"Information Extraction": models, code, and papers
Audio compression-assisted feature extraction for voice replay attack detection
Oct 10, 2023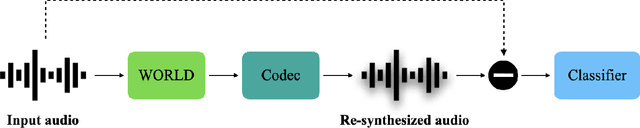
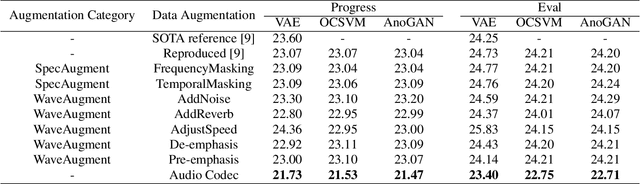
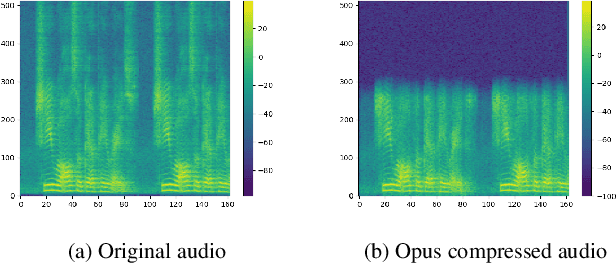
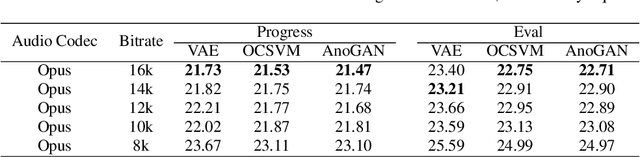
Replay attack is one of the most effective and simplest voice spoofing attacks. Detecting replay attacks is challenging, according to the Automatic Speaker Verification Spoofing and Countermeasures Challenge 2021 (ASVspoof 2021), because they involve a loudspeaker, a microphone, and acoustic conditions (e.g., background noise). One obstacle to detecting replay attacks is finding robust feature representations that reflect the channel noise information added to the replayed speech. This study proposes a feature extraction approach that uses audio compression for assistance. Audio compression compresses audio to preserve content and speaker information for transmission. The missed information after decompression is expected to contain content- and speaker-independent information (e.g., channel noise added during the replay process). We conducted a comprehensive experiment with a few data augmentation techniques and 3 classifiers on the ASVspoof 2021 physical access (PA) set and confirmed the effectiveness of the proposed feature extraction approach. To the best of our knowledge, the proposed approach achieves the lowest EER at 22.71% on the ASVspoof 2021 PA evaluation set.
Enhancing Intra-class Information Extraction for Heterophilous Graphs: One Neural Architecture Search Approach
Nov 20, 2022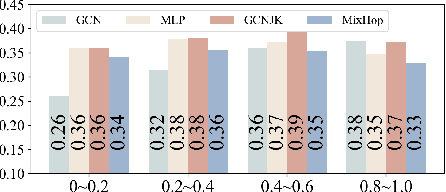
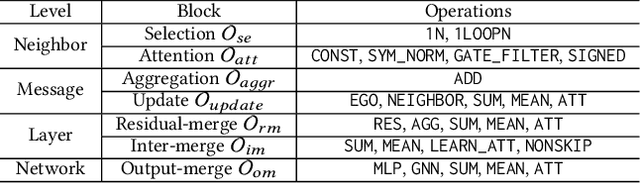
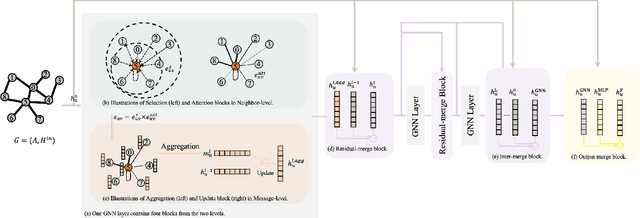
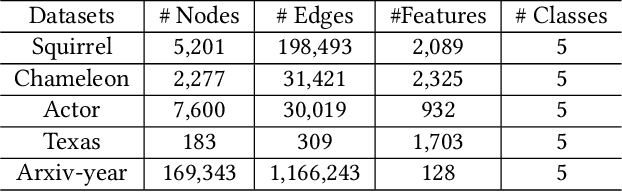
In recent years, Graph Neural Networks (GNNs) have been popular in graph representation learning which assumes the homophily property, i.e., the connected nodes have the same label or have similar features. However, they may fail to generalize into the heterophilous graphs which in the low/medium level of homophily. Existing methods tend to address this problem by enhancing the intra-class information extraction, i.e., either by designing better GNNs to improve the model effectiveness, or re-designing the graph structures to incorporate more potential intra-class nodes from distant hops. Despite the success, we observe two aspects that can be further improved: (a) enhancing the ego feature information extraction from node itself which is more reliable in extracting the intra-class information; (b) designing node-wise GNNs can better adapt to the nodes with different homophily ratios. In this paper, we propose a novel method IIE-GNN (Intra-class Information Enhanced Graph Neural Networks) to achieve two improvements. A unified framework is proposed based on the literature, in which the intra-class information from the node itself and neighbors can be extracted based on seven carefully designed blocks. With the help of neural architecture search (NAS), we propose a novel search space based on the framework, and then provide an architecture predictor to design GNNs for each node. We further conduct experiments to show that IIE-GNN can improve the model performance by designing node-wise GNNs to enhance intra-class information extraction.
GenIE: Generative Information Extraction
Dec 15, 2021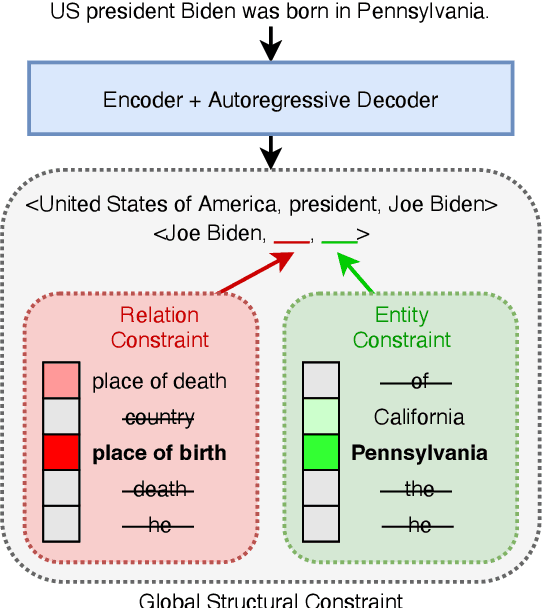
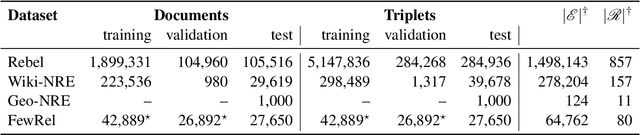
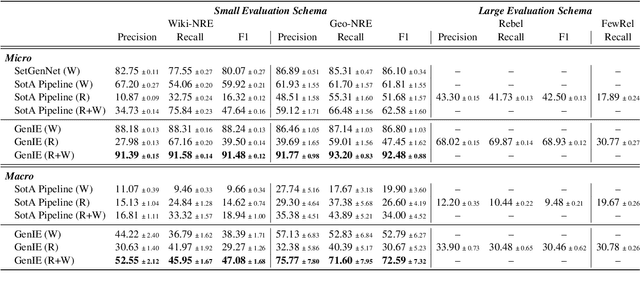
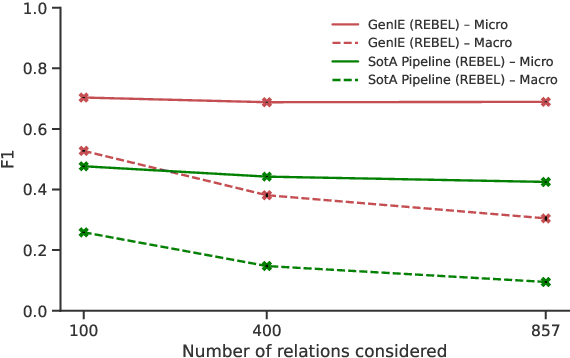
Structured and grounded representation of text is typically formalized by closed information extraction, the problem of extracting an exhaustive set of (subject, relation, object) triplets that are consistent with a predefined set of entities and relations from a knowledge base schema. Most existing works are pipelines prone to error accumulation, and all approaches are only applicable to unrealistically small numbers of entities and relations. We introduce GenIE (generative information extraction), the first end-to-end autoregressive formulation of closed information extraction. GenIE naturally exploits the language knowledge from the pre-trained transformer by autoregressively generating relations and entities in textual form. Thanks to a new bi-level constrained generation strategy, only triplets consistent with the predefined knowledge base schema are produced. Our experiments show that GenIE is state-of-the-art on closed information extraction, generalizes from fewer training data points than baselines, and scales to a previously unmanageable number of entities and relations. With this work, closed information extraction becomes practical in realistic scenarios, providing new opportunities for downstream tasks. Finally, this work paves the way towards a unified end-to-end approach to the core tasks of information extraction. Code and models available at https://github.com/epfl-dlab/GenIE.
Time for aCTIon: Automated Analysis of Cyber Threat Intelligence in the Wild
Jul 14, 2023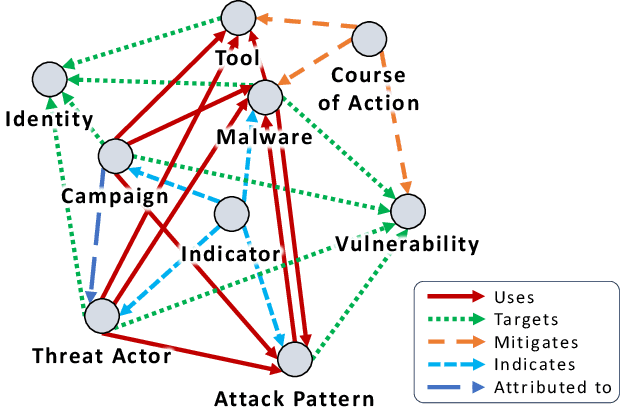
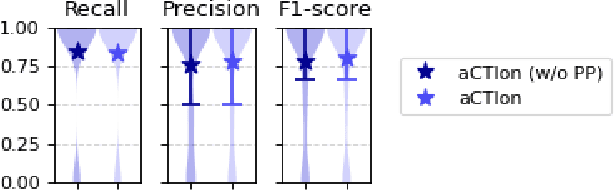
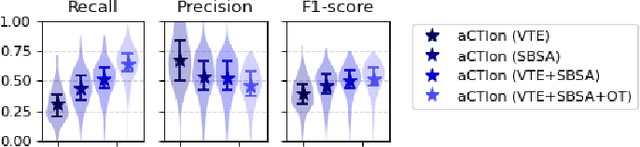
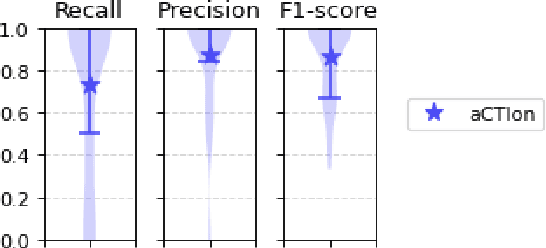
Cyber Threat Intelligence (CTI) plays a crucial role in assessing risks and enhancing security for organizations. However, the process of extracting relevant information from unstructured text sources can be expensive and time-consuming. Our empirical experience shows that existing tools for automated structured CTI extraction have performance limitations. Furthermore, the community lacks a common benchmark to quantitatively assess their performance. We fill these gaps providing a new large open benchmark dataset and aCTIon, a structured CTI information extraction tool. The dataset includes 204 real-world publicly available reports and their corresponding structured CTI information in STIX format. Our team curated the dataset involving three independent groups of CTI analysts working over the course of several months. To the best of our knowledge, this dataset is two orders of magnitude larger than previously released open source datasets. We then design aCTIon, leveraging recently introduced large language models (GPT3.5) in the context of two custom information extraction pipelines. We compare our method with 10 solutions presented in previous work, for which we develop our own implementations when open-source implementations were lacking. Our results show that aCTIon outperforms previous work for structured CTI extraction with an improvement of the F1-score from 10%points to 50%points across all tasks.
Spectral Neural Networks: Approximation Theory and Optimization Landscape
Oct 01, 2023There is a large variety of machine learning methodologies that are based on the extraction of spectral geometric information from data. However, the implementations of many of these methods often depend on traditional eigensolvers, which present limitations when applied in practical online big data scenarios. To address some of these challenges, researchers have proposed different strategies for training neural networks as alternatives to traditional eigensolvers, with one such approach known as Spectral Neural Network (SNN). In this paper, we investigate key theoretical aspects of SNN. First, we present quantitative insights into the tradeoff between the number of neurons and the amount of spectral geometric information a neural network learns. Second, we initiate a theoretical exploration of the optimization landscape of SNN's objective to shed light on the training dynamics of SNN. Unlike typical studies of convergence to global solutions of NN training dynamics, SNN presents an additional complexity due to its non-convex ambient loss function.
Mutual Reinforcement Effects in Japanese Sentence Classification and Named Entity Recognition Tasks
Jul 21, 2023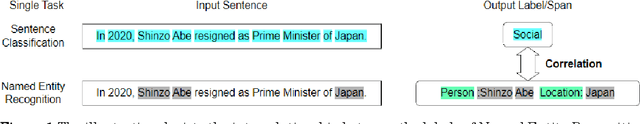
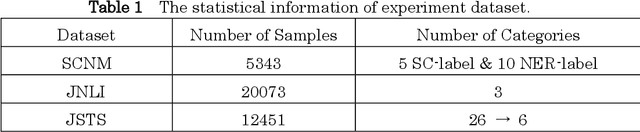

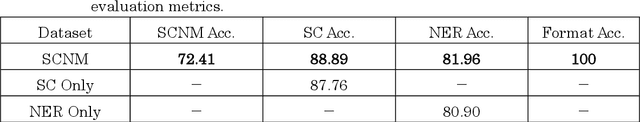
Information extraction(IE) is a crucial subfield within natural language processing. However, for the traditionally segmented approach to sentence classification and Named Entity Recognition, the intricate interactions between these individual subtasks remain largely uninvestigated. In this study, we propose an integrative analysis, converging sentence classification with Named Entity Recognition, with the objective to unveil and comprehend the mutual reinforcement effect within these two information extraction subtasks. To achieve this, we introduce a Sentence Classification and Named Entity Recognition Multi-task (SCNM) approach that combines Sentence Classification (SC) and Named Entity Recognition (NER). We develop a Sentence-to-Label Generation (SLG) framework for SCNM and construct a Wikipedia dataset containing both SC and NER. Using a format converter, we unify input formats and employ a generative model to generate SC-labels, NER-labels, and associated text segments. We propose a Constraint Mechanism (CM) to improve generated format accuracy. Our results show SC accuracy increased by 1.13 points and NER by 1.06 points in SCNM compared to standalone tasks, with CM raising format accuracy from 63.61 to 100. The findings indicate mutual reinforcement effects between SC and NER, and integration enhances both tasks' performance. We additionally implemented the SLG framework on single SC task. It yielded superior accuracies compared to the baseline on two distinct Japanese SC datasets. Notably, in the experiment of few-shot learning, SLG framework shows much better performance than fine-tune method. These empirical findings contribute additional evidence to affirm the efficacy of the SLG framework.
Syntactically Robust Training on Partially-Observed Data for Open Information Extraction
Jan 17, 2023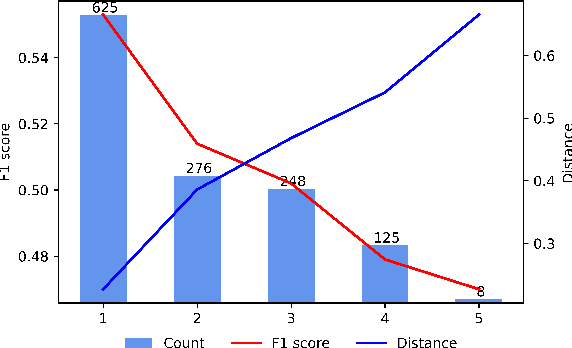
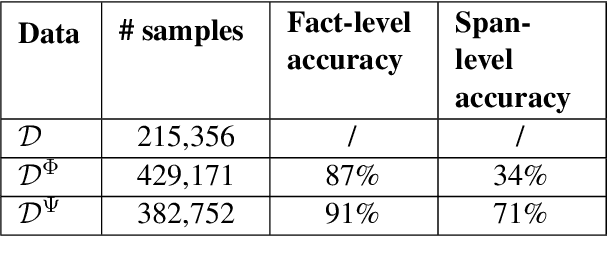
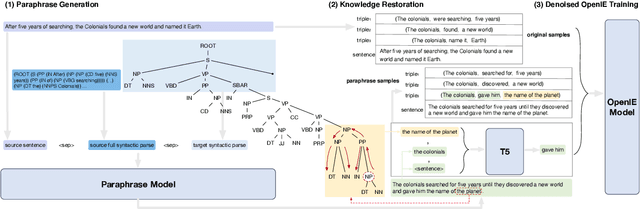
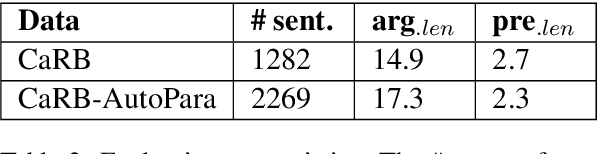
Open Information Extraction models have shown promising results with sufficient supervision. However, these models face a fundamental challenge that the syntactic distribution of training data is partially observable in comparison to the real world. In this paper, we propose a syntactically robust training framework that enables models to be trained on a syntactic-abundant distribution based on diverse paraphrase generation. To tackle the intrinsic problem of knowledge deformation of paraphrasing, two algorithms based on semantic similarity matching and syntactic tree walking are used to restore the expressionally transformed knowledge. The training framework can be generally applied to other syntactic partial observable domains. Based on the proposed framework, we build a new evaluation set called CaRB-AutoPara, a syntactically diverse dataset consistent with the real-world setting for validating the robustness of the models. Experiments including a thorough analysis show that the performance of the model degrades with the increase of the difference in syntactic distribution, while our framework gives a robust boundary. The source code is publicly available at https://github.com/qijimrc/RobustOIE.
Gradient Imitation Reinforcement Learning for General Low-Resource Information Extraction
Nov 14, 2022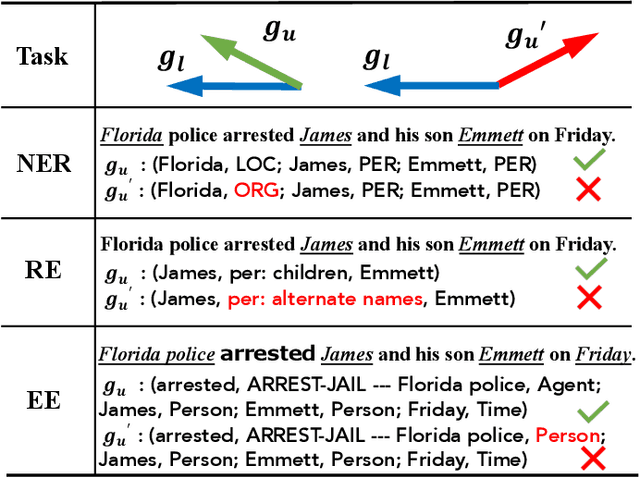
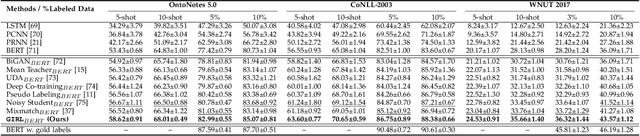
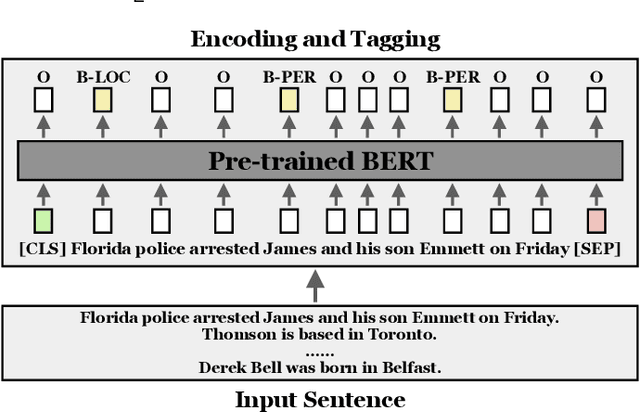
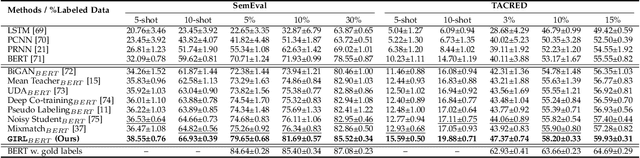
Information Extraction (IE) aims to extract structured information from heterogeneous sources. IE from natural language texts include sub-tasks such as Named Entity Recognition (NER), Relation Extraction (RE), and Event Extraction (EE). Most IE systems require comprehensive understandings of sentence structure, implied semantics, and domain knowledge to perform well; thus, IE tasks always need adequate external resources and annotations. However, it takes time and effort to obtain more human annotations. Low-Resource Information Extraction (LRIE) strives to use unsupervised data, reducing the required resources and human annotation. In practice, existing systems either utilize self-training schemes to generate pseudo labels that will cause the gradual drift problem, or leverage consistency regularization methods which inevitably possess confirmation bias. To alleviate confirmation bias due to the lack of feedback loops in existing LRIE learning paradigms, we develop a Gradient Imitation Reinforcement Learning (GIRL) method to encourage pseudo-labeled data to imitate the gradient descent direction on labeled data, which can force pseudo-labeled data to achieve better optimization capabilities similar to labeled data. Based on how well the pseudo-labeled data imitates the instructive gradient descent direction obtained from labeled data, we design a reward to quantify the imitation process and bootstrap the optimization capability of pseudo-labeled data through trial and error. In addition to learning paradigms, GIRL is not limited to specific sub-tasks, and we leverage GIRL to solve all IE sub-tasks (named entity recognition, relation extraction, and event extraction) in low-resource settings (semi-supervised IE and few-shot IE).
Business Document Information Extraction: Towards Practical Benchmarks
Jun 20, 2022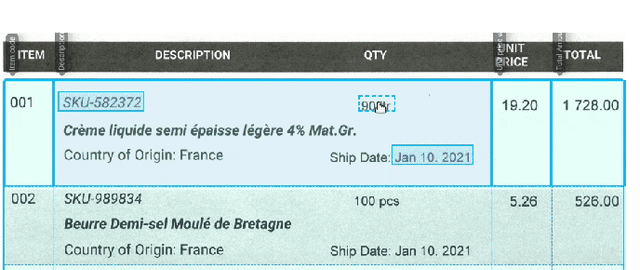
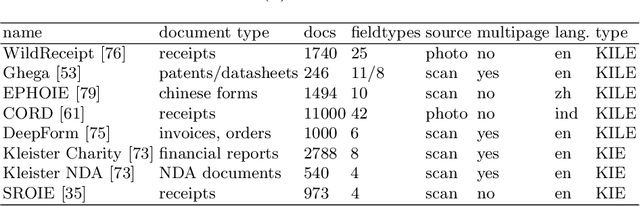
Information extraction from semi-structured documents is crucial for frictionless business-to-business (B2B) communication. While machine learning problems related to Document Information Extraction (IE) have been studied for decades, many common problem definitions and benchmarks do not reflect domain-specific aspects and practical needs for automating B2B document communication. We review the landscape of Document IE problems, datasets and benchmarks. We highlight the practical aspects missing in the common definitions and define the Key Information Localization and Extraction (KILE) and Line Item Recognition (LIR) problems. There is a lack of relevant datasets and benchmarks for Document IE on semi-structured business documents as their content is typically legally protected or sensitive. We discuss potential sources of available documents including synthetic data.
TIER-A: Denoising Learning Framework for Information Extraction
Nov 13, 2022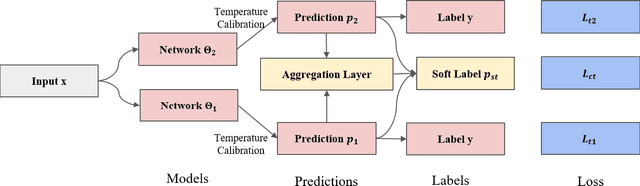
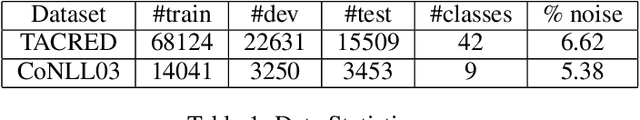
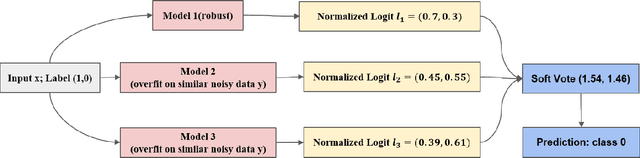
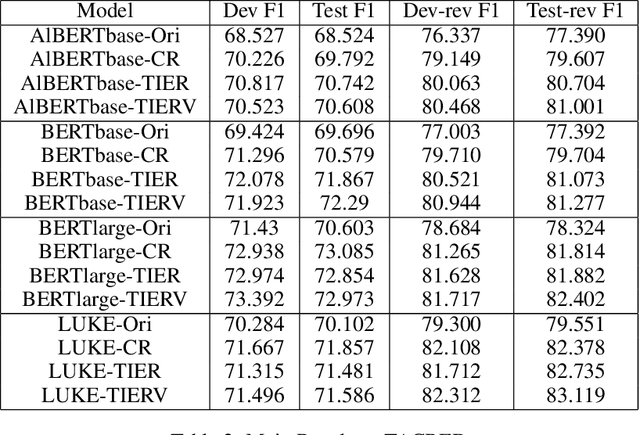
With the development of deep neural language models, great progress has been made in information extraction recently. However, deep learning models often overfit on noisy data points, leading to poor performance. In this work, we examine the role of information entropy in the overfitting process and draw a key insight that overfitting is a process of overconfidence and entropy decreasing. Motivated by such properties, we propose a simple yet effective co-regularization joint-training framework TIER-A, Aggregation Joint-training Framework with Temperature Calibration and Information Entropy Regularization. Our framework consists of several neural models with identical structures. These models are jointly trained and we avoid overfitting by introducing temperature and information entropy regularization. Extensive experiments on two widely-used but noisy datasets, TACRED and CoNLL03, demonstrate the correctness of our assumption and the effectiveness of our framework.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge