Ayana Ghosh
Active Causal Learning for Decoding Chemical Complexities with Targeted Interventions
Apr 05, 2024Abstract:Predicting and enhancing inherent properties based on molecular structures is paramount to design tasks in medicine, materials science, and environmental management. Most of the current machine learning and deep learning approaches have become standard for predictions, but they face challenges when applied across different datasets due to reliance on correlations between molecular representation and target properties. These approaches typically depend on large datasets to capture the diversity within the chemical space, facilitating a more accurate approximation, interpolation, or extrapolation of the chemical behavior of molecules. In our research, we introduce an active learning approach that discerns underlying cause-effect relationships through strategic sampling with the use of a graph loss function. This method identifies the smallest subset of the dataset capable of encoding the most information representative of a much larger chemical space. The identified causal relations are then leveraged to conduct systematic interventions, optimizing the design task within a chemical space that the models have not encountered previously. While our implementation focused on the QM9 quantum-chemical dataset for a specific design task-finding molecules with a large dipole moment-our active causal learning approach, driven by intelligent sampling and interventions, holds potential for broader applications in molecular, materials design and discovery.
Active Deep Kernel Learning of Molecular Functionalities: Realizing Dynamic Structural Embeddings
Mar 02, 2024Abstract:Exploring molecular spaces is crucial for advancing our understanding of chemical properties and reactions, leading to groundbreaking innovations in materials science, medicine, and energy. This paper explores an approach for active learning in molecular discovery using Deep Kernel Learning (DKL), a novel approach surpassing the limits of classical Variational Autoencoders (VAEs). Employing the QM9 dataset, we contrast DKL with traditional VAEs, which analyze molecular structures based on similarity, revealing limitations due to sparse regularities in latent spaces. DKL, however, offers a more holistic perspective by correlating structure with properties, creating latent spaces that prioritize molecular functionality. This is achieved by recalculating embedding vectors iteratively, aligning with the experimental availability of target properties. The resulting latent spaces are not only better organized but also exhibit unique characteristics such as concentrated maxima representing molecular functionalities and a correlation between predictive uncertainty and error. Additionally, the formation of exclusion regions around certain compounds indicates unexplored areas with potential for groundbreaking functionalities. This study underscores DKL's potential in molecular research, offering new avenues for understanding and discovering molecular functionalities beyond classical VAE limitations.
Deep Learning for Automated Experimentation in Scanning Transmission Electron Microscopy
Apr 04, 2023



Abstract:Machine learning (ML) has become critical for post-acquisition data analysis in (scanning) transmission electron microscopy, (S)TEM, imaging and spectroscopy. An emerging trend is the transition to real-time analysis and closed-loop microscope operation. The effective use of ML in electron microscopy now requires the development of strategies for microscopy-centered experiment workflow design and optimization. Here, we discuss the associated challenges with the transition to active ML, including sequential data analysis and out-of-distribution drift effects, the requirements for the edge operation, local and cloud data storage, and theory in the loop operations. Specifically, we discuss the relative contributions of human scientists and ML agents in the ideation, orchestration, and execution of experimental workflows and the need to develop universal hyper languages that can apply across multiple platforms. These considerations will collectively inform the operationalization of ML in next-generation experimentation.
Discovery of structure-property relations for molecules via hypothesis-driven active learning over the chemical space
Jan 06, 2023



Abstract:Discovery of the molecular candidates for applications in drug targets, biomolecular systems, catalysts, photovoltaics, organic electronics, and batteries, necessitates development of machine learning algorithms capable of rapid exploration of the chemical spaces targeting the desired functionalities. Here we introduce a novel approach for the active learning over the chemical spaces based on hypothesis learning. We construct the hypotheses on the possible relationships between structures and functionalities of interest based on a small subset of data and introduce them as (probabilistic) mean functions for the Gaussian process. This approach combines the elements from the symbolic regression methods such as SISSO and active learning into a single framework. Here, we demonstrate it for the QM9 dataset, but it can be applied more broadly to datasets from both domains of molecular and solid-state materials sciences.
Microscopy is All You Need
Oct 12, 2022
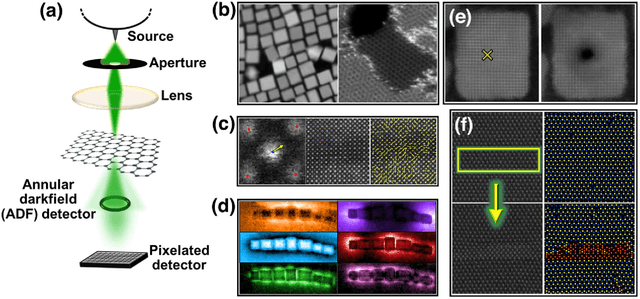

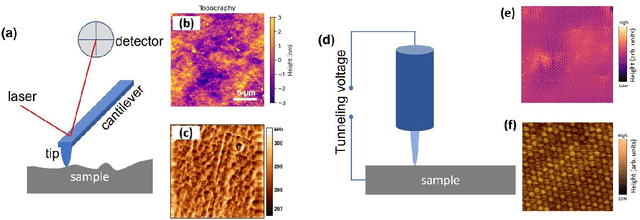
Abstract:We pose that microscopy offers an ideal real-world experimental environment for the development and deployment of active Bayesian and reinforcement learning methods. Indeed, the tremendous progress achieved by machine learning (ML) and artificial intelligence over the last decade has been largely achieved via the utilization of static data sets, from the paradigmatic MNIST to the bespoke corpora of text and image data used to train large models such as GPT3, DALLE and others. However, it is now recognized that continuous, minute improvements to state-of-the-art do not necessarily translate to advances in real-world applications. We argue that a promising pathway for the development of ML methods is via the route of domain-specific deployable algorithms in areas such as electron and scanning probe microscopy and chemical imaging. This will benefit both fundamental physical studies and serve as a test bed for more complex autonomous systems such as robotics and manufacturing. Favorable environment characteristics of scanning and electron microscopy include low risk, extensive availability of domain-specific priors and rewards, relatively small effects of exogeneous variables, and often the presence of both upstream first principles as well as downstream learnable physical models for both statics and dynamics. Recent developments in programmable interfaces, edge computing, and access to APIs facilitating microscope control, all render the deployment of ML codes on operational microscopes straightforward. We discuss these considerations and hope that these arguments will lead to creating a novel set of development targets for the ML community by accelerating both real-world ML applications and scientific progress.
AtomAI: A Deep Learning Framework for Analysis of Image and Spectroscopy Data in Transmission Electron Microscopy and Beyond
May 16, 2021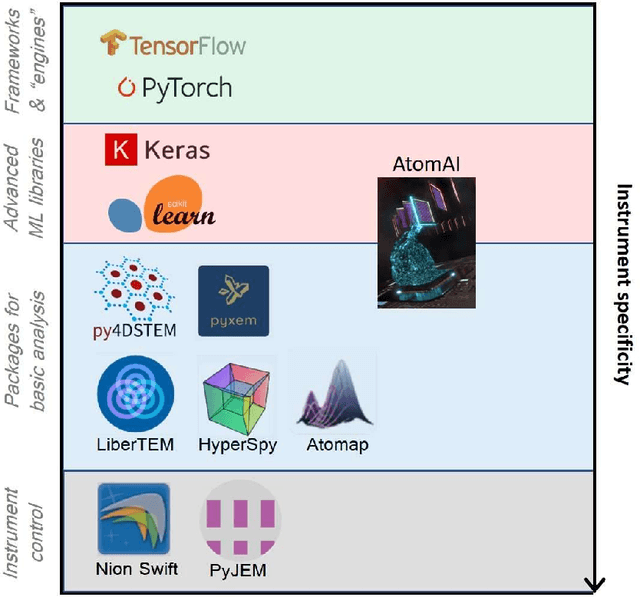
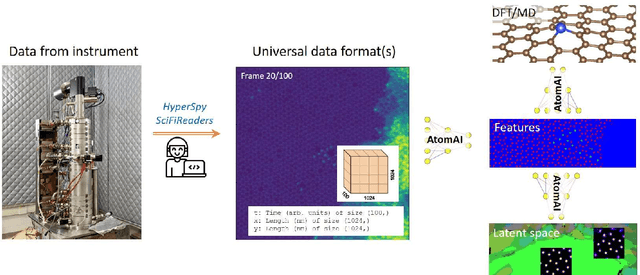
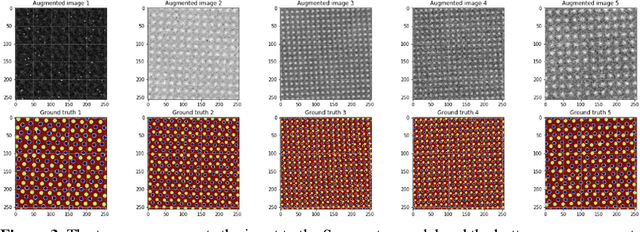
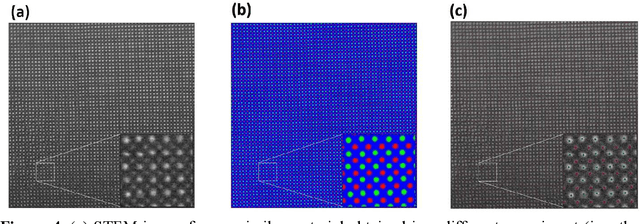
Abstract:AtomAI is an open-source software package bridging instrument-specific Python libraries, deep learning, and simulation tools into a single ecosystem. AtomAI allows direct applications of the deep convolutional neural networks for atomic and mesoscopic image segmentation converting image and spectroscopy data into class-based local descriptors for downstream tasks such as statistical and graph analysis. For atomically-resolved imaging data, the output is types and positions of atomic species, with an option for subsequent refinement. AtomAI further allows the implementation of a broad range of image and spectrum analysis functions, including invariant variational autoencoders (VAEs). The latter consists of VAEs with rotational and (optionally) translational invariance for unsupervised and class-conditioned disentanglement of categorical and continuous data representations. In addition, AtomAI provides utilities for mapping structure-property relationships via im2spec and spec2im type of encoder-decoder models. Finally, AtomAI allows seamless connection to the first principles modeling with a Python interface, including molecular dynamics and density functional theory calculations on the inferred atomic position. While the majority of applications to date were based on atomically resolved electron microscopy, the flexibility of AtomAI allows straightforward extension towards the analysis of mesoscopic imaging data once the labels and feature identification workflows are established/available. The source code and example notebooks are available at https://github.com/pycroscopy/atomai.
Automated and Autonomous Experiment in Electron and Scanning Probe Microscopy
Mar 22, 2021
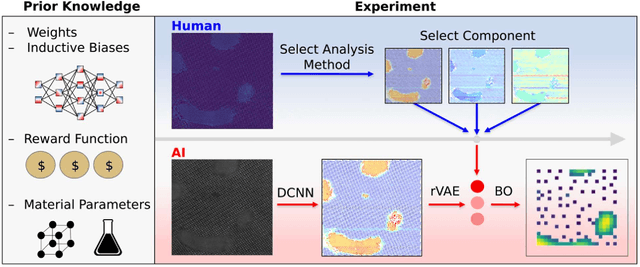
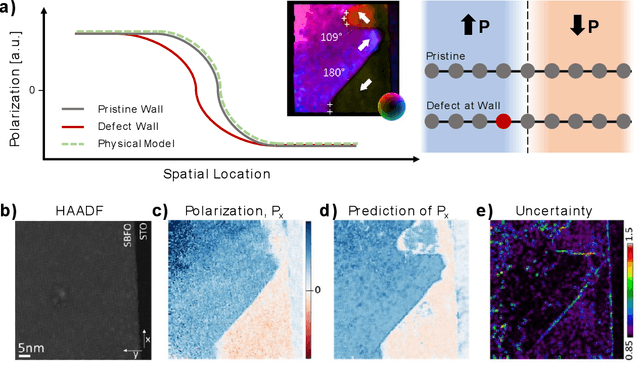
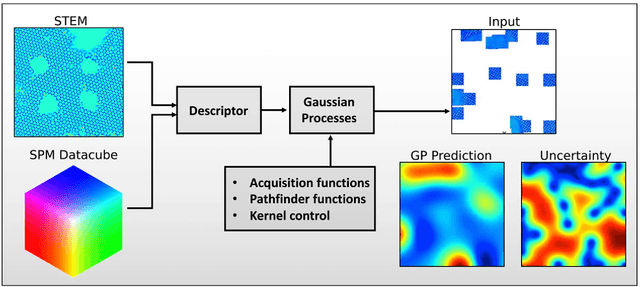
Abstract:Machine learning and artificial intelligence (ML/AI) are rapidly becoming an indispensable part of physics research, with domain applications ranging from theory and materials prediction to high-throughput data analysis. In parallel, the recent successes in applying ML/AI methods for autonomous systems from robotics through self-driving cars to organic and inorganic synthesis are generating enthusiasm for the potential of these techniques to enable automated and autonomous experiment (AE) in imaging. Here, we aim to analyze the major pathways towards AE in imaging methods with sequential image formation mechanisms, focusing on scanning probe microscopy (SPM) and (scanning) transmission electron microscopy ((S)TEM). We argue that automated experiments should necessarily be discussed in a broader context of the general domain knowledge that both informs the experiment and is increased as the result of the experiment. As such, this analysis should explore the human and ML/AI roles prior to and during the experiment, and consider the latencies, biases, and knowledge priors of the decision-making process. Similarly, such discussion should include the limitations of the existing imaging systems, including intrinsic latencies, non-idealities and drifts comprising both correctable and stochastic components. We further pose that the role of the AE in microscopy is not the exclusion of human operators (as is the case for autonomous driving), but rather automation of routine operations such as microscope tuning, etc., prior to the experiment, and conversion of low latency decision making processes on the time scale spanning from image acquisition to human-level high-order experiment planning.
Ensemble learning and iterative training (ELIT) machine learning: applications towards uncertainty quantification and automated experiment in atom-resolved microscopy
Jan 22, 2021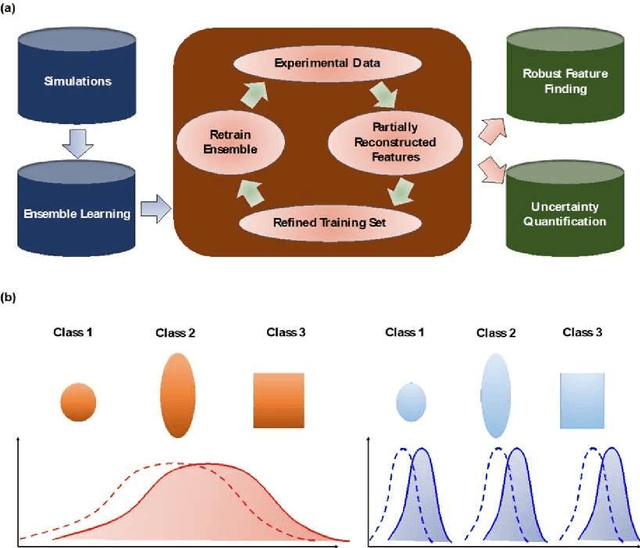
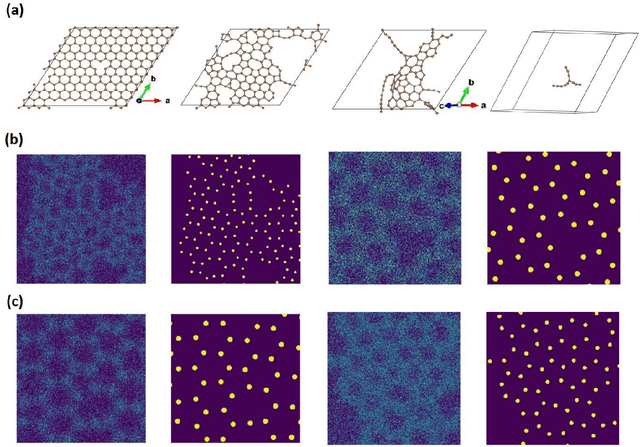
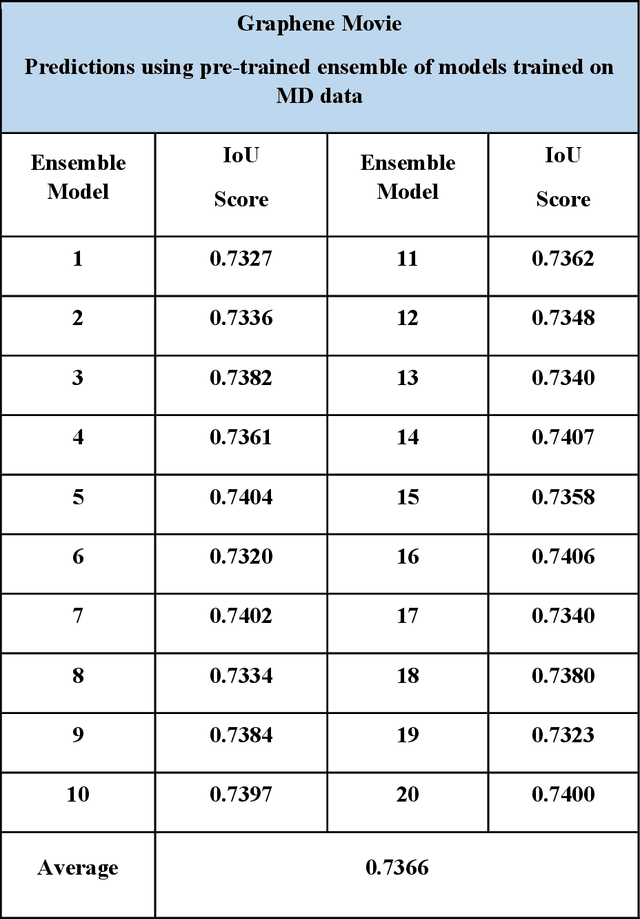
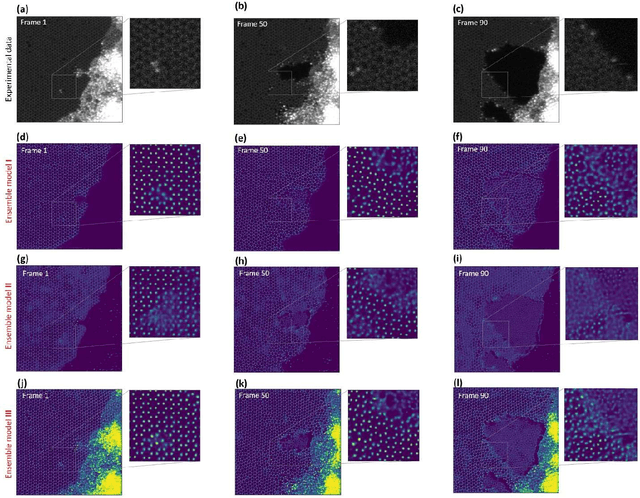
Abstract:Deep learning has emerged as a technique of choice for rapid feature extraction across imaging disciplines, allowing rapid conversion of the data streams to spatial or spatiotemporal arrays of features of interest. However, applications of deep learning in experimental domains are often limited by the out-of-distribution drift between the experiments, where the network trained for one set of imaging conditions becomes sub-optimal for different ones. This limitation is particularly stringent in the quest to have an automated experiment setting, where retraining or transfer learning becomes impractical due to the need for human intervention and associated latencies. Here we explore the reproducibility of deep learning for feature extraction in atom-resolved electron microscopy and introduce workflows based on ensemble learning and iterative training to greatly improve feature detection. This approach both allows incorporating uncertainty quantification into the deep learning analysis and also enables rapid automated experimental workflows where retraining of the network to compensate for out-of-distribution drift due to subtle change in imaging conditions is substituted for a human operator or programmatic selection of networks from the ensemble. This methodology can be further applied to machine learning workflows in other imaging areas including optical and chemical imaging.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge