"Image": models, code, and papers
Semi-supervised Medical Image Classification with Relation-driven Self-ensembling Model
May 15, 2020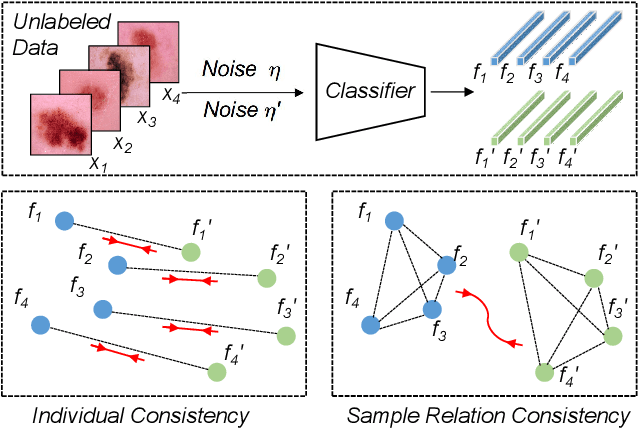
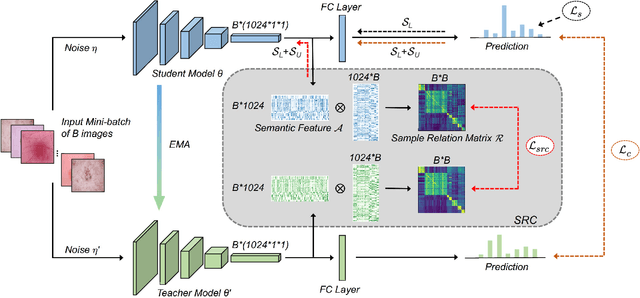
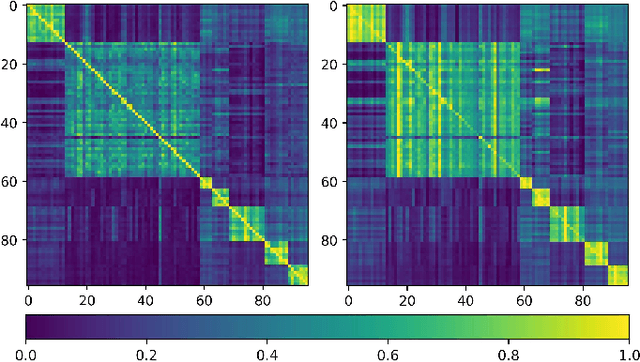
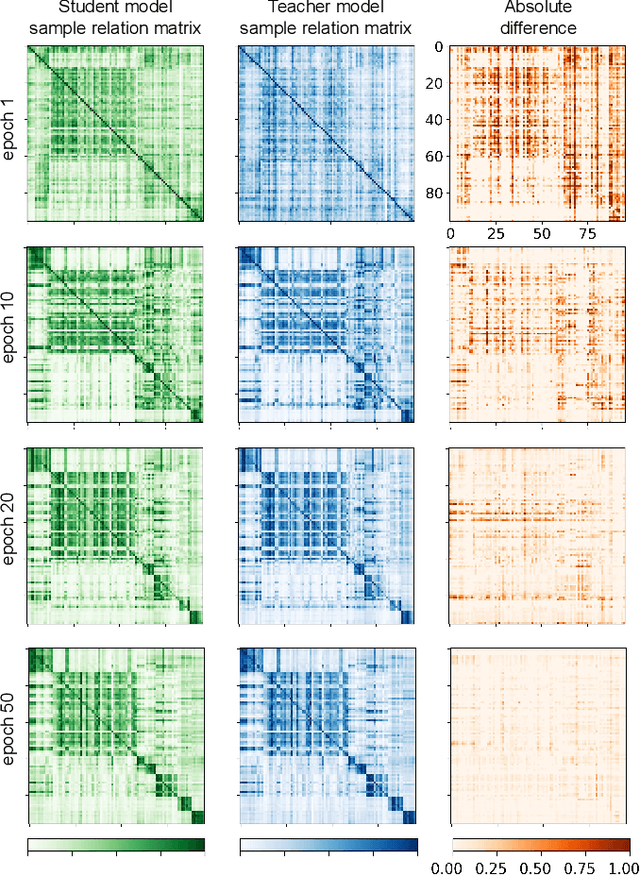
Training deep neural networks usually requires a large amount of labeled data to obtain good performance. However, in medical image analysis, obtaining high-quality labels for the data is laborious and expensive, as accurately annotating medical images demands expertise knowledge of the clinicians. In this paper, we present a novel relation-driven semi-supervised framework for medical image classification. It is a consistency-based method which exploits the unlabeled data by encouraging the prediction consistency of given input under perturbations, and leverages a self-ensembling model to produce high-quality consistency targets for the unlabeled data. Considering that human diagnosis often refers to previous analogous cases to make reliable decisions, we introduce a novel sample relation consistency (SRC) paradigm to effectively exploit unlabeled data by modeling the relationship information among different samples. Superior to existing consistency-based methods which simply enforce consistency of individual predictions, our framework explicitly enforces the consistency of semantic relation among different samples under perturbations, encouraging the model to explore extra semantic information from unlabeled data. We have conducted extensive experiments to evaluate our method on two public benchmark medical image classification datasets, i.e.,skin lesion diagnosis with ISIC 2018 challenge and thorax disease classification with ChestX-ray14. Our method outperforms many state-of-the-art semi-supervised learning methods on both single-label and multi-label image classification scenarios.
"Double-DIP": Unsupervised Image Decomposition via Coupled Deep-Image-Priors
Dec 05, 2018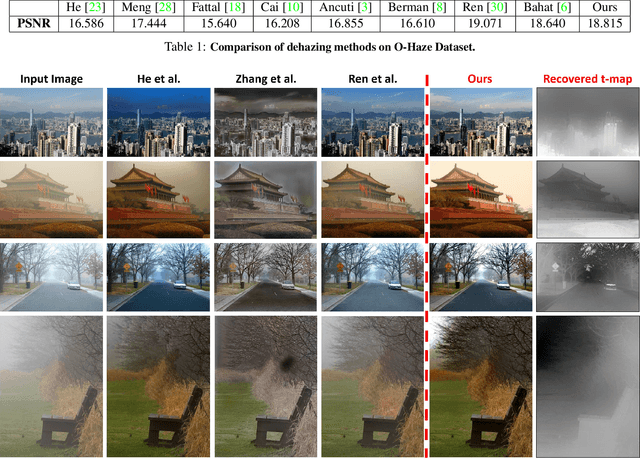
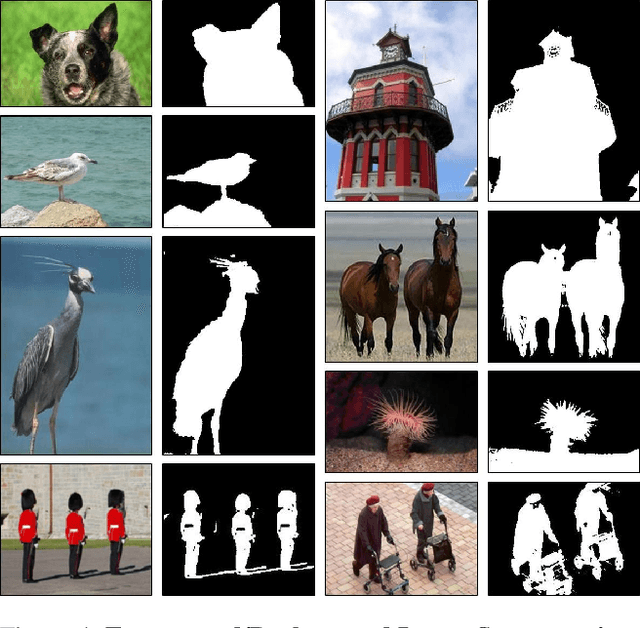
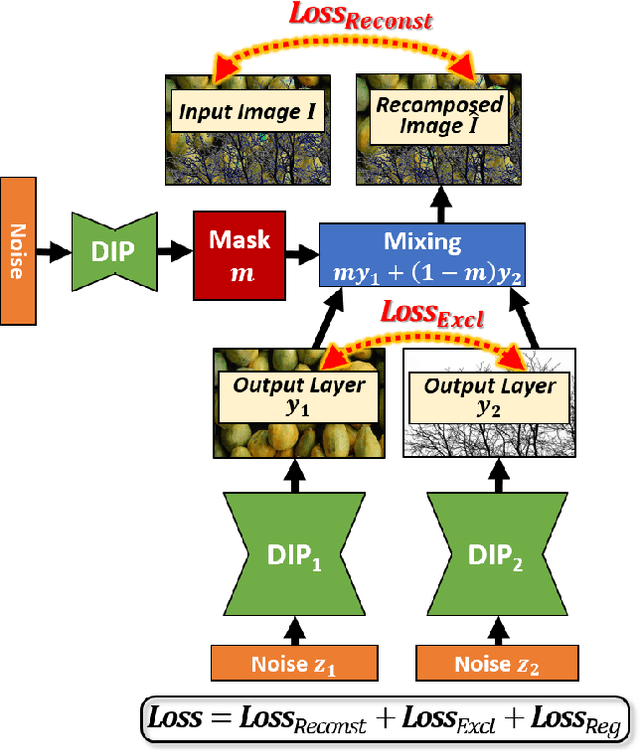
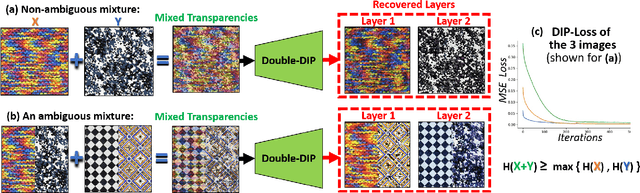

Many seemingly unrelated computer vision tasks can be viewed as a special case of image decomposition into separate layers. For example, image segmentation (separation into foreground and background layers); transparent layer separation (into reflection and transmission layers); Image dehazing (separation into a clear image and a haze map), and more. In this paper we propose a unified framework for unsupervised layer decomposition of a single image, based on coupled "Deep-image-Prior" (DIP) networks. It was shown [Ulyanov et al] that the structure of a single DIP generator network is sufficient to capture the low-level statistics of a single image. We show that coupling multiple such DIPs provides a powerful tool for decomposing images into their basic components, for a wide variety of applications. This capability stems from the fact that the internal statistics of a mixture of layers is more complex than the statistics of each of its individual components. We show the power of this approach for Image-Dehazing, Fg/Bg Segmentation, Watermark-Removal, Transparency Separation in images and video, and more. These capabilities are achieved in a totally unsupervised way, with no training examples other than the input image/video itself.
Learning Hierarchical Graph Neural Networks for Image Clustering
Jul 03, 2021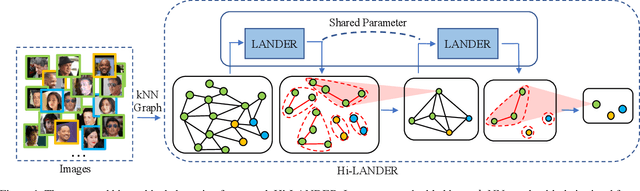
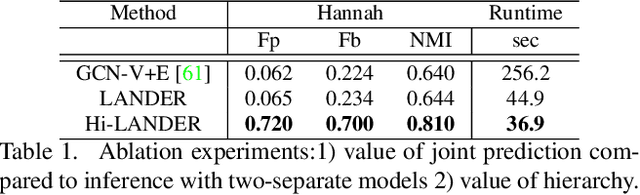
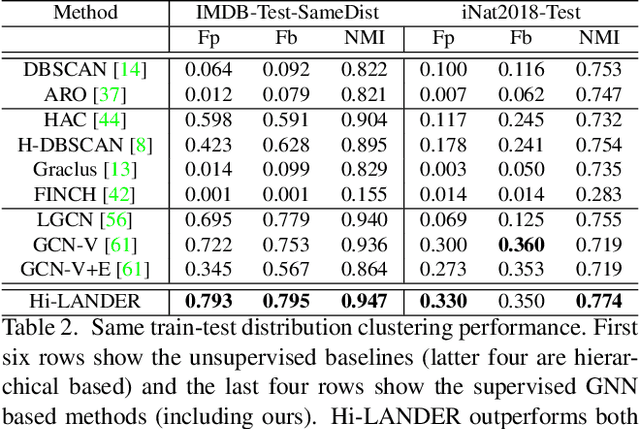
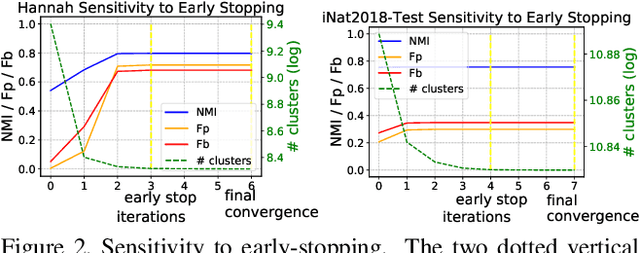
We propose a hierarchical graph neural network (GNN) model that learns how to cluster a set of images into an unknown number of identities using a training set of images annotated with labels belonging to a disjoint set of identities. Our hierarchical GNN uses a novel approach to merge connected components predicted at each level of the hierarchy to form a new graph at the next level. Unlike fully unsupervised hierarchical clustering, the choice of grouping and complexity criteria stems naturally from supervision in the training set. The resulting method, Hi-LANDER, achieves an average of 54% improvement in F-score and 8% increase in Normalized Mutual Information (NMI) relative to current GNN-based clustering algorithms. Additionally, state-of-the-art GNN-based methods rely on separate models to predict linkage probabilities and node densities as intermediate steps of the clustering process. In contrast, our unified framework achieves a seven-fold decrease in computational cost. We release our training and inference code at https://github.com/dmlc/dgl/tree/master/examples/pytorch/hilander.
ALBRT: Cellular Composition Prediction in Routine Histology Images
Aug 26, 2021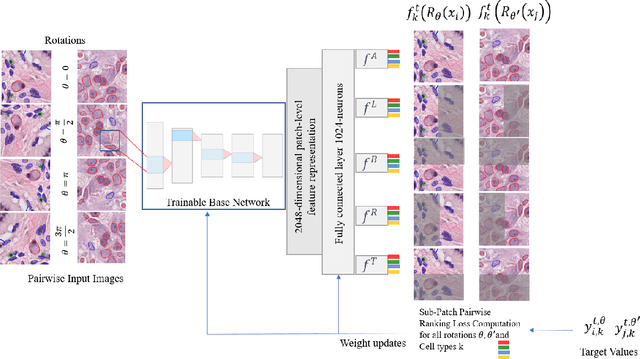
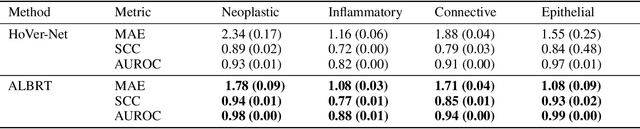
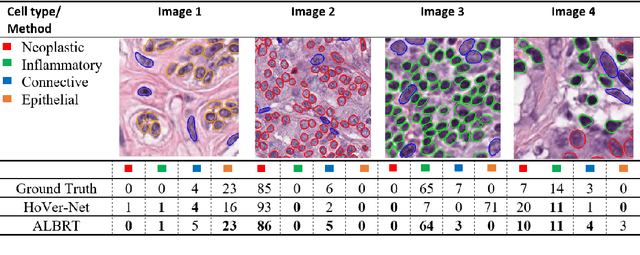
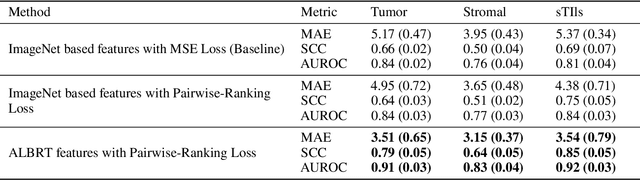
Cellular composition prediction, i.e., predicting the presence and counts of different types of cells in the tumor microenvironment from a digitized image of a Hematoxylin and Eosin (H&E) stained tissue section can be used for various tasks in computational pathology such as the analysis of cellular topology and interactions, subtype prediction, survival analysis, etc. In this work, we propose an image-based cellular composition predictor (ALBRT) which can accurately predict the presence and counts of different types of cells in a given image patch. ALBRT, by its contrastive-learning inspired design, learns a compact and rotation-invariant feature representation that is then used for cellular composition prediction of different cell types. It offers significant improvement over existing state-of-the-art approaches for cell classification and counting. The patch-level feature representation learned by ALBRT is transferrable for cellular composition analysis over novel datasets and can also be utilized for downstream prediction tasks in CPath as well. The code and the inference webserver for the proposed method are available at the URL: https://github.com/engrodawood/ALBRT.
Approximating Human Judgment of Generated Image Quality
Nov 30, 2019

Generative models have made immense progress in recent years, particularly in their ability to generate high quality images. However, that quality has been difficult to evaluate rigorously, with evaluation dominated by heuristic approaches that do not correlate well with human judgment, such as the Inception Score and Fr\'echet Inception Distance. Real human labels have also been used in evaluation, but are inefficient and expensive to collect for each image. Here, we present a novel method to automatically evaluate images based on their quality as perceived by humans. By not only generating image embeddings from Inception network activations and comparing them to the activations for real images, of which other methods perform a variant, but also regressing the activation statistics to match gold standard human labels, we demonstrate 66% accuracy in predicting human scores of image realism, matching the human inter-rater agreement rate. Our approach also generalizes across generative models, suggesting the potential for capturing a model-agnostic measure of image quality. We open source our dataset of human labels for the advancement of research and techniques in this area.
FDA-GAN: Flow-based Dual Attention GAN for Human Pose Transfer
Dec 01, 2021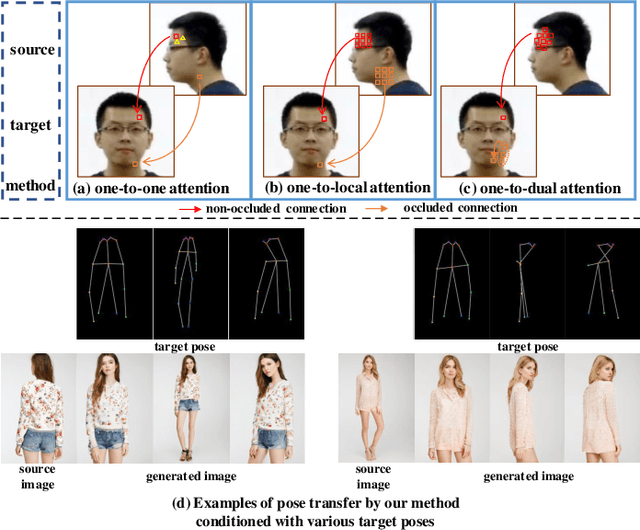
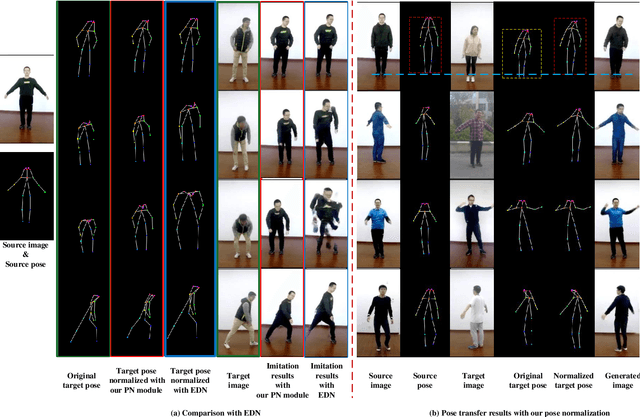


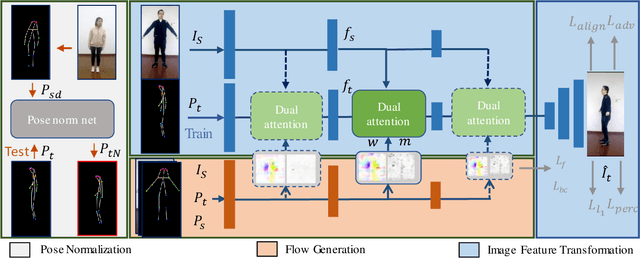
Human pose transfer aims at transferring the appearance of the source person to the target pose. Existing methods utilizing flow-based warping for non-rigid human image generation have achieved great success. However, they fail to preserve the appearance details in synthesized images since the spatial correlation between the source and target is not fully exploited. To this end, we propose the Flow-based Dual Attention GAN (FDA-GAN) to apply occlusion- and deformation-aware feature fusion for higher generation quality. Specifically, deformable local attention and flow similarity attention, constituting the dual attention mechanism, can derive the output features responsible for deformable- and occlusion-aware fusion, respectively. Besides, to maintain the pose and global position consistency in transferring, we design a pose normalization network for learning adaptive normalization from the target pose to the source person. Both qualitative and quantitative results show that our method outperforms state-of-the-art models in public iPER and DeepFashion datasets.
A Robust Completed Local Binary Pattern (RCLBP) for Surface Defect Detection
Dec 07, 2021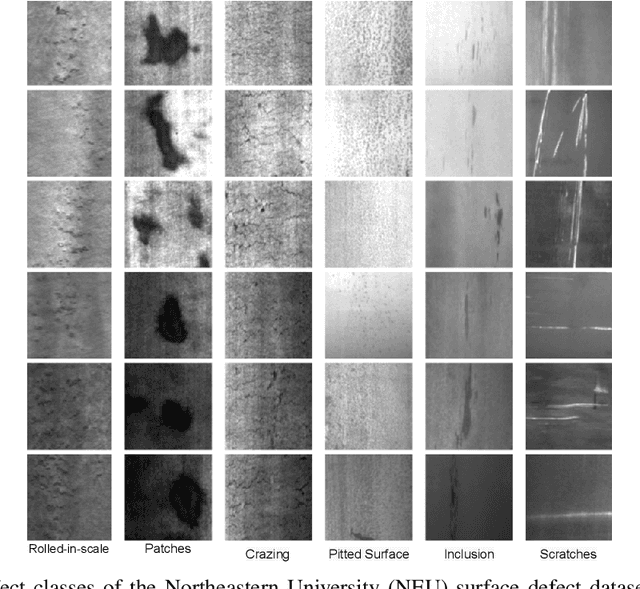
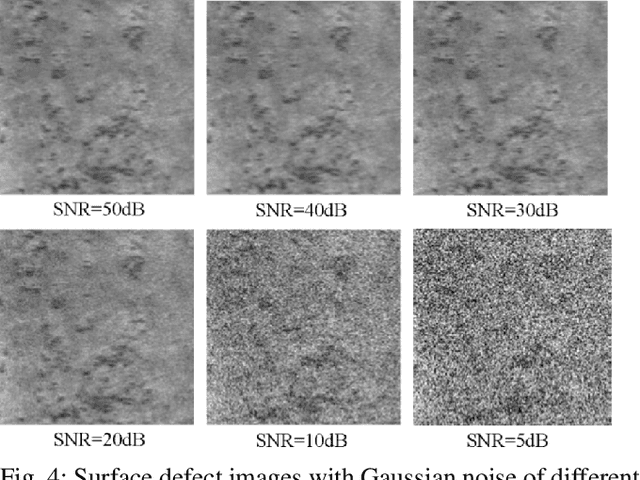
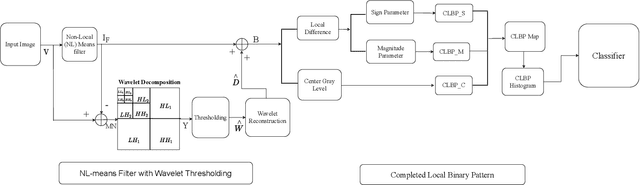
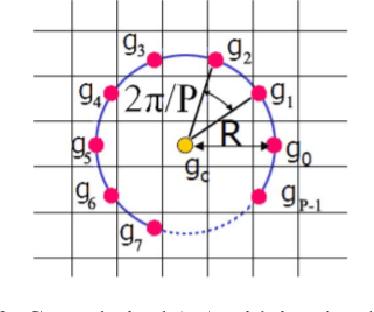

In this paper, we present a Robust Completed Local Binary Pattern (RCLBP) framework for a surface defect detection task. Our approach uses a combination of Non-Local (NL) means filter with wavelet thresholding and Completed Local Binary Pattern (CLBP) to extract robust features which are fed into classifiers for surface defects detection. This paper combines three components: A denoising technique based on Non-Local (NL) means filter with wavelet thresholding is established to denoise the noisy image while preserving the textures and edges. Second, discriminative features are extracted using the CLBP technique. Finally, the discriminative features are fed into the classifiers to build the detection model and evaluate the performance of the proposed framework. The performance of the defect detection models are evaluated using a real-world steel surface defect database from Northeastern University (NEU). Experimental results demonstrate that the proposed approach RCLBP is noise robust and can be applied for surface defect detection under varying conditions of intra-class and inter-class changes and with illumination changes.
HVTR: Hybrid Volumetric-Textural Rendering for Human Avatars
Dec 19, 2021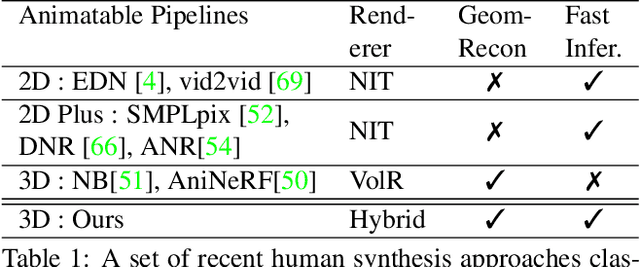
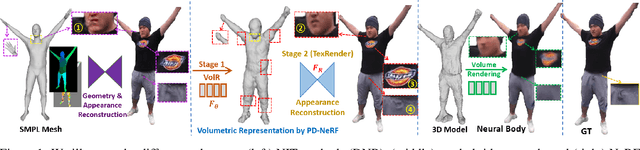
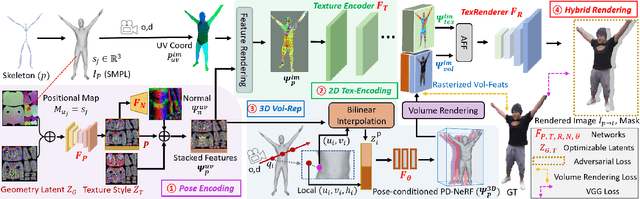
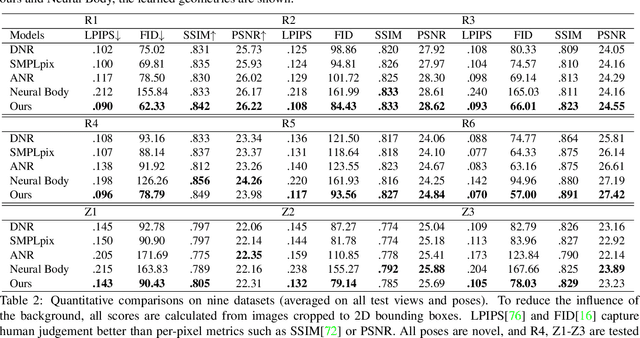
We propose a novel neural rendering pipeline, Hybrid Volumetric-Textural Rendering (HVTR), which synthesizes virtual human avatars from arbitrary poses efficiently and at high quality. First, we learn to encode articulated human motions on a dense UV manifold of the human body surface. To handle complicated motions (e.g., self-occlusions), we then leverage the encoded information on the UV manifold to construct a 3D volumetric representation based on a dynamic pose-conditioned neural radiance field. While this allows us to represent 3D geometry with changing topology, volumetric rendering is computationally heavy. Hence we employ only a rough volumetric representation using a pose-conditioned downsampled neural radiance field (PD-NeRF), which we can render efficiently at low resolutions. In addition, we learn 2D textural features that are fused with rendered volumetric features in image space. The key advantage of our approach is that we can then convert the fused features into a high resolution, high-quality avatar by a fast GAN-based textural renderer. We demonstrate that hybrid rendering enables HVTR to handle complicated motions, render high-quality avatars under user-controlled poses/shapes and even loose clothing, and most importantly, be fast at inference time. Our experimental results also demonstrate state-of-the-art quantitative results.
Automatic Recognition of Abdominal Organs in Ultrasound Images based on Deep Neural Networks and K-Nearest-Neighbor Classification
Oct 09, 2021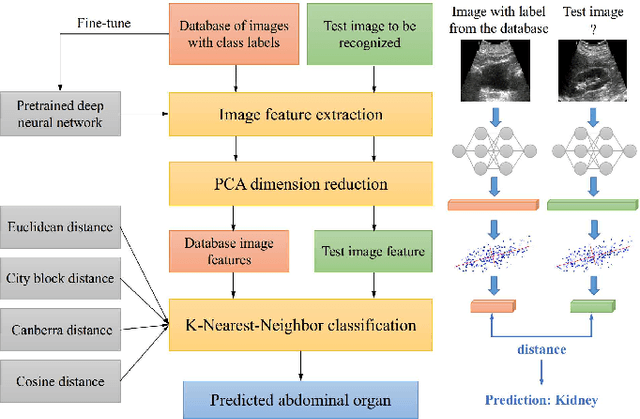
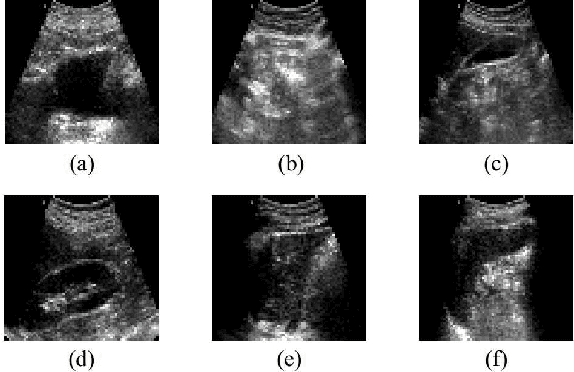
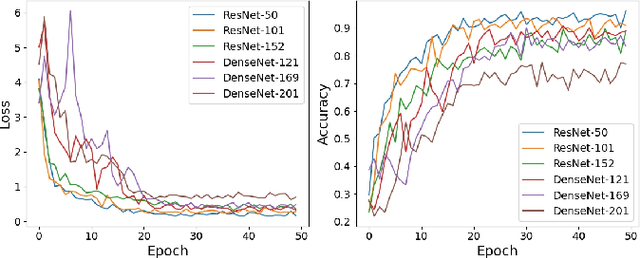
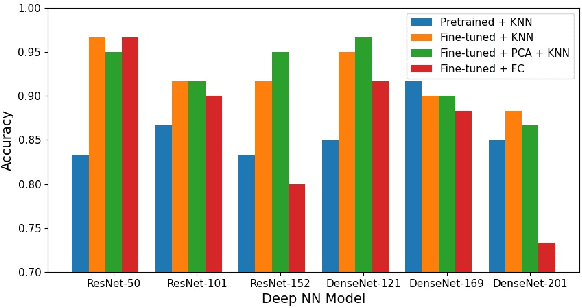
Abdominal ultrasound imaging has been widely used to assist in the diagnosis and treatment of various abdominal organs. In order to shorten the examination time and reduce the cognitive burden on the sonographers, we present a classification method that combines the deep learning techniques and k-Nearest-Neighbor (k-NN) classification to automatically recognize various abdominal organs in the ultrasound images in real time. Fine-tuned deep neural networks are used in combination with PCA dimension reduction to extract high-level features from raw ultrasound images, and a k-NN classifier is employed to predict the abdominal organ in the image. We demonstrate the effectiveness of our method in the task of ultrasound image classification to automatically recognize six abdominal organs. A comprehensive comparison of different configurations is conducted to study the influence of different feature extractors and classifiers on the classification accuracy. Both quantitative and qualitative results show that with minimal training effort, our method can "lazily" recognize the abdominal organs in the ultrasound images in real time with an accuracy of 96.67%. Our implementation code is publicly available at: https://github.com/LeeKeyu/abdominal_ultrasound_classification.
Deformed2Self: Self-Supervised Denoising for Dynamic Medical Imaging
Jun 23, 2021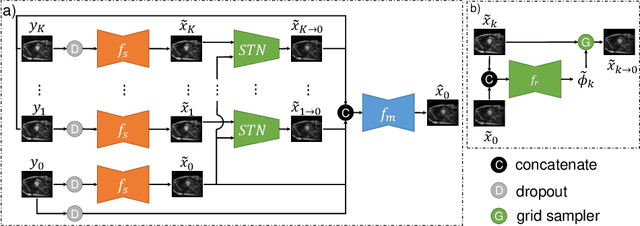
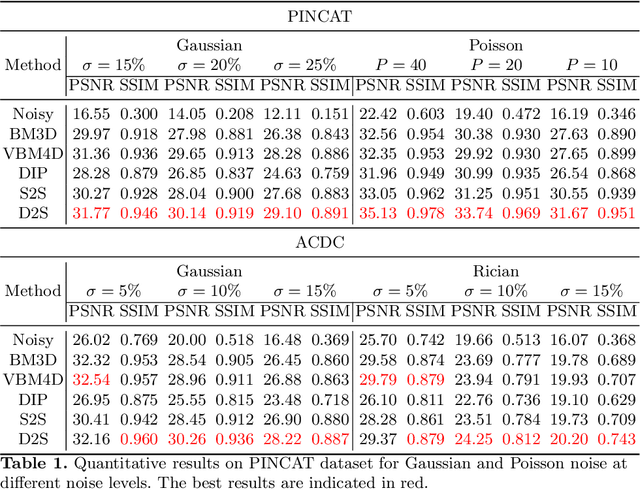
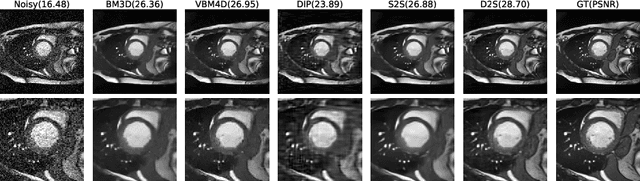
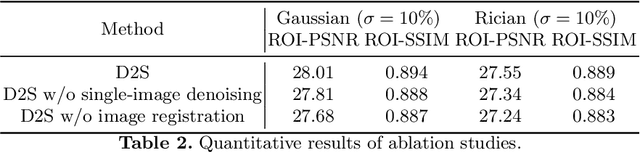
Image denoising is of great importance for medical imaging system, since it can improve image quality for disease diagnosis and downstream image analyses. In a variety of applications, dynamic imaging techniques are utilized to capture the time-varying features of the subject, where multiple images are acquired for the same subject at different time points. Although signal-to-noise ratio of each time frame is usually limited by the short acquisition time, the correlation among different time frames can be exploited to improve denoising results with shared information across time frames. With the success of neural networks in computer vision, supervised deep learning methods show prominent performance in single-image denoising, which rely on large datasets with clean-vs-noisy image pairs. Recently, several self-supervised deep denoising models have been proposed, achieving promising results without needing the pairwise ground truth of clean images. In the field of multi-image denoising, however, very few works have been done on extracting correlated information from multiple slices for denoising using self-supervised deep learning methods. In this work, we propose Deformed2Self, an end-to-end self-supervised deep learning framework for dynamic imaging denoising. It combines single-image and multi-image denoising to improve image quality and use a spatial transformer network to model motion between different slices. Further, it only requires a single noisy image with a few auxiliary observations at different time frames for training and inference. Evaluations on phantom and in vivo data with different noise statistics show that our method has comparable performance to other state-of-the-art unsupervised or self-supervised denoising methods and outperforms under high noise levels.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge