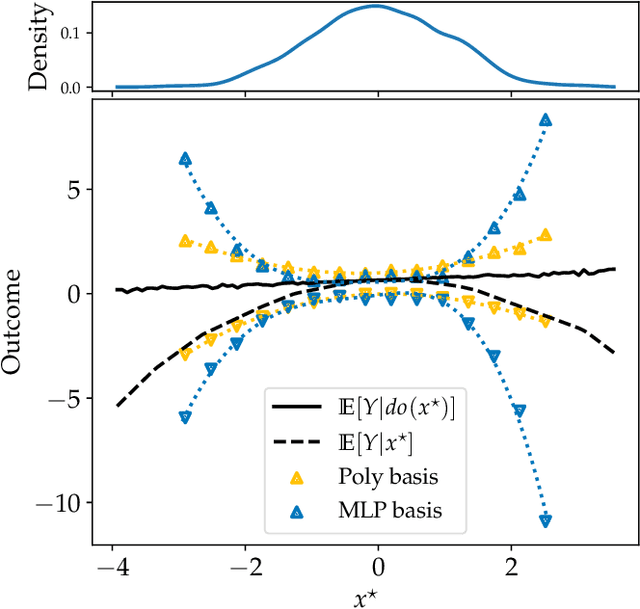
Abstract:Causal effect estimation is important for numerous tasks in the natural and social sciences. However, identifying effects is impossible from observational data without making strong, often untestable assumptions. We consider algorithms for the partial identification problem, bounding treatment effects from multivariate, continuous treatments over multiple possible causal models when unmeasured confounding makes identification impossible. We consider a framework where observable evidence is matched to the implications of constraints encoded in a causal model by norm-based criteria. This generalizes classical approaches based purely on generative models. Casting causal effects as objective functions in a constrained optimization problem, we combine flexible learning algorithms with Monte Carlo methods to implement a family of solutions under the name of stochastic causal programming. In particular, we present ways by which such constrained optimization problems can be parameterized without likelihood functions for the causal or the observed data model, reducing the computational and statistical complexity of the task.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge