"Image": models, code, and papers
Semi-supervised Learning for COVID-19 Image Classification via ResNet
Feb 27, 2021
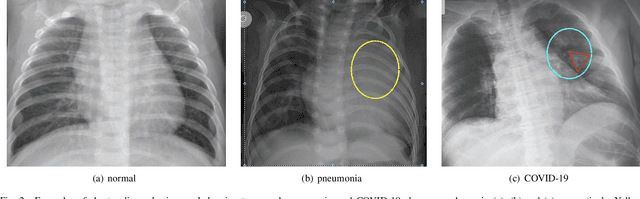
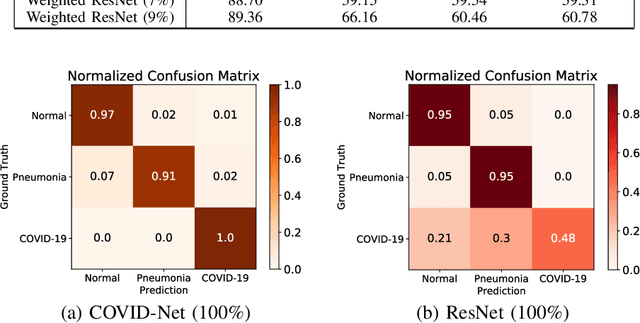
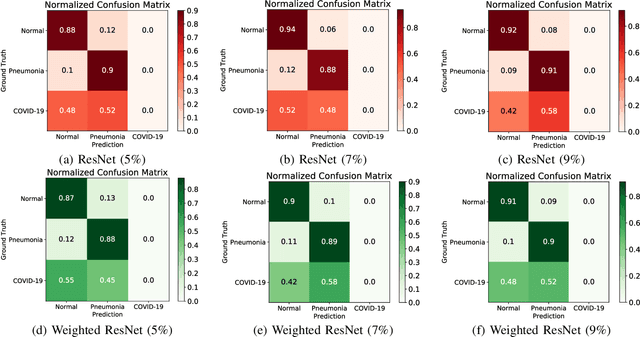
Coronavirus disease 2019 (COVID-19) is an ongoing global pandemic in over 200 countries and territories, which has resulted in a great public health concern across the international community. Analysis of X-ray imaging data can play a critical role in timely and accurate screening and fighting against COVID-19. Supervised deep learning has been successfully applied to recognize COVID-19 pathology from X-ray imaging datasets. However, it requires a substantial amount of annotated X-ray images to train models, which is often not applicable to data analysis for emerging events such as COVID-19 outbreak, especially in the early stage of the outbreak. To address this challenge, this paper proposes a two-path semi-supervised deep learning model, ssResNet, based on Residual Neural Network (ResNet) for COVID-19 image classification, where two paths refer to a supervised path and an unsupervised path, respectively. Moreover, we design a weighted supervised loss that assigns higher weight for the minority classes in the training process to resolve the data imbalance. Experimental results on a large-scale of X-ray image dataset COVIDx demonstrate that the proposed model can achieve promising performance even when trained on very few labeled training images.
YOLO-Pose: Enhancing YOLO for Multi Person Pose Estimation Using Object Keypoint Similarity Loss
Apr 14, 2022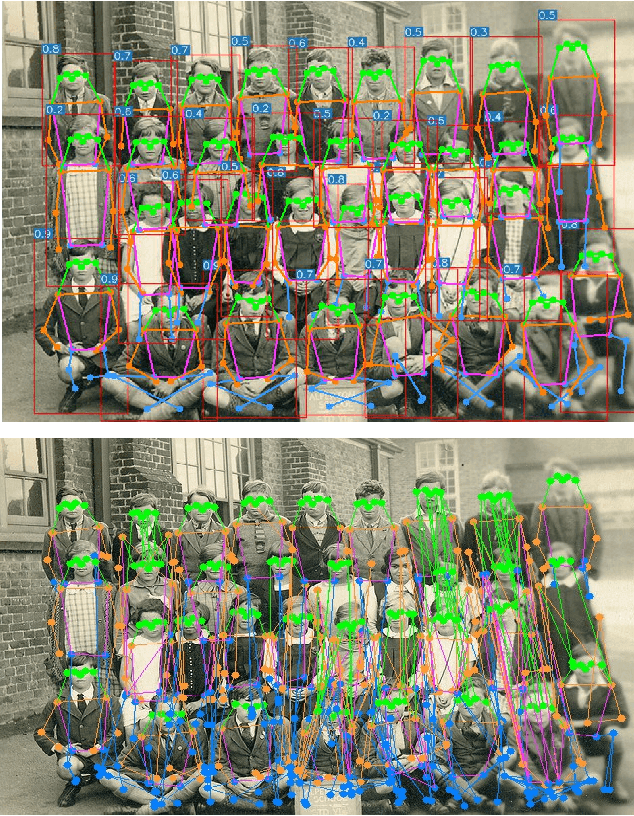
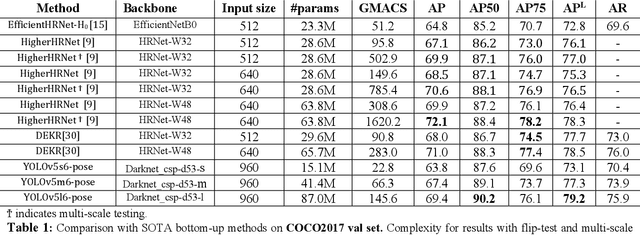
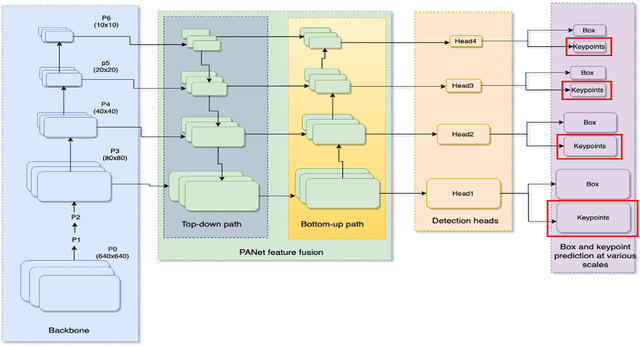
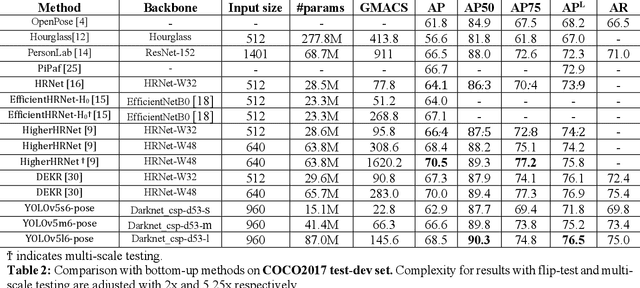
We introduce YOLO-pose, a novel heatmap-free approach for joint detection, and 2D multi-person pose estimation in an image based on the popular YOLO object detection framework. Existing heatmap based two-stage approaches are sub-optimal as they are not end-to-end trainable and training relies on a surrogate L1 loss that is not equivalent to maximizing the evaluation metric, i.e. Object Keypoint Similarity (OKS). Our framework allows us to train the model end-to-end and optimize the OKS metric itself. The proposed model learns to jointly detect bounding boxes for multiple persons and their corresponding 2D poses in a single forward pass and thus bringing in the best of both top-down and bottom-up approaches. Proposed approach doesn't require the postprocessing of bottom-up approaches to group detected keypoints into a skeleton as each bounding box has an associated pose, resulting in an inherent grouping of the keypoints. Unlike top-down approaches, multiple forward passes are done away with since all persons are localized along with their pose in a single inference. YOLO-pose achieves new state-of-the-art results on COCO validation (90.2% AP50) and test-dev set (90.3% AP50), surpassing all existing bottom-up approaches in a single forward pass without flip test, multi-scale testing, or any other test time augmentation. All experiments and results reported in this paper are without any test time augmentation, unlike traditional approaches that use flip-test and multi-scale testing to boost performance. Our training codes will be made publicly available at https://github.com/TexasInstruments/edgeai-yolov5 and https://github.com/TexasInstruments/edgeai-yolox
A Joint Morphological Profiles and Patch Tensor Change Detection for Hyperspectral Imagery
Jan 20, 2022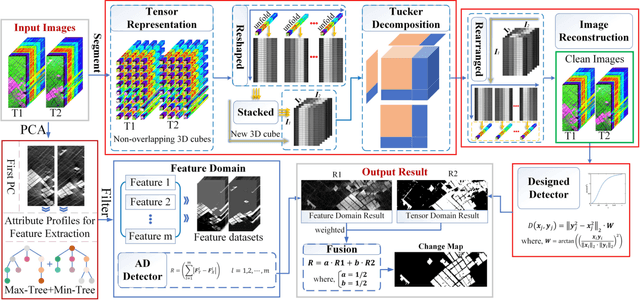
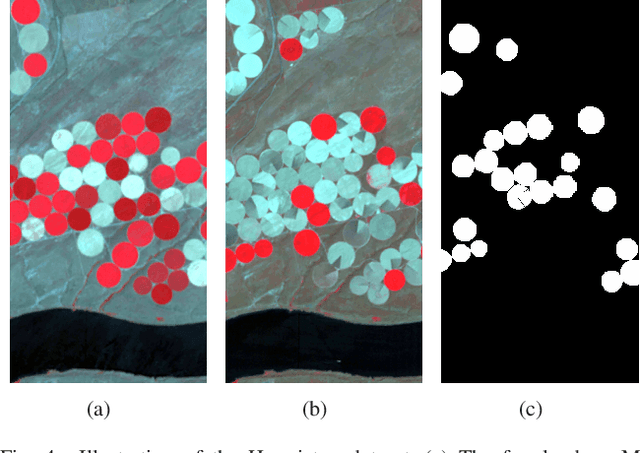
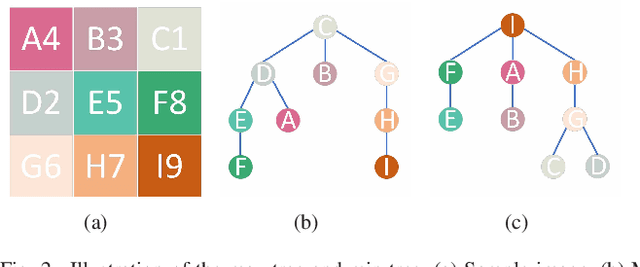
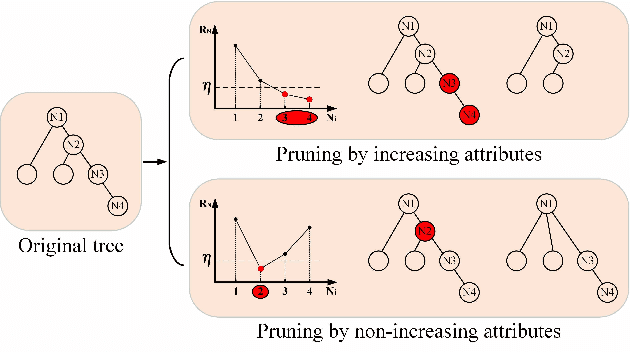

Multi-temporal hyperspectral images can be used to detect changed information, which has gradually attracted researchers' attention. However, traditional change detection algorithms have not deeply explored the relevance of spatial and spectral changed features, which leads to low detection accuracy. To better excavate both spectral and spatial information of changed features, a joint morphology and patch-tensor change detection (JMPT) method is proposed. Initially, a patch-based tensor strategy is adopted to exploit similar property of spatial structure, where the non-overlapping local patch image is reshaped into a new tensor cube, and then three-order Tucker decompositon and image reconstruction strategies are adopted to obtain more robust multi-temporal hyperspectral datasets. Meanwhile, multiple morphological profiles including max-tree and min-tree are applied to extract different attributes of multi-temporal images. Finally, these results are fused to general a final change detection map. Experiments conducted on two real hyperspectral datasets demonstrate that the proposed detector achieves better detection performance.
Image Inpainting Guided by Coherence Priors of Semantics and Textures
Dec 15, 2020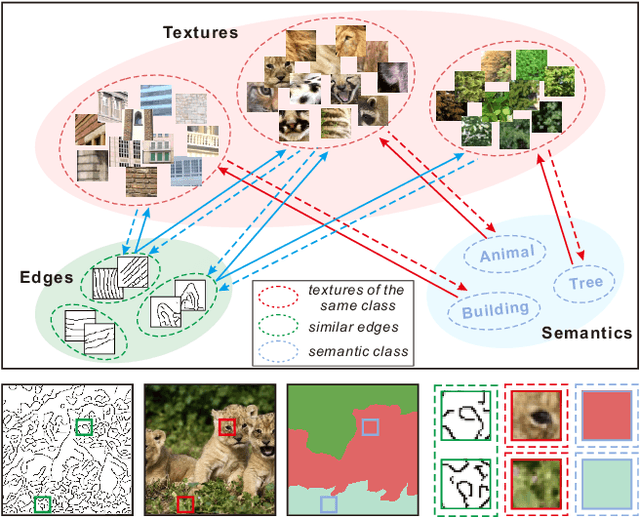
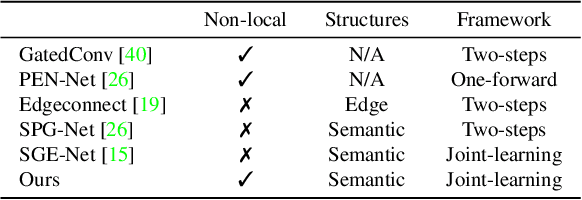
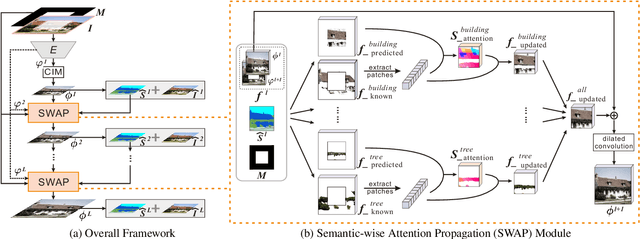
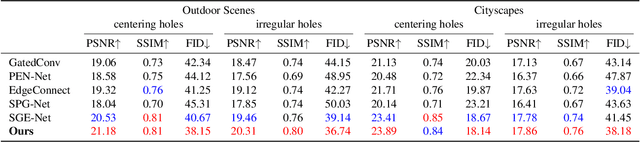
Existing inpainting methods have achieved promising performance in recovering defected images of specific scenes. However, filling holes involving multiple semantic categories remains challenging due to the obscure semantic boundaries and the mixture of different semantic textures. In this paper, we introduce coherence priors between the semantics and textures which make it possible to concentrate on completing separate textures in a semantic-wise manner. Specifically, we adopt a multi-scale joint optimization framework to first model the coherence priors and then accordingly interleavingly optimize image inpainting and semantic segmentation in a coarse-to-fine manner. A Semantic-Wise Attention Propagation (SWAP) module is devised to refine completed image textures across scales by exploring non-local semantic coherence, which effectively mitigates mix-up of textures. We also propose two coherence losses to constrain the consistency between the semantics and the inpainted image in terms of the overall structure and detailed textures. Experimental results demonstrate the superiority of our proposed method for challenging cases with complex holes.
Multi-institutional Collaborations for Improving Deep Learning-based Magnetic Resonance Image Reconstruction Using Federated Learning
Mar 03, 2021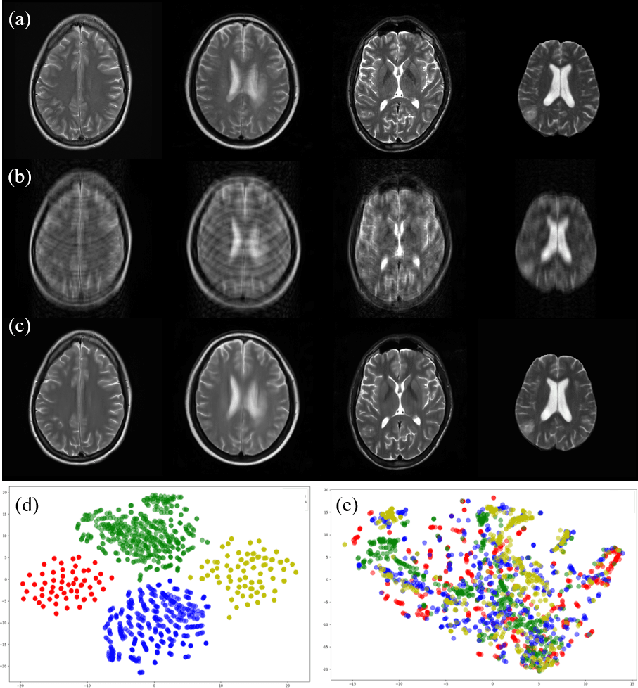
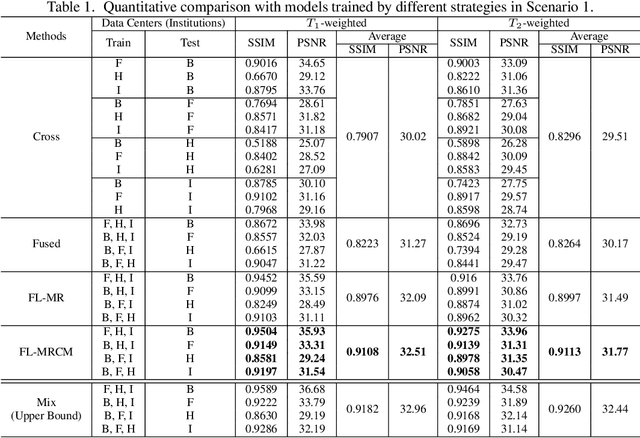
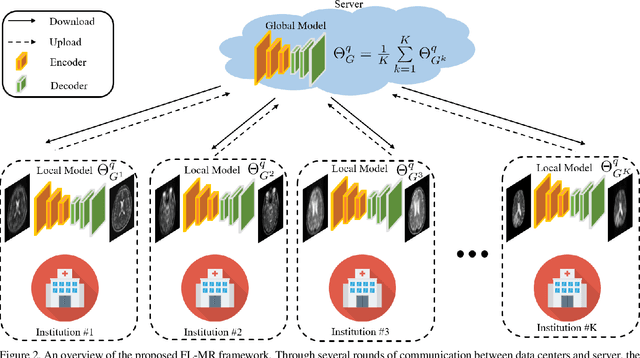
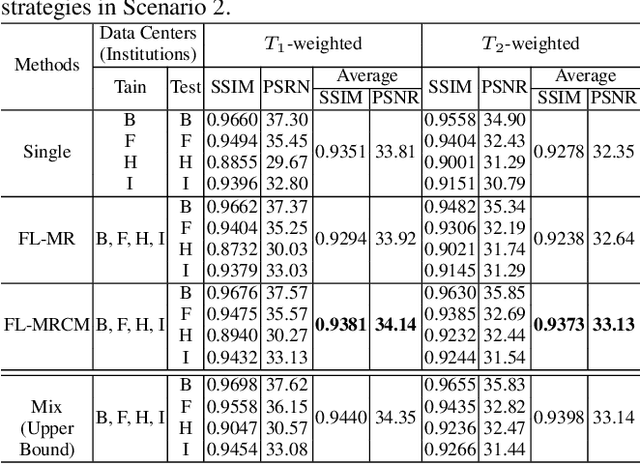
Fast and accurate reconstruction of magnetic resonance (MR) images from under-sampled data is important in many clinical applications. In recent years, deep learning-based methods have been shown to produce superior performance on MR image reconstruction. However, these methods require large amounts of data which is difficult to collect and share due to the high cost of acquisition and medical data privacy regulations. In order to overcome this challenge, we propose a federated learning (FL) based solution in which we take advantage of the MR data available at different institutions while preserving patients' privacy. However, the generalizability of models trained with the FL setting can still be suboptimal due to domain shift, which results from the data collected at multiple institutions with different sensors, disease types, and acquisition protocols, etc. With the motivation of circumventing this challenge, we propose a cross-site modeling for MR image reconstruction in which the learned intermediate latent features among different source sites are aligned with the distribution of the latent features at the target site. Extensive experiments are conducted to provide various insights about FL for MR image reconstruction. Experimental results demonstrate that the proposed framework is a promising direction to utilize multi-institutional data without compromising patients' privacy for achieving improved MR image reconstruction. Our code will be available at https://github.com/guopengf/FLMRCM.
Automatic image annotation base on Naive Bayes and Decision Tree classifiers using MPEG-7
Jan 27, 2021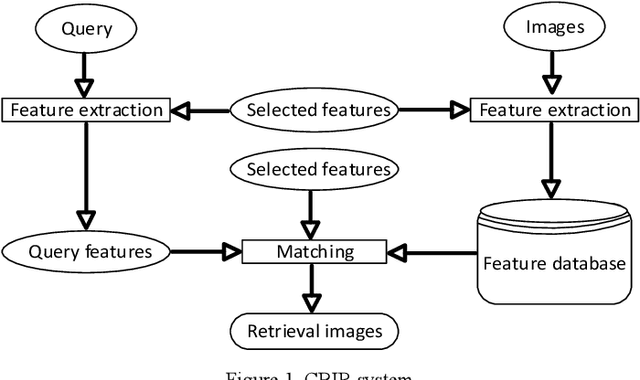
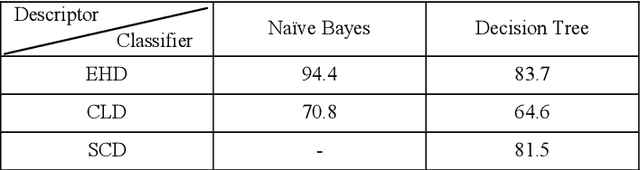
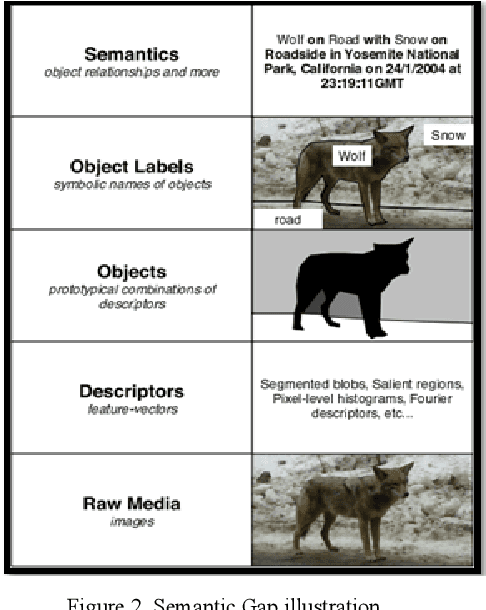


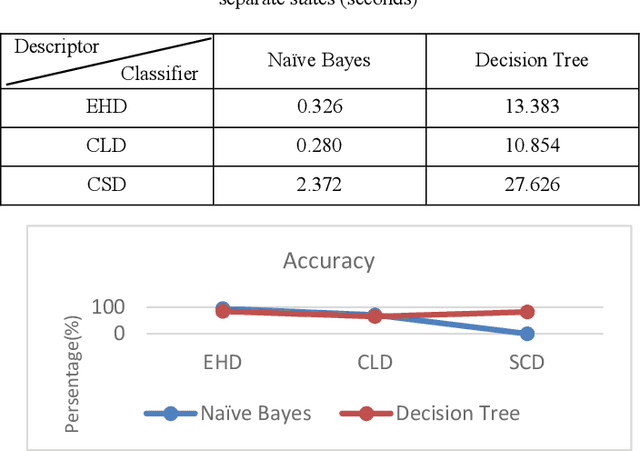
Recently it has become essential to search for and retrieve high-resolution and efficient images easily due to swift development of digital images, many present annotation algorithms facing a big challenge which is the variance for represent the image where high level represent image semantic and low level illustrate the features, this issue is known as semantic gab. This work has been used MPEG-7 standard to extract the features from the images, where the color feature was extracted by using Scalable Color Descriptor (SCD) and Color Layout Descriptor (CLD), whereas the texture feature was extracted by employing Edge Histogram Descriptor (EHD), the CLD produced high dimensionality feature vector therefore it is reduced by Principal Component Analysis (PCA). The features that have extracted by these three descriptors could be passing to the classifiers (Naive Bayes and Decision Tree) for training. Finally, they annotated the query image. In this study TUDarmstadt image bank had been used. The results of tests and comparative performance evaluation indicated better precision and executing time of Naive Bayes classification in comparison with Decision Tree classification.
Sample Efficient Grasp Learning Using Equivariant Models
Feb 18, 2022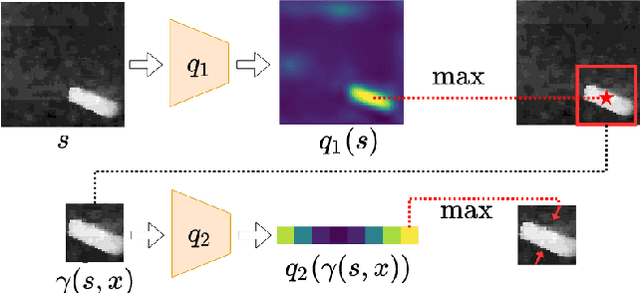
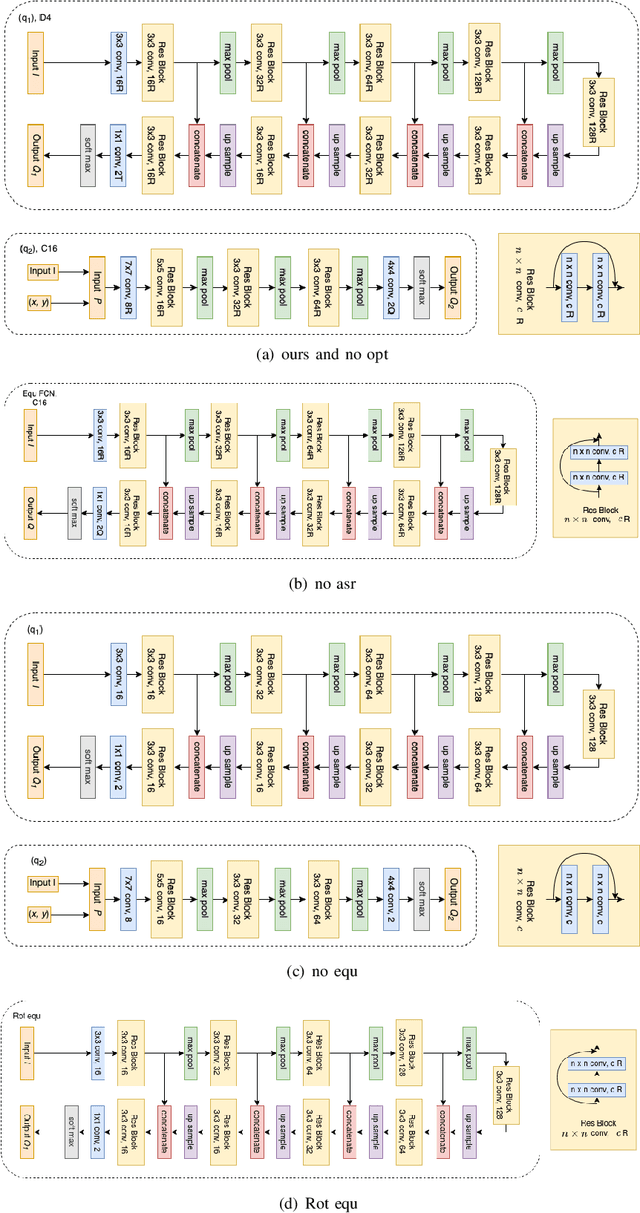
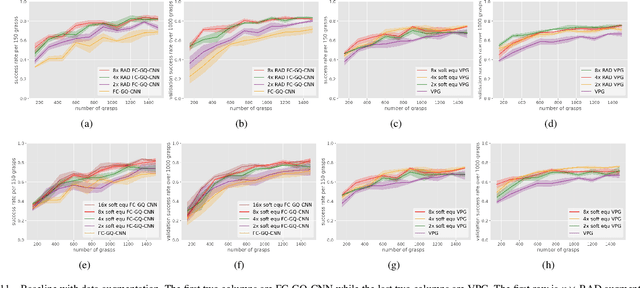
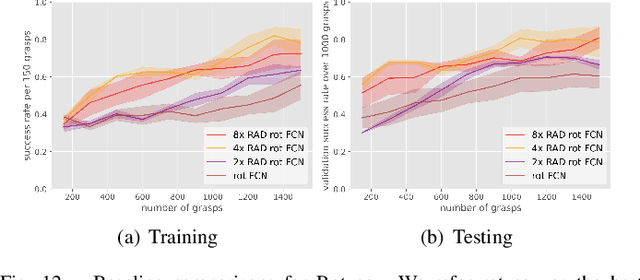
In planar grasp detection, the goal is to learn a function from an image of a scene onto a set of feasible grasp poses in $\mathrm{SE}(2)$. In this paper, we recognize that the optimal grasp function is $\mathrm{SE}(2)$-equivariant and can be modeled using an equivariant convolutional neural network. As a result, we are able to significantly improve the sample efficiency of grasp learning, obtaining a good approximation of the grasp function after only 600 grasp attempts. This is few enough that we can learn to grasp completely on a physical robot in about 1.5 hours.
MedSelect: Selective Labeling for Medical Image Classification Combining Meta-Learning with Deep Reinforcement Learning
Mar 26, 2021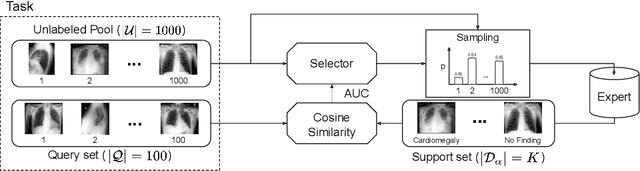
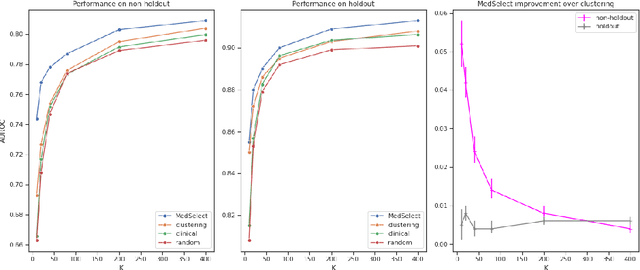
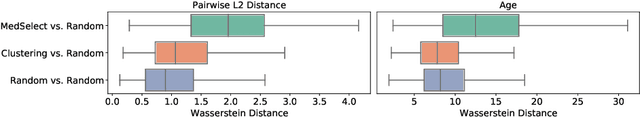
We propose a selective learning method using meta-learning and deep reinforcement learning for medical image interpretation in the setting of limited labeling resources. Our method, MedSelect, consists of a trainable deep learning selector that uses image embeddings obtained from contrastive pretraining for determining which images to label, and a non-parametric selector that uses cosine similarity to classify unseen images. We demonstrate that MedSelect learns an effective selection strategy outperforming baseline selection strategies across seen and unseen medical conditions for chest X-ray interpretation. We also perform an analysis of the selections performed by MedSelect comparing the distribution of latent embeddings and clinical features, and find significant differences compared to the strongest performing baseline. We believe that our method may be broadly applicable across medical imaging settings where labels are expensive to acquire.
AtomAI: A Deep Learning Framework for Analysis of Image and Spectroscopy Data in (Scanning) Transmission Electron Microscopy and Beyond
May 16, 2021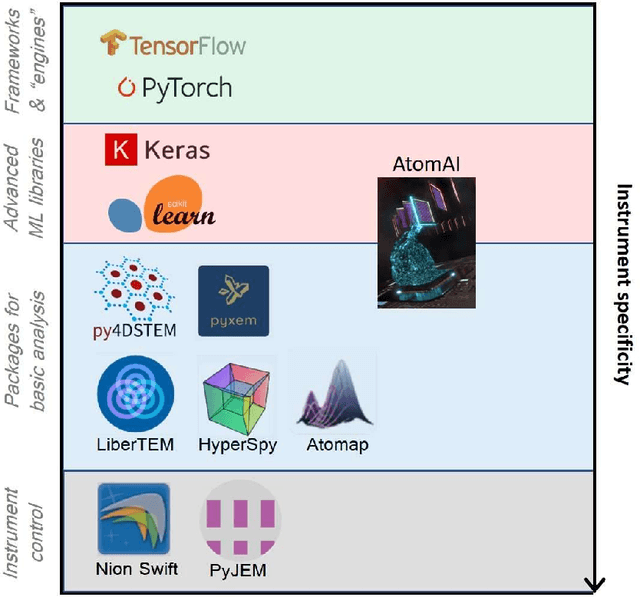
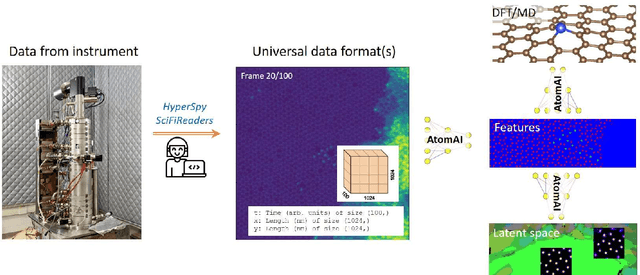
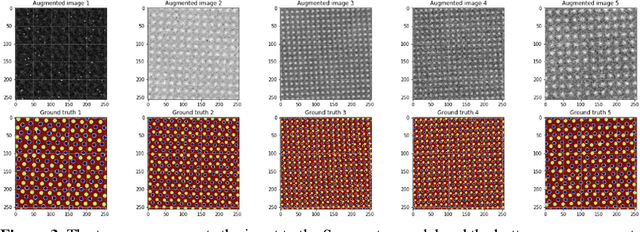
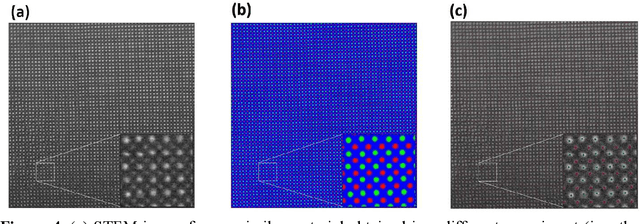
AtomAI is an open-source software package bridging instrument-specific Python libraries, deep learning, and simulation tools into a single ecosystem. AtomAI allows direct applications of the deep convolutional neural networks for atomic and mesoscopic image segmentation converting image and spectroscopy data into class-based local descriptors for downstream tasks such as statistical and graph analysis. For atomically-resolved imaging data, the output is types and positions of atomic species, with an option for subsequent refinement. AtomAI further allows the implementation of a broad range of image and spectrum analysis functions, including invariant variational autoencoders (VAEs). The latter consists of VAEs with rotational and (optionally) translational invariance for unsupervised and class-conditioned disentanglement of categorical and continuous data representations. In addition, AtomAI provides utilities for mapping structure-property relationships via im2spec and spec2im type of encoder-decoder models. Finally, AtomAI allows seamless connection to the first principles modeling with a Python interface, including molecular dynamics and density functional theory calculations on the inferred atomic position. While the majority of applications to date were based on atomically resolved electron microscopy, the flexibility of AtomAI allows straightforward extension towards the analysis of mesoscopic imaging data once the labels and feature identification workflows are established/available. The source code and example notebooks are available at https://github.com/pycroscopy/atomai.
Delving into Inter-Image Invariance for Unsupervised Visual Representations
Aug 26, 2020
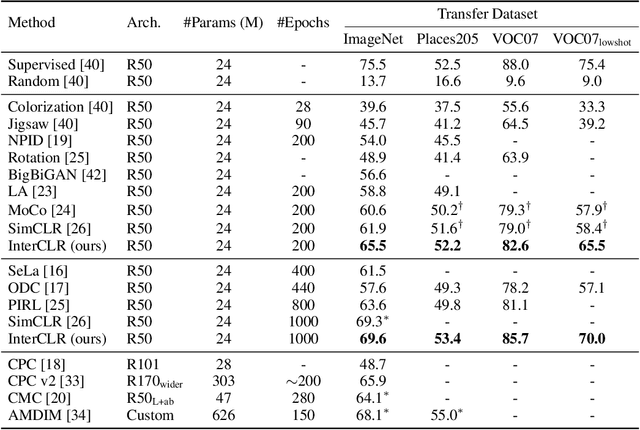
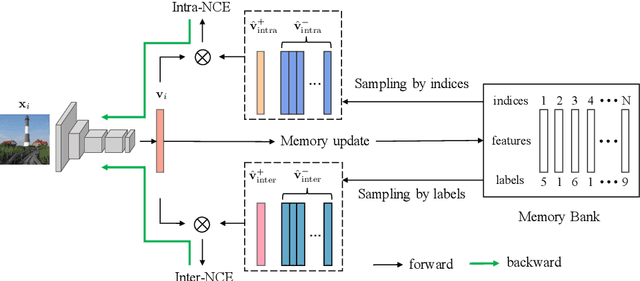
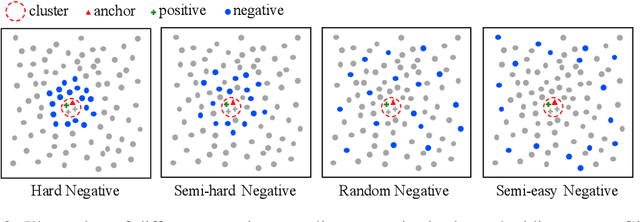
Contrastive learning has recently shown immense potential in unsupervised visual representation learning. Existing studies in this track mainly focus on intra-image invariance learning. The learning typically uses rich intra-image transformations to construct positive pairs and then maximizes agreement using a contrastive loss. The merits of inter-image invariance, conversely, remain much less explored. One major obstacle to exploit inter-image invariance is that it is unclear how to reliably construct inter-image positive pairs, and further derive effective supervision from them since there are no pair annotations available. In this work, we present a rigorous and comprehensive study on inter-image invariance learning from three main constituting components: pseudo-label maintenance, sampling strategy, and decision boundary design. Through carefully-designed comparisons and analysis, we propose a unified framework that supports the integration of unsupervised intra- and inter-image invariance learning. With all the obtained recipes, our final model, namely InterCLR, achieves state-of-the-art performance on standard benchmarks. Code and models will be available at https://github.com/open-mmlab/OpenSelfSup.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge